WO2007139767A2 - Compositions and methods for the delivery of nitric oxide - Google Patents
Compositions and methods for the delivery of nitric oxide Download PDFInfo
- Publication number
- WO2007139767A2 WO2007139767A2 PCT/US2007/012133 US2007012133W WO2007139767A2 WO 2007139767 A2 WO2007139767 A2 WO 2007139767A2 US 2007012133 W US2007012133 W US 2007012133W WO 2007139767 A2 WO2007139767 A2 WO 2007139767A2
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- nox
- protein
- wild
- type
- tengcongensis
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/46—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- C07K14/47—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K38/00—Medicinal preparations containing peptides
- A61K38/02—Peptides of undefined number of amino acids; Derivatives thereof
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P13/00—Drugs for disorders of the urinary system
- A61P13/12—Drugs for disorders of the urinary system of the kidneys
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P17/00—Drugs for dermatological disorders
- A61P17/02—Drugs for dermatological disorders for treating wounds, ulcers, burns, scars, keloids, or the like
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P25/00—Drugs for disorders of the nervous system
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P43/00—Drugs for specific purposes, not provided for in groups A61P1/00-A61P41/00
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P7/00—Drugs for disorders of the blood or the extracellular fluid
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P7/00—Drugs for disorders of the blood or the extracellular fluid
- A61P7/04—Antihaemorrhagics; Procoagulants; Haemostatic agents; Antifibrinolytic agents
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P7/00—Drugs for disorders of the blood or the extracellular fluid
- A61P7/06—Antianaemics
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P7/00—Drugs for disorders of the blood or the extracellular fluid
- A61P7/08—Plasma substitutes; Perfusion solutions; Dialytics or haemodialytics; Drugs for electrolytic or acid-base disorders, e.g. hypovolemic shock
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P9/00—Drugs for disorders of the cardiovascular system
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P9/00—Drugs for disorders of the cardiovascular system
- A61P9/04—Inotropic agents, i.e. stimulants of cardiac contraction; Drugs for heart failure
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P9/00—Drugs for disorders of the cardiovascular system
- A61P9/10—Drugs for disorders of the cardiovascular system for treating ischaemic or atherosclerotic diseases, e.g. antianginal drugs, coronary vasodilators, drugs for myocardial infarction, retinopathy, cerebrovascula insufficiency, renal arteriosclerosis
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61P—SPECIFIC THERAPEUTIC ACTIVITY OF CHEMICAL COMPOUNDS OR MEDICINAL PREPARATIONS
- A61P9/00—Drugs for disorders of the cardiovascular system
- A61P9/12—Antihypertensives
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/195—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from bacteria
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/195—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from bacteria
- C07K14/33—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from bacteria from Clostridium (G)
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/43504—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from invertebrates
- C07K14/43536—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from invertebrates from worms
- C07K14/4354—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from invertebrates from worms from nematodes
- C07K14/43545—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from invertebrates from worms from nematodes from Caenorhabditis
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/43504—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from invertebrates
- C07K14/43563—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from invertebrates from insects
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/43504—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from invertebrates
- C07K14/43563—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from invertebrates from insects
- C07K14/43577—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from invertebrates from insects from flies
- C07K14/43581—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from invertebrates from insects from flies from Drosophila
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K38/00—Medicinal preparations containing peptides
-
- Y—GENERAL TAGGING OF NEW TECHNOLOGICAL DEVELOPMENTS; GENERAL TAGGING OF CROSS-SECTIONAL TECHNOLOGIES SPANNING OVER SEVERAL SECTIONS OF THE IPC; TECHNICAL SUBJECTS COVERED BY FORMER USPC CROSS-REFERENCE ART COLLECTIONS [XRACs] AND DIGESTS
- Y02—TECHNOLOGIES OR APPLICATIONS FOR MITIGATION OR ADAPTATION AGAINST CLIMATE CHANGE
- Y02A—TECHNOLOGIES FOR ADAPTATION TO CLIMATE CHANGE
- Y02A50/00—TECHNOLOGIES FOR ADAPTATION TO CLIMATE CHANGE in human health protection, e.g. against extreme weather
- Y02A50/30—Against vector-borne diseases, e.g. mosquito-borne, fly-borne, tick-borne or waterborne diseases whose impact is exacerbated by climate change
-
- Y—GENERAL TAGGING OF NEW TECHNOLOGICAL DEVELOPMENTS; GENERAL TAGGING OF CROSS-SECTIONAL TECHNOLOGIES SPANNING OVER SEVERAL SECTIONS OF THE IPC; TECHNICAL SUBJECTS COVERED BY FORMER USPC CROSS-REFERENCE ART COLLECTIONS [XRACs] AND DIGESTS
- Y10—TECHNICAL SUBJECTS COVERED BY FORMER USPC
- Y10T—TECHNICAL SUBJECTS COVERED BY FORMER US CLASSIFICATION
- Y10T436/00—Chemistry: analytical and immunological testing
- Y10T436/10—Composition for standardization, calibration, simulation, stabilization, preparation or preservation; processes of use in preparation for chemical testing
- Y10T436/102499—Blood gas standard or control
Definitions
- H-NOX proteins and methods of using them to deliver nitric oxide (NO).
- H-NOX proteins provide a new therapeutic tool for delivering NO to humans and, for veterinary purposes, to animals.
- NO acts as a chemical messenger in the control of many important processes in vivo, including vasodilation, neurotransmission, inflammation, platelet aggregation, and regulation of gastrointestinal and vascular smooth muscle tone.
- GTN nitroglycerin
- NO which is synthesized in endothelial cells, diffuses to smooth muscle cells and activates soluble guanylate cyclase (sGC) to produce cyclic GMP, and thereby induce vasodilation.
- sGC soluble guanylate cyclase
- the clinical mechanism of action of organic nitrates is presumed to require their biotransformation to NO and subsequent activation of sGC.
- organic nitrates cease to be effective in patients after 24-48 hours, due to a phenomenon called tolerance.
- compounds such as ⁇ -blockers and ACE inhibitors are used, although they too have limitations and side effects.
- nitrovasodilators are most useful in treating acute situations where rapid vasodilation is required to alleviate symptoms such as angina and myocardial infarction.
- Prolonged administration of organic nitrates results in reduced efficacy, and the vasculature becomes non-responsive; this tolerance prevents their further use both in chronic and acute cases.
- non-continuous nitrovasodilator use is employed with limited effect.
- other avenues of treatment are employed, typically using a mixed regimen of organic nitrates and NO-independent blood pressure medications, with mixed success.
- a competing theory posits that the response to NO from organic nitrates becomes dampened in the target tissue, perhaps because the generation of NO and the by-products of the reaction eventually inhibit the response to NO, or because acute activation of the NO pathway has a feedback mechanism that desensitizes it to further stimulation.
- This theory is known as end-organ tolerance.
- a unifying theory has been proposed that includes aspects of the biotransformation of organic nitrates as well as end-organ desensitization to NO. Essentially, biotransformation of organic nitrates appears to result in higher levels of superoxide (O 2 " ) in tissues. Superoxide reacts at the rate of diffusion with NO to produce peroxynitrite (OONO).
- This reaction essentially traps and destroys basal NO, preventing it from activating sGC.
- Reduced NO levels leads to vasoconstriction, and OONO ' is a powerful oxidant that damages tissues.
- Prolonged treatment with organic nitrates such as GTN can result in hypertension and tissue damage in patients, and this can be moderated with co-administration of antioxidants such as ascorbate.
- antioxidants such as ascorbate.
- NO is also needed to mediate certain inflammatory responses.
- NO produced by the endothelium inhibits platelet aggregation. Consequently, as NO is bound by cell- free hemoglobin (with or without O 2 bound), platelet aggregation may increase. As platelets aggregate, they release potent vasoconstrictor compounds such as thromboxane A 2 and serotonin. These compounds may act synergistically with the reduced NO levels caused by hemoglobin scavenging to produce significant vasoconstriction. In addition to inhibiting platelet aggregation, NO also inhibits neutrophil attachment to cell walls, which in turn can lead to cell wall damage. Endothelial cell wall damage has been observed with the infusion of certain hemoglobin solutions.
- Hemoglobin-based NO carriers are also hindered by the rapid clearance of cell-free hemoglobin from plasma due the presence of receptors for hemoglobin that remove cell-free hemoglobin from plasma.
- Cell-free hemoglobin may also cause kidney toxicity, possibly due to NO depletion in glomeruli, causing constriction and subsequent dysfunction.
- NO carriers that produce less tolerance are needed.
- NO carriers with a low rate of inactivation by NO in the presence of O 2 are desired, such as NO carriers that have a low NO reactivity and/or a low affinity for O 2 .
- NO carriers with NO dissociation constants or NO dissociation rates that are appropriate for particular clinical or industrial applications are also needed.
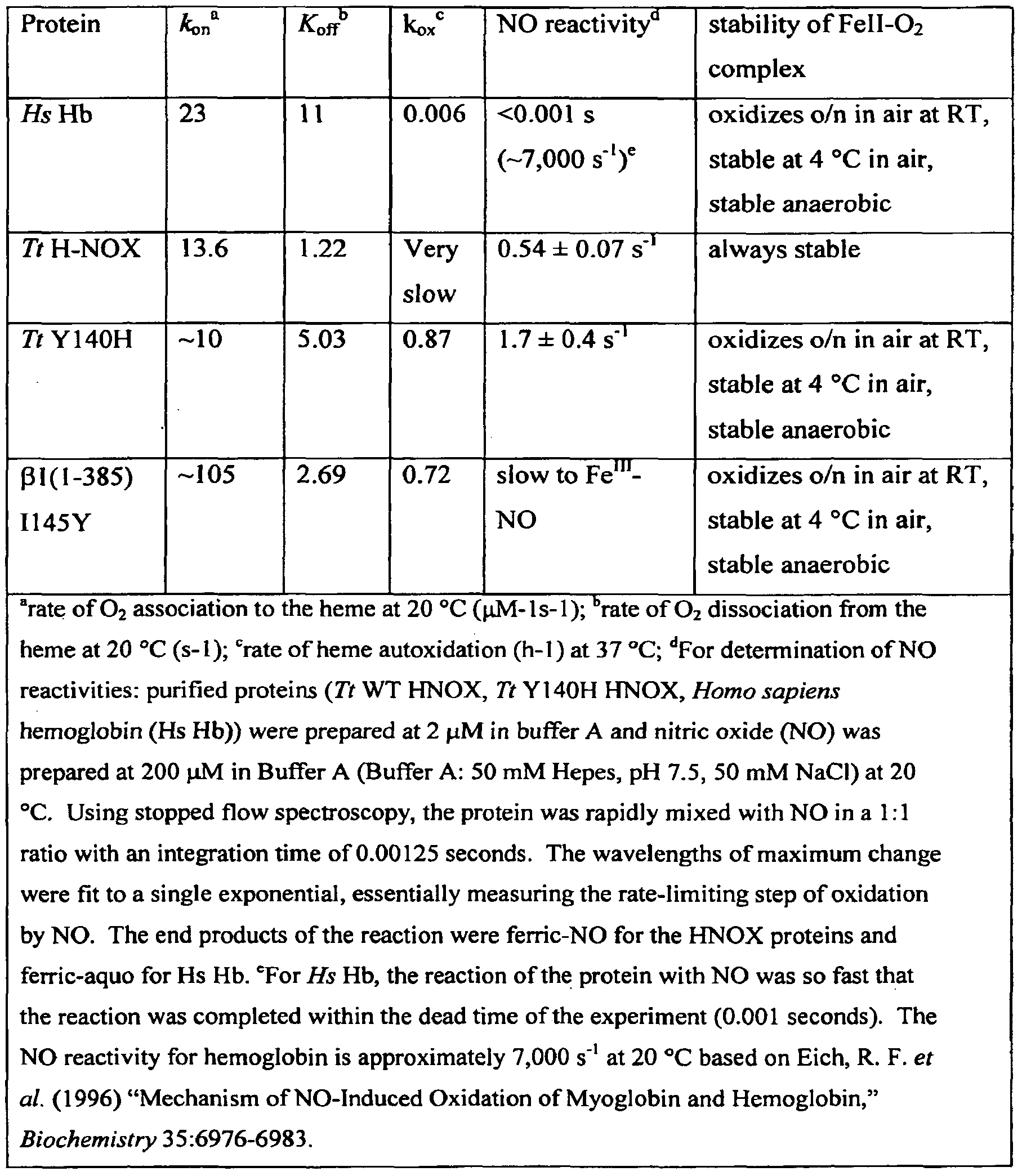
- the present invention is based in part on the surprising discovery that wild-type and mutant H-NOX proteins have a much lower NO reactivity than hemoglobin and thus are desirable NO carriers. If desired, mutations can be introduced into H-NOX proteins to alter their binding of NO and O 2 ligands to further optimize the use of H-NOX proteins as NO carriers. In some embodiments, use of an H-NOX protein as an NO carrier produces less tolerance than the use of current vasodilators, such as organic nitrates.
- the invention features mutant H-NOX proteins. Accordingly, in some embodiments, the invention provides an isolated H-NOX protein comprising at least one mutation that alters the NQ dissociation constant or NO reactivity compared to that of a corresponding wild- type H-NOX protein.
- the NO dissociation constant of the mutant H-NOX protein is within 2 orders of magnitude of that of hemoglobin, and the NO reactivity of the mutant H-NOX protein is at least 10-fold lower than that of hemoglobin.
- the NO reactivity of the mutant H-NOX protein is at least 100-fold lower than that of hemoglobin, such as at least 1,000-fold lower than that of hemoglobin.
- the k ⁇ , k t , or k 2 for NO of the mutant H-NOX protein is between about 1 x 10 " V 1 to about IO s '1 at 37 0 C, such as about 1 . x 10 "4 S 1 to about 0.012 s "1 or about 1 x 10 ⁇ V' to about 1 x 10 ' V 1 at 37 0 C.
- the O 2 dissociation constant of the mutant H-NOX protein is at least about 1 ⁇ M at 37 °C, such as at least about 10 ⁇ M or at least about 50 ⁇ M at 37 "C.
- the invention provides an isolated H-NOX protein comprising at least one mutation that alters the kofr, k ⁇ , or k 2 for NO or alters the O 2 dissociation constant compared to that of a corresponding wild-type H-NOX protein.
- the k o i ⁇ , ki, or k 2 for NO of the mutant H-NOX protein is between about 1 x 10 "4 S 1 to about 10 s "1 at 37 "C, and the O 2 dissociation constant of the mutant H-NOX protein is at least about 1 ⁇ M at 37 0 C.
- the k off , kj, or k 2 for NO of the mutant H-NOX protein is between about 1 x 10 " V to about 0.012 s "1 or about 1 x 10 "4 S “1 to about 1 x lO ' V 1 at 37 "C.
- the O 2 dissociation constant of the mutant H-NOX protein is at least about 10 ⁇ M, such as at least about 50 ⁇ M at 37 "C.
- the NO reactivity of the mutant H-NOX protein is at least 10- fold lower than that of hemoglobin, such as at least 100-fold lower than that of hemoglobin or at least 1, 000-fold lower than that of hemoglobin.
- the invention provides an isolated H-NOX protein selected from the group consisting of T. tengcongensis H-NOX I5A, T. tengcongensis H-NOX I5L, T. tengcongensis H-NOX I5L-P1 15A, T. tengcongensis H-NOX W9F, T. tengcongensis H-NOX W9F- Y140L, T. tengcongensis H-NOX W9F-Y140Hr. tengcongensis H-NOX W9F-N74A, T. tengcongensis H-NOX W9Y, T.
- the ⁇ l or ⁇ 2 protein is derived from a R. norvegicus or H. sapiens ⁇ l or ⁇ 2 protein.
- the invention provides an isolated H-NOX protein selected from the group consisting of T. tengcongensis H-NOX I5A, T. tengcongensis H-NOX 15L, T. tengcongensis H-NOX 15 L-Pl 15 A, T. tengcongensis H-NOX W9F-Y140L, T. tengcongensis H- NOX W9F-Y140H, T. tengcongensis H-NOX W9F-N74A, T. tengcongensis H-NOX W9Y, T.
- the ⁇ l or ⁇ 2 protein is derived from a R. norvegicus or H. sapiens ⁇ l or ⁇ 2 protein.
- the NO dissociation constant of the H-NOX protein is between 0.1 to 10-fold of that of hemoglobin, such as between 0.5 to 2-fold of that of hemoglobin. In some embodiments of the isolated H-NOX proteins, the NO dissociation constant of the H-NOX protein is within 2 orders of magnitude of that of Homo sapiens hemoglobin alpha, such as an NO dissociation constant between 0.1 to 10-fold or between 0.5 to 2-fold of that of Homo sapiens hemoglobin alpha.
- the NO reactivity of the H-NOX protein is at least 10- fold lower than that of Homo sapiens hemoglobin alpha, such as at least 100-fold or 1, 000-fold lower than that of Homo sapiens hemoglobin alpha.
- the NO reactivity of the H-NOX protein is less than about 700 s "1 at 20 "C, such as less than about 600 s "1 , 500 400 300 200 s ⁇ 100 s " ⁇ 75 s “1 , 50 s 1 , 25 s “1 , 20 s 1 , 10 s “1 , 50 s “1 , 3 s “1 , 2 s “1 , 1.8 s '1 , 1.5 s “1 , 1.2 s “1 , 1.0 s *1 , 0.8 s " ⁇ 0.7 s "1 , or 0.6 s '1 at 20 0 C.
- the O 2 dissociation constant of the H-NOX protein is at least about 1 ⁇ M at 37 °C, such as at least about 10 ⁇ M or at least about 50 ⁇ M at 37 "C.
- the k off , k], or k 2 for NO of the H-NOX protein is between about 1 x 10 ⁇ s "1 to about 10 s ⁇ ' at 37 "C, and the O 2 dissociation constant of the H-NOX protein is at least about 1 ⁇ M at 37 "C.
- the kof f , ki, or k 2 for NO of the H-NOX protein is between about 1 x 10 "4 S ' 1 to about 10 s '1 at 37 “C, and the NO reactivity of the H-NOX protein is less than about 700 s "1 at 20 °C (e.g., less than about 600 s 1 , 500 s ⁇ ⁇ 100 s ⁇ ⁇ 20 s "1 , or 1.8 s 'at 20 °C).
- the O 2 dissociation constant of the H-NOX protein is at least about 1 ⁇ M at 37 0 C, and the NO reactivity of the H-NOX protein is less than about 700 s "1 at 20 "C (e.g., less than about 600 s ' ⁇ 500 s "1 , 100 s "1 , 20 s ' ⁇ or 1.8 s 'at 20 °C).
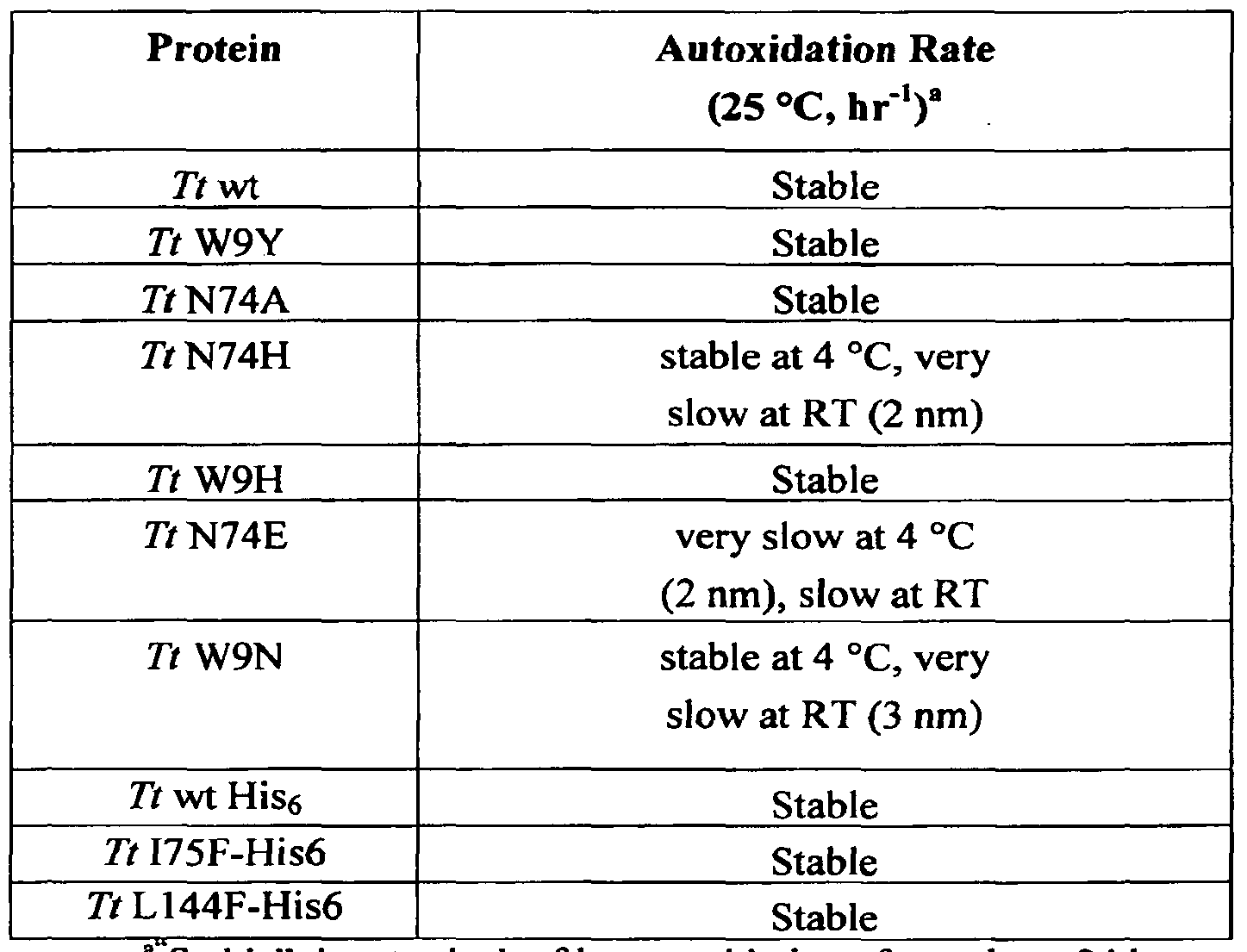
- the rate of heme autoxidation of the H-NOX protein is less than about 1 h ' 'at 37 C C.
- the k Of ⁇ , ki, or k 2 for NO of the H- NOX protein is between about 1 x 10 "4 S “1 to about 10 s '1 at 37 °C, and the rate of heme autoxidation of the H-NOX protein is less than about 1 h 'at 37 "C.
- the O 2 dissociation constant of the H-NOX protein is at least about 1 ⁇ M at 37 "C, and the rate of heme autoxidation of the H-NOX protein is less than about 1 h 'at 37 "C.
- the rate of heme autoxidation of the H-NOX protein is less than about 1 h 'at 37 0 C, and the NO reactivity of the H-NOX protein is less than about 700 s " 1 at 20 "C (e.g., less than about 600 500 s "1 , 100 20 s "1 , or 1.8 s 'at 20 'C).
- the H-NOX protein contains one or more mutations (e.g., 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 mutations) compared to the H-NOX protein from which it was derived.
- the H-NOX protein contains less than 20, 15, 12, 10, 9, 8, 7, 6, 5, 4, 3, or 2 mutations compared to the H-NOX protein from which it was derived.
- the H-NOX protein has at least one distal pocket mutation.
- the H-NOX protein has at least one mutation that is not in the distal pocket.
- the H-NOX protein has at least one mutation in which a residue that corresponds to Tyrl40 of 7! tengcongensis H-NOX or PheI42 of L. pneumophila 2 is replaced by any other amino acid.
- the H-NOX protein has at least two mutations, wherein at least one mutation is the replacement of a residue that corresponds to Tyrl40 of T. tengcongensis H-NOX or Phel42 of L. pneumophila 2 by any other amino acid.
- the mutation in the H-NOX protein corresponds to a Y 140F mutation or a Yl 4OL mutation of T. tengcongensis or a F142Y mutation of L. pneumophila 2.
- the H-NOX protein is a deletion that contains the first 194, 217, or 385 amino acids of an H-NOX protein such as R. norvegicus or H. sapiens ⁇ l or ⁇ 2 protein.
- the H-NOX protein is derived from a mammalian protein ⁇ e.g., a human protein such as ⁇ l).
- the H-NOX protein derived from a bacterial protein (e.g., a T. tengcongensis protein).
- the H-NOX protein is covalently bound to another molecule or moiety, such as polyethylene glycol. Heme may or may not be bound to the H-NOX protein.
- NO is bound to the H-NOX protein.
- the H-NOX protein is a fusion protein that includes an H-NOX domain and part or all of another protein, such as albumin (e.g., human serum albumin).
- the H-NOX protein is not T. tengcongensis H-NOX Y40L, wild-type T. tengcongensis H-NOX, wild-type R. norvegicus sGC, or L. pneumophilia 2 H-NOX F 142 Y. In some embodiments of the isolated H-NOX proteins, the H- NOX protein is not T. tengcongensis H-NOX F78Y/Y140L. In some embodiments of the isolated H-NOX proteins, the H-NOX protein is not wild-type L. pneumophilia 2 H-NOX, wiJd-type H. sapiens ⁇ l H-NOX, R.
- norvegicus sGC ⁇ l H-NOX (1-385), wild-type R. norvegicus ⁇ l H-NOX, wild-type D. melangaster ⁇ l H-NOX, wild-type D. melangaster CG14885-PA H-NOX, wild-type C. elegans GCY-35 H-NOX, wild-type N. punctiforme H-NOX, wild-type C. crescentus H-NOX, wild-type S. oneidensis H-NOX, or wild-type C. acetobutylicum H-NOX.
- the H-NOX protein is not T. tengcongensis H-NOX W9F, T.
- the H- NOX protein is not R. norvegicus ⁇ 2(l-217), R. norvegicus ⁇ l(l -194), R. norvegicus ⁇ l(l-385), or R.
- the H-NOX protein is not T. tengcongensis H-NOX W9F, T. tengcongensis H-NOX Y 140F, or H. sapiens ⁇ l H- NOX (1-385) I145Y. In some embodiments of the isolated H-NOX proteins, the H-NOX protein is not T. tengcongensis H-NOX Y140H, H. sapiens ⁇ l I140Y, or H. sapiens ⁇ l I145Y. In some embodiments of the isolated H-NOX proteins, the H-NOX protein is not T.
- norvegicus sGC ⁇ l H-NOX (1-385) I145Y, R. norvegicus sGC ⁇ l H-NOX H105G, R. norvegicus sGC ⁇ l H-NOX H105F, R. norvegicus sGC ⁇ l H-NOX 1145 Y, wild-type R. norvegicus ⁇ l H-NOX, wild-type D. melangaster ⁇ l H-NOX, wild- type D. melangaster CG14885-PA H-NOX, wild-type C. elegans GCY-35 H-NOX 5 wild-type N. punctiforme H-NOX, wild-type C. crescentus H-NOX, wild-type S.
- the H- NOX protein is not any of the following H-NOX proteins that are listed by their gene name, followed by their species abbreviation and Genbank Identifiers (such as the following protein sequences available as of May 21, 2006; May 22, 2006; May 21, 2007; or May 22, 2007): Npun5905_Npu 23129606, alr2278_Ana 17229770, SO2144_Sone_24373702, Mdeg 1343_Mde_23027521 , VCA0720_Vch_ 15601476, CC2992_Ccr_ 16127222, Rsph2043_Rhsp_22958463 (gi:46l92757), Mmcl0739_Mcsp_22999020, Tar4_Tte_20807169, Ddes2822_Dde_23475919, CAC3243_
- the H-NOX protein is not any of the following H- NOX proteins that are listed by their organism name and Pfam database accession number (such as the following protein sequences available as of May 21, 2006; May 22, 2006; May 17, 2007; May 21, 2007; or May 22, 2007): Caenorhabditis briggsae Q622M5 CAEBR, Caenorhabditis briggsae Q61P44_CAEBR, Caenorhabditis briggsae Q61 R54 C AEBR 5 Caenorhabditis briggsae Q61V90_CAEBR, Caenorhabditis briggsae Q61A94 CAEBR, Caenorhabditis briggsae Q60TP4 CAEBR, Caenorhabditis briggsae Q60M 1 O CAEBR, Caenorhabditis elegans GCY37_CAEEL, Caenorhabdititis briggsae Q622M
- the invention features a recombinant nucleic acid encoding any one or more of the mutant H-NOX proteins described herein.
- the nucleic acid includes a segment of or the entire nucleic acid sequence of any of the nucleic acids shown in FIGS. 2-4D or 8A-8DD.
- the nucleic acid encodes a fusion protein that includes an H-NOX domain and part or all of another protein, such as albumin (e.g., human serum albumin).
- the nucleic acid includes at least about any of 50, 100, 150, 200, 300, 400, 500, 600, 700, 800, or more contiguous nucleotides from an H-NOX nucleic acid and contains one or more mutations (e.g., 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 mutations) compared to the H-NOX nucleic acid from which it was derived.
- a mutant H-NOX nucleic acid contains less than about any of 20, 15, 12, 10, 9, 8, 7, 6, 5, 4, 3, or 2 mutations compared to the H-NOX nucleic acid from which it was derived.
- the invention also features degenerate variants of any nucleic acid encoding a mutant H-NOX protein.
- the invention provides a vector that includes any one or more of the mutant H-NOX nucleic acids described herein. In another aspect, the invention features a cell that includes any one or more of the mutant H-NOX nucleic acids described herein. In one aspect, the invention features a cell that includes any vector described herein.
- the invention features a method of producing an H-NOX protein. This method involves culturing a cell having a nucleic acid encoding any one or more of the mutant H- NOX proteins described herein under conditions suitable for production of the mutant H-NOX protein. In some embodiments, the invention further includes the step of purifying the mutant H- NOX protein.
- the invention features pharmaceutical compositions that include one or more H-NOX proteins, such as any of the wild-type or mutant H-NOX proteins described herein.
- the pharmaceutical composition includes a pharmaceutically acceptable amount of an H-NOX protein described herein and a pharmaceutically acceptable carrier.
- the k off , ki, or k 2 for NO of the H-NOX protein is between about 1 x 10 ' V 1 to about 10 s "1 at 37 °C, and the O 2 dissociation constant of the H-NOX protein is at least about 1 ⁇ M at 37 °C.
- the NO dissociation constant of the H-NOX protein is within 2 orders of magnitude of that of hemoglobin, and the NO reactivity of the H-NOX protein is at least 10-fold lower than that of hemoglobin.
- the NO dissociation constant of the H-NOX protein is within 2 orders of magnitude of that of Homo sapiens hemoglobin alpha, such as an NO dissociation constant between 0.1 to 10-fold or between 0.5 to 2-fold of that of Homo sapiens hemoglobin alpha.
- the NO reactivity of the H-NOX protein is at least 10-fold lower than that of Homo sapiens hemoglobin alpha, such as at least 100-fold or 1, 000-fold lower than that of Homo sapiens hemoglobin alpha.
- the H-NOX protein is a wild-type protein.
- the H-NOX protein is a mutant protein as described herein.
- the H-NOX protein has at least one mutation that alters the NO dissociation constant, the k ⁇ , f r for NO, the kj for NO, the k 2 for NO, the O 2 dissociation constant, the NO stability, the NO reactivity the rate of heme autoxidation, or any combination of two or more of the foregoing compared to that of a corresponding wild-type protein.
- the H- NOX protein is a selected from the group consisting of wild-type T. tengcongensis H-NOX, T.
- tengcongensis H-NOX Y 140H T. tengcongensis H-NOX Y140A, T. tengcongensis I75F-His6, T. tengcongensis I75F, T. tengcongensis L144F-His6, T. tengcongensis L144F, L. pneumophilia 2 H-NOX F 142 Y, wild-type L. pneumophilia 1 H-NOX, wild-type L. pneumophilia 2 H-NOX, L. pneumophilia 2 F9W-F142Y, wild-type D. desulfuricans H-NOX, D.
- H. sapiens ⁇ l H-NOX H-NOX(728-899)
- D. desulfuricans H-NOX Y139L wild-type H. sapiens ⁇ l H-NOX, H. sapiens ⁇ l 114OY, H sapiens ⁇ l 1145 Y, H. sapiens ⁇ l( 1-385), H. sapiens ⁇ l(l-385) I145Y, H. sapiens ⁇ l(l-385) I145H, H. sapiens ⁇ l(l-194), H. sapiens ⁇ l(l-194) I145Y, H. sapiens ⁇ l(l-194) L9W-I145Y, H. sapiens ⁇ 2(l-217), H. sapiens ⁇ 2(l-217) 1142Y, H.
- norvegicus ⁇ 2(l-217) R. norvegicus ⁇ 2(l-217) 1142Y
- R. norvegicus ⁇ l H-NOX H105G R. norvegicus ⁇ l H-NOX H105F
- R. norvegicus sGC ⁇ l H-NOX C78S R. norvegicus sGC ⁇ l H-NOX C78E
- C. botulinum H-NOX(I-175) C. botulinum H- NOX(1-186), wild-type C. acetobutylicum H-NOX, C. acetobutylicum H-NOX(1-197), C.
- H-NOX Taurus H-NOX, wild-type R. norvegicus; wild-type ⁇ C laevis H-NOX, wild-type O. latipes H-NOX, wild-type O. curivatus H-NOX, wild- type F. rubripes H-NOX, wild-type A. gambiae H-NOX, wild-type M. sexta H-NOX; wild-type C. elegans gcy-31, C. elegans gcy-32, wild-type C. elegans gcy-33, wild-type C. elegans gcy-34, wild- type C. elegans gcy-35, wild-type C.
- the H-NOX protein is a selected from the group consisting of wild-type R. norvegicus sGC, wild-type R. norvegicus ⁇ 1(1-385), R. norvegicus ⁇ l(l-217), R. norvegicus ⁇ l(l-194), wild-type T. tengcongensis H-NOX, T.
- the pharmaceutical composition includes one or more liposomes or nanoparticles that include or encapsulate the H-NOX protein.
- the H-NOX protein is not
- the H-NOX protein is not T. tengcongensis H-NOX F78Y/Y140L. In some embodiments of the pharmaceutical compositions, the H-NOX protein is not wild-type L. pneumophilia 2 H-NOX, wild- type H. sapiens ⁇ l H-NOX, R. norvegicus sGC ⁇ l H-NOX (1 -385), wild-type R.
- the H-NOX protein is not T. tengcongensis H- NOX W9F, T. tengcongensis H-NOX Y 140F, R.
- the H-NOX protein is not R. norvegicus ⁇ 2(l-217), R. norvegicus ⁇ 1(1- 194), R. norvegicus ⁇ l(l-385), or R. norvegicus ⁇ l(l-385) I145Y.
- the H-NOX protein is not T. tengcongensis H-NOX W9F, T. tengcongensis H-NOX Y 140F, or H. sapiens ⁇ l H-NOX (1-385) 1145 Y. In some embodiments of the pharmaceutical compositions, the H-NOX protein is not T. tengcongensis H-NOX Y 140H, H. sapiens ⁇ l 114OY, or H. sapiens ⁇ l 1145 Y. In some embodiments of the pharmaceutical compositions, the H-NOX protein is not T. tengcongensis H-NOX Y40L, T.
- norvegicus sGC ⁇ l H-NOX (1-385) I145Y, R. norvegicus sGC ⁇ l H- NOX H105G, R. norvegicus sGC ⁇ l H-NOX H105F, R. norvegicus sGC ⁇ l H-NOX 1145 Y, wild- type R. norvegicus ⁇ l H-NOX, wild-type D. melangaster ⁇ l H-NOX, wild-type D. melangaster CG14885-PA H-NOX, wild-type C. elegans GCY-35 H-NOX, wild-type N. punctiforme H-NOX, wild-type C. crescentus H-NOX, wild-type S.
- the H-NOX protein is not any of the following H-NOX proteins that are listed by their gene name, followed by their species abbreviation and Genbank Identifiers (such as the following protein sequences available as of May 21, 2006; May 22, 2006; May 21, 2007; or May 22, 2007): Npun5905_Npu_23129606, alr2278_Ana_l 7229770, SO2144_Sone_24373702, Mdegl343_Mde_23027521, VCA0720_Vch_l 5601476, CC2992_Ccr_l 6127222, Rsph2043_Rhsp_22958463 (gi:46192757), Mmc 10739_Mcsp_22999020, Tar4_Tte_20807169, Ddes2822_Dde_23475919, CAC3243_
- the H-NOX protein is not any of the following H-NOX proteins that are listed by their organism name and Pfam database accession number (such as the following protein sequences available as of May 21, 2006; May 22, 2006; May 17, 2007; May 21, 2007; or May 22, 2007): Caenorhabditis briggsae Q622M5 CAEBR, Caenorhabditis briggsae Q61P44 CAEBR, Caenorhabditis briggsae Q61R54 CAEBR, Caenorhabditis briggsae Q61 V90_CAEBR, Caenorhabditis briggsae Q61A94 CAEBR, Caenorhabditis briggsae Q60TP4 CAEBR, Caenorhabditis briggsae Q60M 1 O CAEBR, Caenorhabditis elegans GCY37 CAEEL, Caenorhabditis elegans G
- H-NOX proteins described herein may be used in any of the pharmaceutical compositions described herein.
- the H-NOX protein may or may not have heme and/or NO bound and may or may not be covalently bound to another molecule or moiety, such as polyethylene glycol.
- the H-NOX protein is a fusion protein that includes an H-NOX domain and part or all of another protein, such as albumin (e.g., human serum albumin).
- the invention provides methods of delivering NO to an individual (e.g., a mammal, such as a primate (e.g., a human, a monkey, a gorilla, an ape, a lemur, etc), a bovine, an equine, a porcine, a canine, or a feline) using an H-NOX protein.
- the individual is suffering from or at risk for a cardiovascular condition, hypertension, a condition exacerbated by hypertension, a vasoconstrictive condition, stroke, or a functional NO deficiency.
- the condition exacerbated by hypertension is heart failure, renal failure, or a stroke.
- the invention provides a method of delivering
- the k off , ki, or k 2 for NO of the H-NOX protein is between about 1 x 10 "4 S "1 to about 10 s "1 at 37 "C, and the O 2 dissociation constant of the H-NOX protein is at least about 1 ⁇ M at 37 0 C.
- the NO dissociation constant of the H-NOX protein is within 2 orders of magnitude of that of hemoglobin, and the NO reactivity of the H-NOX protein is at least 10-fold lower than that of hemoglobin.
- NO is bound to the H-NOX protein prior to the administration of the H-NOX protein to the individual. In some embodiments of the methods, NO is not bound to the H-NOX protein prior to the administration of the H-NOX protein to the individual, and the H-NOX protein transports NO from one location in the individual to another location in the individual. In some embodiments of the methods, the H-NOX protein is administered orally, rectally, or to the blood of the individual. In particular embodiments of the methods, the H-NOX protein is administered to the blood of the individual. In some embodiments of the methods, the H-NOX protein is administered to the individual at least twice.
- the NO dissociation constant of the H-NOX protein is within 2 orders of magnitude of that of Homo sapiens hemoglobin alpha, such as an NO dissociation constant between 0.1 to 10-fold or between 0.5 to 2-fold of that of Homo sapiens hemoglobin alpha.
- the NO reactivity of the H-NOX protein is at least 10-fold lower than that of Homo sapiens hemoglobin alpha, such as at least 100-fold or 1, 000-fold lower than that of Homo sapiens hemoglobin alpha.
- the H-NOX protein is a wild-type protein.
- the H- NOX protein is a mutant protein as described herein.
- the H-NOX protein has at least one mutation that alters the NO dissociation constant, the k ⁇ , f r for NO, the ki for NO, the k 2 for NO, the O 2 dissociation constant, the NO stability, the NO reactivity the rate of heme autoxidation, or any combination of two or more of the foregoing compared to that of a corresponding wild-type protein.
- the H-NOX protein is a selected from the group consisting of wild-type T. tengcongensis H-NOX, T.
- tengcongensis H-NOX Y 140H T. tengcongensis H-NOX Y 140A, T. tengcongensis 175F-His6, T. tengcongensis I75F, T. tengcongensis L144F-His6, T. tengcongensis L144F, L. pneumophilia 2 H- NOX F142Y, wild-type L. pneumophilia 1 H-NOX, wild-type L. pneumophilia 2 H-NOX, L. pneumophilia 2 F9W-F142Y, wild-type D. desulfuricans H-NOX, D.
- H. sapiens ⁇ l H-NOX H. sapiens ⁇ l 1140Y 5 H. sapiens ⁇ l I145Y, H. sapiens ⁇ l(l-385), H. sapiens ⁇ l(l-385) I145Y, H. sapiens ⁇ l(l-385) 1145H, H. sapiens ⁇ l(l-194), H. sapiens ⁇ l(l -194) I 145Y, H. sapiens ⁇ l(l-194) L9W-I 145Y, H. sapiens ⁇ 2(l-217), H.
- sapiens ⁇ 2(l-217) I142Y, H. sapiens ⁇ l H-NOX H105G, H. sapiens ⁇ l H-NOX H105F, H. sapiens ⁇ l H-NOX C78S, H. sapiens ⁇ l H-NOX C78E, wild-type R. norvegicus ⁇ l H- NOX, R. norvegicus ⁇ l(l-385), R. norvegicus ⁇ l(l-385) I145Y, R. norvegicus ⁇ l(l-385) I145H, R. norvegicus ⁇ l(l-194), R. norvegicus ⁇ l(l-194) I145Y, R.
- musculus H-NOX wild-type C. familiaris H-NOX, wild-type B. Taurus H-NOX, wild-type R. norvegicus; wild-type X. laevis H-NOX, wild-type O. latipes H-NOX, wild-type O. curivatus H-NOX, wild-type F. rubripes H-NOX, wild-type A. gambiae H-NOX, wild-type M. sexta H-NOX; wild-type C. elegans gcy-31, C. elegans gcy-32, wild-type C. elegans gcy-33, wild-type C.
- the H-NOX protein is a selected from the group consisting of wild-type R. norvegicus sGC, wild-type R. norvegicus ⁇ l(l - 385), R. norvegicus ⁇ l(l-217), R.
- one or more liposomes or nanoparticles that include or encapsulate the H-NOX protein are examples of the H-NOX protein.
- the H-NOX protein is not T. tengcongensis H-
- H-NOX Y40L wild-type T. tengcongensis H-NOX, wild-type R. norvegicus sGC, or L. pneumophilia 2 H-NOX F 142 Y.
- the H-NOX protein is not T. tengcongensis H-NOX F78Y/Y140L.
- the H-NOX protein is not wild-type L. pneumophilia 2 H-NOX, wild-type H. sapiens ⁇ l H-NOX, R. norvegicus sGC ⁇ l H-NOX (1-385), wild-type R. norvegicus ⁇ l H-NOX, wild-type D.
- the H-NOX protein is not T. tengcongensis H-NOX W9F, T. tengcongensis H-NOX Y 140F, R. norvegicus sGC ⁇ l H-NOX H105G, R.
- the H-NOX protein is not R. norvegicus ⁇ 2(l-217), R. norvegicus ⁇ 1(1 -194), R. norvegicus ⁇ l(l-385), or R. norvegicus ⁇ l(l-385) I145Y. In some embodiments of the methods, the H-NOX protein is not T.
- the H-NOX protein is not T. tengcongensis H-NOX Y140H, H. sapiens ⁇ l I140Y, or H. sapiens ⁇ l 1145Y. In some embodiments of the methods, the H-NOX protein is not T. tengcongensis H-NOX Y40L, T. tengcongensis H-NOX F78Y/Y140L, T. tengcongensis H-NOX W9F, T.
- norvegicus sGC ⁇ l H-NOX H105F R. norvegicus sGC ⁇ l H-NOX 1145 Y
- wild-type R. norvegicus ⁇ l H-NOX wild-type D. melangaster ⁇ l H-NOX, wild- type D. melangaster CG14885-PA H-NOX, wild-type C. elegans GCY-35 H-NOX, wild-type N. punctiforme H-NOX 5 wild-type C. crescentus H-NOX, wild-type S. oneidensis H-NOX, or wild- type C. acetobutylicum H-NOX.
- the H-NOX protein is not any of the following H-NOX proteins that are listed by their gene name, followed by their species abbreviation and Genbank Identifiers (such as the following protein sequences available as of May 21, 2006; May 22, 2006; May 21, 2007; or May 22, 2007): Npun5905_Npu_23129606, alr2278_Ana_l 7229770, SO2144_Sone_24373702, Mdegl343_Mde_23027521 , VCA0720_Vch_l 5601476, CC2992_Ccr_l 6127222, Rsph2043_Rhsp_22958463 (gi:46192757), Mmc 10739_Mcsp_22999020, Tar4_Tte_20807169, Ddes2822_Dde_23475919, CAC3243_Cac_l 5896488 , gcy-31_Ce_ 17568389, CG148
- the H-NOX protein is not any of the following H-NOX proteins that are listed by their organism name and Pfam database accession number (such as the following protein sequences available as of May 21, 2006; May 22, 2006; May 17, 2007; May 21, 2007; or May 22, 2007): Caenorhabditis briggsae Q622M5_CAEBR, Caenorhabditis briggsae Q61P44 CAEBR, Caenorhabditis briggsae Q61R54 CAEBR, Caenorhabditis briggsae Q61V90_CAEBR, Caenorhabditis briggsae Q61A94_CAEBR, Caenorhabditis briggsae Q60TP4 CAEBR, Caenorhabditis briggsae Q60M10 CAEBR, Caenorhabditis elegans GCY37 CAEEL, Caenorhabditis elegans GCY
- the H-NOX protein does not have a mutation in the Y-S-R motif, which includes Tyrl35, Serl37, and Argl39 of human H-NOX.
- the H-NOX protein may or may not have heme and/or NO bound and may or may not be covalently bound to another molecule or moiety, such as polyethylene glycol.
- the H-NOX protein is a fusion protein that includes an H-NOX domain and part or all of another protein, such as albumin (e.g., human serum albumin).
- the invention features kits that include one or more H-NOX proteins.
- the invention provides a kit that includes an H-NOX protein and instructions for using the kit to deliver NO to an individual.
- the kom ki, or k 2 for NO of the H-NOX protein is between about 1 x 10 " V 1 to about 10 s "1 at 37 "C, and the O 2 dissociation constant of the H-NOX protein is at least about 1 ⁇ M at 37 "C.
- the NO dissociation constant of the H-NOX protein is within 2 orders of magnitude of that of hemoglobin, and the NO reactivity of the H-NOX protein is at least 10- fold lower than that of hemoglobin.
- the H-NOX protein may or may not have heme and/or NO bound and may or may not be covalently bound to another molecule or moiety, such as polyethylene glycol.
- the H- NOX protein is a fusion protein that includes an H-NOX domain and part or all of another protein, such as albumin (e.g., human serum albumin).
- the invention features an H-NOX protein (such as any of the wild-type or mutant proteins described herein) for use as a medicament.
- the invention features an H-NOX protein for use in a method of delivering NO to an individual.
- the H-NOX protein is used to treat any condition for which delivery of NO is beneficial, such as a cardiovascular condition, hypertension, a condition exacerbated by hypertension (e.g., heart failure, renal failure, or a stroke) , a vasoconstrictive condition, stroke, or a functional NO deficiency.
- the invention features the use of an H-NOX protein (such as any of the wild-type or mutant proteins described herein) for the manufacture of a medicament, such as a medicament for delivering NO to an individual.
- the invention features the use of an H-NOX protein for delivering NO to an individual.
- the H- NOX protein is used to treat any condition for which delivery of NO is beneficial, such as a cardiovascular condition, hypertension, a condition exacerbated by hypertension (e.g., heart failure, renal failure, or a stroke) , a vasoconstrictive condition, stroke, or a functional NO deficiency.
- FIG. IA is a picture of the three dimensional structure of distal pocket residues of
- FIG. 1 A is based on the three-dimensional structure of T. tengcongensis H-NOX reported by Pellicena, P. et al. (August 31, 2004). "Crystal Structure of An Oxygen-Binding Heme Domain Related to Soluble Guanylate Cyclases," Proc Natl. AcadSci USA 101(35):12854-12859. [0036]
- FIG. 1 B is a stereo side view of the three dimensional structure of T. tengcongensis
- HNOX illustrating structural features of the H-NOX domain.
- the protein fold is represented by ribbon diagrams.
- the heme, dioxygen ligand, and proximal histidine are shown as ball-and-stick models, ⁇ -helices are labeled A-G according to the nomenclature shown in FIG. 5B.
- ⁇ -strands are labeled 1-4.
- FlG. IB is from Pellicena, P. et al. (August 31, 2004). "Crystal Structure of An Oxygen-Binding Heme Domain Related to Soluble Guanylate Cyclases," Proc Natl. AcadSci USA 101(35): 12854-12859.
- FIGS. 1C- IH are pictures of the three dimensional structure of T. tengcongensis
- FIG. 1C-1H illustrating exemplary distal pocket residues in T. tengcongensis HNOX.
- the following residues depicted in FIGS. 1C-1H are the main residues comprising the H-NOX distal pocket: Thr4, Ile5, Thr8, Trp9, Trp67, Asn74, Ile75, Phe78, Phe82, Tyrl40, and Leul44, which are contained within helices A, D, E, and G.
- FIGS. 1C- IH were created using PYMOL (DeLano Scientific, LLP).
- FIG. 2 is a sequence alignment of the following H-NOX proteins that bind or are predicted to bind O 2 and NO: Majority (SEQ ID NO: I); Ce.
- H-NOX proteins are predicted to bind O 2 as well as NO because they have a tyrosine at the position corresponding to Y140 of T. tengcongensis H-NOX.
- FIG. 3A-3D are a sequence alignment of the following H-NOX proteins that bind or are predicted to bind NO but not O 2 : Majority (SEQ ID NO: 10); Dm. sGC betal protein (SEQ ID NO:1 1); sGC betal protein (SEQ ID NO:12); hs. sGC betal protein (SEQ ID NO:13); hs. beta2 protein (SEQ ID NO: 14); Ms. sGC betal protein (SEQ ID NO: 15); Mm. sGCbetal protein (SEQ ID NO: 16); Np. betal HD-like (SEQ ID NO: 17); Tr.
- Majority SEQ ID NO: 10
- Dm. sGC betal protein SEQ ID NO:1 1
- sGC betal protein SEQ ID NO:12
- hs. sGC betal protein SEQ ID NO:13
- hs. beta2 protein SEQ ID NO: 14
- Ms. sGC betal protein SEQ ID NO: 15
- sGC betal protein (SEQ ID NO: 18); Anopheles_gambiae
- FIGS. 4A-4D are a sequence alignment of H-NOX proteins from FIGS. 2 and 3A-
- 3D Majority (SEQ ID NO:27); Dm. sGC betal protein (SEQ ID NO:1 1); sGC betal protein (SEQ ID NO: 12); hs. sGC betal protein (SEQ ID NO: 13); hs. beta2 protein (SEQ ID NO: 14); Mm. sGC betal protein (SEQ ID NO:16); Np. betalHD-like (SEQ ID NO:17); Tr.
- sGC betal protein (SEQ ID NO: 18); Chlamydomonas_reinhardtii
- sGc beta 3 protein SEQ ID NO:7; CG 14885 (SEQ ID NO:5); and Dm. sGC short variant (SEQ ID NO:6).
- the alignment was generated using the default parameters in the program MegAlign.
- Dm. sGC betal protein denotes Drosophila melanogaster ⁇ l H-NOX
- sGC betal protein denotes Rattus norvegicus ⁇ l H-NOX
- hs. sGC betal protein denotes Homo sapiens ⁇ l H-NOX
- hs. beta2 protein denotes Homo sapiens ⁇ 2 H-NOX
- sGC betal protein denotes Mus musculus ⁇ l H-NOX
- Np. betalHD-like denotes Nostoc punctiforme H-NOX
- Tr. sGC betal protein denotes Takifugu rubripes ⁇ l H-NOX
- XP_310919 denotes Anopheles gambiae ⁇ l H-NOX
- NP_001011632 denotes Apis mellifera ⁇ l H-NOX
- sGC betal protein denotes Bos taurus ⁇ 1 H-NOX
- AAR02 denotes Chlamydomonas reinhardtii ⁇ l H-NOX
- BAC98396 denotes Oryzias curvinotus ⁇ l H-NOX
- BAA76691 denotes Oryzias latipes ⁇ l H-NOX
- X denotes Strongylocentrotus purpuratus ⁇ l H-NOX
- NP_001018042+” denotes Sus scrofa ⁇ l H-NOX
- gcy-31a denotes Caenorhabditis elegans Gcy-31a H-NOX
- gcy-33 denotes Caenorhab
- HNOX denotes Clostridium acetobutiylicum H- NOX
- T. betal HD-like denotes Thermoanaerobacter tengcongensis H-NOX
- Ms. sGc beta 3 protein denotes Manduca sexta ⁇ 3 H-NOX
- CG 14885" denotes Drosophila melanogaster CG 14885 H-NOX
- Dm. sGC short variant denotes Drosophila melanogaster Gcy-88-E-S H- NOX
- Dm. CG4154 HNOX denotes Drosophila melanogaster CG4154 H-NOX.
- 5A is a sequence alignment of members of the H-NOX family.
- the sequence numbering is that of T. tengcongensis H-NOX.
- Invariant residues are indicated by a "V”, very highly conserved residues are indicated by "s”.
- H-NOX tengcongensis H-NOX is indicated by a "H.”
- Predicted distal pocket tyrosine residues that may stabilize an Fe"- ⁇ 2 complex in other H- NOX proteins are: position 70 for Caenorhabditis elegans GCY-35; position 140 in Drosophila melanogaster CG14885-PA; position 138 of Caenorhabditis elegans GCY-35; position 140 of Clostridium acetobutylicum; numbered according to Thermoanaerobacter tengcongensis.
- Accession numbers are: Homo sapiens ⁇ l [gi:2746083] (SEQ ID NO:28), Rattus norvegicus ⁇ l [gi:27127318] (SEQ ID NO:29), Drosophila melangaster ⁇ l [gi:861203] (SEQ ID NO:30), Drosophila melangaster CG14885-PA [gi:23171476] (SEQ ID NO:31), Caenorhabditis elegans GCY-35 [gi:52782806] (SEQ ID NO:32), Nostoc punctiforme [gi:23129606] (SEQ ID NO:33), Caulobacter crescentus [gi: 16127222] (SEQ ID NO:34), Shewanella oneidensis [gi .24373702] (SEQ ID NO:35), Legionella pneumophila (ORF 2) [CUCGC_272624] (SEQ ID NO:36), Clostridium aceto
- FlG. 5B is a sequence alignment of exemplary H-NOX domains. The secondary structure annotations and the numbering on top of the alignment correspond to the H-NOX domain from T. tengcongensis. ⁇ -helices are represented by spirals, and ⁇ -strands by arrows. The distal pocket is defined by ⁇ -helices ⁇ A, ⁇ D, ⁇ E, and ⁇ G.
- Pubmed/NCBI accession numbers are as follows: Ther tengcongensis gi 120807169 I (SEQ ID NO:39), Clos_acetobutylicum gi I 15896488 I (SEQ ID NO:40), Clos tetani Gl:75543266 (SEQ ID NO:41), Desu_desulfuricans gi 123475919 I (SEQ ID NO:42), Vibr vulnificus gi I 27361734
- FIG. 5 B The alignments were generated using the program MULTALIN (Corpet, F. (1988) Nucleic Acids Res. 16:10881-10890), and FIG. 5B was prepared using the program ESPRIPT (Gouet, P. et al. (1999) Bioinformatics 15: 305-308.)-
- FIGS. 6A and 6B are pictures of the three dimensional structure of the heme environment of the T. tengcongensis H-NOX domain.
- FIG.S. 6A and 6B are from Pellicena, P. et al. (August 31, 2004). "Crystal Structure of An Oxygen-Binding Heme Domain Related to Soluble Gua ⁇ ylate Cyclases," Proc Natl. AcadSci USA 101(35):12854-12859.
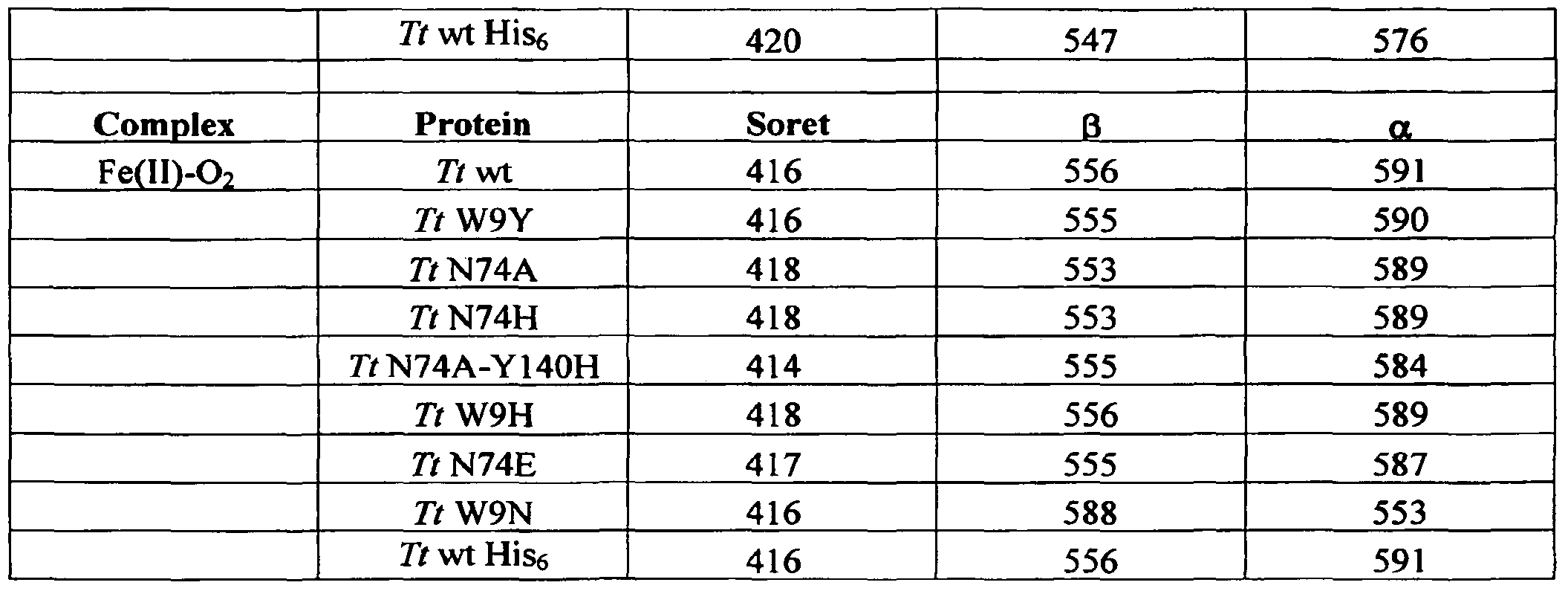
- FIGS. 7A-7F are graphs of the UV-visible spectroscopy of H-NOX proteins after anaerobic reduction (Fe" unligated complexes; top line in each graph) before and after being exposed to air (Fe"-O 2 complexes; bottom line in each graph) for Tt H-NOX (FIG. 7A), Tt Y 140L (FIG. 7B), Tt W9F-Y140L (FIG. 7C), Tt F78Y-Y140L (FIG. 7D), L2 H-NOX and L2 F142Y (FIG. 7E), and ⁇ l(l-385) and ⁇ l(l-385) I145Y (FIG. 7F).
- FIGS. 7A-7F are from Boon, E. M. et al. (2005). "Molecular Basis For NO Selectivity in Soluble Guanylate Cyclase,” Nature Chem. Biol. 1 :53-59. [004S] FIGS.
- 8A-8DD contain polynucleotide sequences of exemplary nucleic acids that encode H-NOX proteins and the amino acid sequences of the corresponding H-NOX proteins (SEQ ID NOS:53-162). DETAILED DESCRIPTION OF THE INVENTION
- the present invention is based in part on the surprising discovery that H-NOX proteins have a much lower NO reactivity than hemoglobin. This intrinsic low NO reactivity (and high NO stability) makes wild-type and mutant H-NOX proteins desirable NO carriers because of the lower probability of inactivation of H-NOX proteins by NO in the presence of O 2 . Importantly, the presence of a distal pocket tyrosine in some H-NOX proteins (Pellicena, P. et al. (August 31, 2004). "Crystal Structure of An Oxygen-Binding Heme Domain Related to Soluble Guanylate Cyclases," Proc Natl.
- AcadSci USA 101 (35): 12854- 12859) is suggestive of undesirable, AfgA NO reactivity, contraindicating use as an NO carrier.
- a Mycobacterium tuberculosis hemoglobin protein with a structurally analogous distal pocket tyrosine, reacts extremely rapidly with NO, and is used by the Mycobacterium to effectively scavenge and avoid defensive NO produced by an infected host (Ouellet, H. et al. (April 30, 2002).
- "Truncated Hemoglobin HbN Protects Mycobacterium Bovis From Nitric Oxide” Proc. Natl. Acad. Sci. USA 99(9):5902-5907).
- H-NOX proteins actually have a much lower NO reactivity than that of hemoglobin making their use as NO carriers possible.
- the usefulness of H-NOX proteins as NO carriers can be improved by modifying their affinities for NO or O 2 to maximize the amount of NO that is bound to the H-NOX protein and to reduce the amount of H-NOX protein that is oxidized by the reaction of NO with O 2 bound to the H-NOX protein.
- the affinity of H-NOX proteins for NO or O 2 and the ability of H-NOX proteins to discriminate between NO and O 2 ligands can be altered by the introduction of one or more amino acid mutations, allowing H-NOX proteins to be tailored to bind NO or O 2 with desired affinities.
- the dissociation constant or dissociation rate for NO or O 2 binding by H-NOX proteins can be altered the introduction of a single amino acid mutation. Additional mutations can be introduced to further alter the affinity for NO and/or O 2 .
- the H-NOX protein family can therefore be manipulated to exhibit improved or optimal kinetic and thermodynamic properties for NO delivery.
- mutant H-NOX proteins have been generated with altered dissociation constants and/or dissociation rates for NO binding that improve the usefulness of H-NOX proteins for a variety of clinical and industrial applications.
- an H-NOX protein with a low affinity for O 2 (such as an O 2 dissociation constant of at least about 1 ⁇ M at 37 "C) is used to minimize the amount of O 2 that binds the H-NOX protein, thereby facilitating the binding of NO to the H-NOX protein and reducing the amount of H-NOX protein that is oxidized due to the reaction of NO with O2 bound to the heme of the H-NOX protein.
- H-NOX proteins for NO delivery.
- Organic nitrates are effective for a limited length of time due to tolerance. Since H-NOX proteins delivery NO directly to individuals without requiring the bioconversion of nitrates to NO, the effectiveness of H-NOX proteins as NO carriers is not limited by inhibition of this bioconversion pathway.
- hemoglobin-based NO carriers Major limitations of hemoglobin-based NO carriers are their high affinity for O 2 and their propensity to be inactivated by NO. As mentioned above, destruction of even low levels of NO by hemoglobin- based carriers can have serious effects on the tonic resting state of the vasculature and organs and leads to hypertension and gastrointestinal distress. Intra- and inter-molecular cross-linking have been used to minimize the toxicity of hemoglobin-based vehicles when used as oxygen carriers ("Blood Substitutes," R. Winslow ed. Academic Press, 2006). While these modifications overcame some of the severe toxicity issues related to extravasation of hemoglobin, the high NO reactivity remained.
- H-NOX proteins have a much lower NO reactivity than hemoglobin. This lower reactivity leads to less destruction of NO, O 2 , and H-NOX protein since less NO reacts with O 2 bound to the H-NOX protein.
- the ability to select H-NOX proteins with desired dissociation constants and dissociation rates for NO can also minimize side-effects by preventing too much NO from being released (causing hypotension) and prevent NO from being released at undesired sites ⁇ e.g., sites that are not vasoconstricted).
- Engineering H-NOX proteins to bind and deliver NO with minimal NO reactivity provides a new blood gas NO carrier where the H-NOX proteins deliver NO without being inactivated by NO.
- H-NOX proteins represent an important alternative that overcomes the persistent problem of tolerance with current nitrovasodilators.
- the use of H- NOX proteins as delivery vehicles for NO provides a new therapeutic venue for treating diseases exacerbated by chronic hypertension.
- H-NOX protein means a protein that has an H-NOX domain (named for Heme-Nitric oxide and OXygen binding domain).
- An H-NOX protein may or may not contain one or more other domains in addition to the H-NOX domain.
- H-NOX proteins are members of a highly-conserved, well-characterized family of hemoproteins (Iyer, L. M. et al. (February 3, 2003).
- H- NOX proteins are also referred to as Pfam 07700 proteins or HNOB proteins (Pfam - A database of protein domain family alignments and Hidden Markov Models, Copyright (C) 1996-2006 The Pfam Consortium; GNU LGPL Free Software Foundation, Inc., 59 Temple Place - Suite 330, Boston, MA 021 11-1307, USA).
- an H-NOX protein has, or is predicted to have, a secondary structure that includes six alpha-helices, followed by two beta-strands, followed by one alpha-helix, followed by two beta-strands.
- An H-NOX protein can be an apoprotein that is capable of binding heme or a holoprotein with heme bound.
- H-NOX protein can covalently or non- covalently bind a heme group. Some H-NOX proteins bind NO but not O 2 , and others bind both NO and O 2 . H-NOX domains from facultative aerobes that have been isolated bind NO but not O 2 . H-NOX proteins from obligate aerobic prokaryotes, C. elegans, and D. melanogaster bind NO and O 2 Mammals have two H-NOX proteins: ⁇ l and ⁇ 2. An alignment of mouse, rat, cow, and human H-NOX sequences shows that these species share >99% identity.
- the H- NOX domain of an H-NOX protein or the entire H-NOX protein is at least about any of 10, 15, 20, 25, 30, 40, 50, 60, 70, 80, 90, 95, 97, 98, 99, or 99.5% identical to that of the corresponding region of a naturally-occurring Thermoanaerobacter tengcongensis H-NOX protein or a naturally- occurring sGC protein (e.g., a naturally-occurring sGC ⁇ l protein).
- an H-NOX protein may optionally contain one or more mutations relative to the corresponding naturally-occurring H-NOX protein.
- the H-NOX protein includes one or more domains in addition to the H-NOX domain.
- the H-NOX protein includes one or more domains or the entire sequence from another protein.
- the H- NOX protein may be a fusion protein that includes an H-NOX domain and part or all of another protein, such as albumin (e.g., human serum albumin). In some embodiments, only the H-NOX domain is present.
- a "protein” includes proteins and fragments of proteins whether isolated from natural sources, produced by recombinant techniques, or chemically synthesized.
- a protein may have one or more modifications, such as a post-translational modification (e.g., glycosylation, etc) or any other modification (e.g., PEGylation, etc).
- the protein may contain one or more non -naturally-occurring amino acids (e.g., such as an amino acid with a side chain modification).
- the H-NOX protein has at least about 50, 100, 150, 181, 200, 250, 300, 350, 400, or more amino acids.
- the H-NOX proteins may include from about 50 to about 600 amino acids, such as about 100 to about 500 amino acids, about 150 to about 400 amino acids, about 150 to about 300 amino acids, or about 175 to about 200 amino acids.
- H-NOX proteins from any genus or species can be used in the compositions, kits, and methods described herein.
- the H-NOX protein is a protein from a mammal (e.g., a primate (e.g., human, monkey, gorilla, ape, lemur, etc), a bovine, an equine, a porcine, a canine, or a feline), an insect, a yeast, or a bacteria or is derived from such a protein.
- exemplary mammalian H-NOX proteins include wild-type human and rat soluble guanylate cyclase (such as the ⁇ l subunit).
- H-NOX proteins include wild-type mammalian H-NOX proteins, e.g. H. sapiens, M. musculus, C. familiaris, B. taunts and R. norvegicus; and wild-type non-mammalian vertebrate H-NOX proteins, e.g.. X. laevis, O. latipes, O. curivatus, and F. rubripes.
- non-mammalian wild-type NO-binding H-NOX proteins include wild-type H-NOX proteins of D. melanogaster, A.
- examples of non-mammalian wild- type O2-binding H-NOX proteins include wild-type H-NOX proteins of C. elegans gcy-31, gcy-32, gcy-33, gcy-34, gcy-35, gcy-36, and gcy-37; D. melanogaster CG14885, CG14886, and CG4154; and M. sexta beta-3;
- examples of prokaryotic wild-type H-NOX proteins include T. tengcongensis, V. cholera, V. fischerii, N. punctiforme, D. desulfuricans, L. pneumophila 1, L. pneumophila 2, and C. acetobutylicum.
- NCBI Accession numbers for exemplary H-NOX proteins include the following:
- Exemplary H-NOX protein also include the following H-NOX proteins that are listed by their gene name, followed by their species abbreviation and Genbank Identifiers (such as the following protein sequences available as of May 21, 2006; May 22, 2006; May 21, 2007; or May 22, 2007, which are each hereby incorporated by reference in their entireties): Npun5905_Npu_23129606, alr2278_Ana_l 7229770, SO2144_Sone_24373702, Mdegl343_Mde_23027521, VCA0720_Vch_15601476,CC2992_Ccr_16127222, Rsph2043 Rhsp_22958463 (gi:46192757), Mmcl0739_Mcsp_22999020,
- H-NOX proteins include the following H-NOX proteins that are listed by their organism name and Pfam database accession number (such as the following protein sequences available as of May 21, 2006; May 22, 2006; May 17, 2007; May 21, 2007; or May 22, 2007, which are each hereby incorporated by reference in their entireties): Caenorhabditis briggsae Q622M5_CAEBR, Caenorhabditis briggsae Q61P44_CAEBR, Caenorhabditis briggsae Q61R54 CAEBR, Caenorhabditis briggsae Q61V90 CAEBR, Caenorhabditis briggsae Q61A94 CAEBR, Caenorhabditis briggsae Q60TP4 CAEBR, Caenorhabditis briggsae Q60M10_CAEBR, Caenorhabditis elegans GCY37 CAEEL, Caenorhabditis elegans
- H-NOX proteins and nucleic acids which may be suitable for use in the pharmaceutical compositions and methods described herein, can be identified using standard methods. For example, standard sequence alignment and/or structure prediction programs can be used to identify additional H-NOX proteins and nucleic acids based on the similarity of their primary and/or predicted protein secondary structure with that of known H-NOX proteins and nucleic acids.
- the Pfam database uses defined alignment algorithms and Hidden Markov Models (such as Pfam 21.0) to categorize proteins into families, such as the H-NOX protein family (Pfam - A database of protein domain family alignments and Hidden Markov Models, Copyright (C) 1996-2006 The Pfam Consortium; GNU LGPL Free Software Foundation, Inc., 59 Temple Place - Suite 330, Boston, MA 021 1 1-1307, USA).
- Standard databases such as the swissprot-trembl database (world-wide web at "expasy.org", Swiss Institute of Bioinformatics Swiss-Prot group CMU - 1 rue Michel Servet CH-121 1 Geneva 4 , Switzerland) can also be used to identify members of the H-NOX protein family.
- the secondary and/or tertiary structure of an H- NOX protein can be predicted using the default settings of standard structure prediction programs, such as PredictProtein (630 West, 168 Street, BB217, New York, N. Y. 10032, USA). Alternatively, the actual secondary and/or tertiary structure of an H-NOX protein can be determined using standard methods.
- the H-NOX protein has the same amino acid in the corresponding position as any of following distal pocket residues in T. tengcongensis H-NOX: Thr4, Ile5, Thr8, Trp9, Trp67, Asn74, Ile75, Phe78, Phe82, Tyrl40, Leu 144, or any combination of two or more of the foregoing.
- the H-NOX protein has a proline or an arginine in a position corresponding to that of Prol 15 orArgl35 of T. tengcongensis H-NOX, respectively, based on sequence alignment of their amino acid sequences.
- the H-NOX protein has a histidine that corresponds to HislO5 of R. norvegicus ⁇ l H-NOX.
- the H-NOX protein has or is predicted to have a secondary structure that includes six alpha-helices, followed by two beta-strands, followed by one alpha-helix, followed by two beta-strands. This secondary structure has been reported for H-NOX proteins.
- a newly identified H-NOX protein can be tested to determine whether it binds heme using standard methods.
- the ability of an H-NOX protein to function as an NO carrier can be tested by determining whether the H-NOX protein binds NO using standard methods, such as those described herein.
- one or more of the mutations described herein can be introduced into the H-NOX protein to optimize its characteristics as an NO carrier. For example, one or more mutations can be introduced to alter its NO dissociation constant, ko t r for NO, ki for NO, k 2 for NO, O 2 dissociation constant, NO stability, NO reactivity, rate of heme autoxidation, or any combination of two or more of the foregoing. Standard techniques such as those described herein can be used to measure these parameters.
- mutant H-NOX proteins may be derived by mutagenesis from these or other natural wild-type source sequences (e.g., the sequences listed in FlG. 2-4D or 8A-8DD or any other sequence described herein).
- derived from refers to the source of the protein into which one or more mutations is introduced.
- a protein that is "derived from a mammalian protein” refers to protein of interest that results from introducing one or more mutations into the sequence of a wild-type (i.e., a sequence occurring in nature) mammalian protein.
- an H-NOX protein may contain one or more mutations, such as a mutation that alters the NO dissociation constant, the k ot r for NO, the O 2 dissociation constant, the kofr for O2, the rate of heme autoxidation, the NO reactivity, the NO stability, or any combination of two or more of the foregoing compared to that of the corresponding wild-type protein.
- Panels of engineered H-NOX proteins may be generated by random mutagenesis followed by empirical screening for requisite or desired dissociation constants, dissociation rates, NO- reactivity, stability, physio-compatibility, or any combination of two or more of the foregoing in view of the teaching provided herein using techniques as described herein and, additionally, as known by the skilled artisan.
- mutagenesis can be selectively targeted to particular regions or residues such as distal pocket residues apparent from the experimentally determined or predicted three-dimensional structure of an H-NOX protein (FIG. IA herein; and see, for example, Boon, E. M. et al. (2005).
- a mutant protein means a protein with one or more mutations compared to a protein occurring in nature.
- the mutant protein has a sequence that differs from that of all proteins occurring in nature.
- the amino acid sequence of the mutant protein is at least about any of 10, 15, 20, 25, 30, 40, 50, 60, 70, 80, 90, 95, 97, 98, 99, or 99.5% identical to that of the corresponding region of a protein occurring in nature.
- the mutant protein is a protein fragment that contains at least about any of 25, 50, 75, 100, 150, 200, 300, or 400 contiguous amino acids from a full-length protein.
- Sequence identity can be measured, for example, using sequence analysis software with the default parameters specified therein (e.g., Sequence Analysis Software Package of the Genetics Computer Group, University of Wisconsin Biotechnology Center, 1710 University Avenue, Madison, WI 53705). This software program matches similar sequences by assigning degrees of homology to various amino acids replacements, deletions, and other modifications.
- sequence analysis software with the default parameters specified therein (e.g., Sequence Analysis Software Package of the Genetics Computer Group, University of Wisconsin Biotechnology Center, 1710 University Avenue, Madison, WI 53705). This software program matches similar sequences by assigning degrees of homology to various amino acids replacements, deletions, and other modifications.
- a “mutation” means an alteration in a reference nucleic acid or amino acid sequence occurring in nature.
- Exemplary nucleic acid mutations include an insertion, deletion, frameshift mutation, silent mutation, nonsense mutation, or missense mutation.
- the nucleic acid mutation is not a silent mutation.
- Exemplary protein mutations include the insertion of one or more amino acids (e.g., the insertion of 2, 3, 4, 5, 6, 7, 8, 9, or 10 amino acids), the deletion of one or more amino acids (e.g., a deletion of N-terminal, C-terminal, and/or internal residues, such as the deletion of at least about any of 5, 10, 15, 25, 50, 75, 100, 150, 200, 300, or more amino acids or a deletion of about any of 5, 10, 15, 25, 50, 75, 100, 150, 200, 300, or 400 amino acids), the replacement of one or more amino acids (e.g., the replacement of 2, 3, 4, 5, 6, 7, 8, 9, or 10 amino acids), or combinations of two or more of the foregoing.
- the deletion of one or more amino acids e.g., the insertion of 2, 3, 4, 5, 6, 7, 8, 9, or 10 amino acids
- the deletion of one or more amino acids e.g., a deletion of N-terminal, C-terminal, and/or internal residues, such as the deletion of at least about any of 5,
- An exemplary functional truncation of an H-NOX protein includes residues 1-385 of the ⁇ l sequence.
- a mutant protein has at least one amino acid alteration compared to a protein occurring in nature.
- a mutant nucleic acid sequence encodes a protein that has at least one amino acid alteration compared to a protein occurring in nature.
- the nucleic acid is not a degenerate version of a nucleic acid occurring in nature that encodes a protein with an amino acid sequence identical to a protein occurring in nature. The nomenclature used in referring to a particular amino acid mutation first identifies the wild-type amino acid, followed by the residue number and finally the substitute amino acid.
- Y 140L means that tyrosine has been replaced by a leucine at residue number 140.
- An "evolutionary conserved mutation” is the replacement of an amino acid in one protein by an amino acid in the corresponding position of another protein in the same protein family.
- Exemplary evolutionary conserved mutations are listed in Table IA. In Table 1 A, mutations are numbered/annotated according to the sequence of human ⁇ l H-NOX, but are analogous for all H-NOX sequences. Thus, the corresponding position in any other H-NOX protein can be mutated to the indicated residue.
- Phe4 of human ⁇ l H-NOX can be mutated to a tyrosine since other H-NOX proteins have a tyrosine in this position.
- the corresponding phenylalanine residue can be mutated to a tyrosine in any other H-NOX protein.
- the one or more mutations are confined to evolutionarily conserved residues.
- the one or more mutations may include at least one evolutionarily conserved mutation and at least one non-evolutionarily conserved mutation. If desired, these mutant H-NOX proteins are subjected to empirical screening for NO/O 2 dissociation constants, NO- reactivity, stability, and physio-compatibility in view of the teaching provided herein.
- the mutation is a distal pocket mutation, such as mutation of a residue in alpha-helix A, D, E, or G (Pellicena, P. et al. (August 31, 2004). "Crystal Structure of An Oxygen-Binding Heme Domain Related to Soluble Guanylate Cyclases," Proc Natl. Acad Sci USA 101 (35): 12854-12859).
- Exemplary distal pocket mutations are listed in Table IB. In Table IB, mutations are numbered/annotated according to the sequence of human ⁇ 1 H-NOX, but are analogous for all H-NOX sequences.
- the residue at each indicated position can be changed to any other naturally or non-naturally-occurring amino acid (denoted "X").
- Such mutations can produce H-NOX proteins with a variety of desired affinity, stability, and reactivity characteristics.
- the mutation is a heme distal pocket mutation.
- a crucial molecular determinant that prevents O 2 binding in NO-binding members of the H-NOX family is the lack of a H-bond donor in the distal pocket of the heme.
- the mutation alters H-bonding between the H-NOX domain and the ligand within the distal pocket.
- the mutation disrupts an H-bond donor of the distal pocket and/or imparts reduced O 2 ligand-binding relative to the corresponding wild-type H-NOX domain.
- Exemplary distal pocket residues include Thr4, Ile5, Thr8, Trp9, Trp67, As ⁇ 74, Ile75, Phe78, Phe82, Tyrl40, and Leul44of T. tengcongensis H-NOX and the corresponding residues in any other H-NOX protein.
- the H-NOX protein has one or more mutations outside of the distal pocket. Examples of residues that can be mutated but are not in the distal pocket include Prol 15 and Argl35 of T. tengcongensis H-NOX. In some embodiments, the mutation is in the proximal pocket which includes His 105 as a residue that Iigates to the heme iron.
- the amino acid sequence of the H-NOX protein is not identical to the sequence of a protein that is produced by an organism in nature. In some embodiments, the amino acid sequence of the H-NOX protein is not identical to a sequence found in any database on May 21, 2006 or May 22, 2006 (such as all known sequences predicted or known to be an H-NOX nucleic acid or amino acid sequence). In some embodiments, the amino acid sequence of the H-NOX protein is not identical to a sequence found in any database on May 21, 2007 or May 22, 2007 (such as all known sequences predicted or known to be an H-NOX nucleic acid or amino acid sequence).
- amino acids in an H-NOX protein can be mutated to the corresponding amino acids in a human H-NOX.
- one or more amino acids on the surface of the tertiary structure of a non-human H-NOX protein can be mutated to the corresponding amino acid in a human H-NOX proteins.
- mutation of one or more surface amino acids may be combined with mutation of two or more distal pocket residues, mutation of one or more residues outside of the distal pocket (e.g., a mutation in the proximal pocket), or combinations of two or more of the foregoing.
- Table 2 can be mutated to any other amino acid.
- the invention also relates to any combination of mutation described herein, such as double, triple, or higher multiple mutations.
- combinations of any of the mutations described herein can be made in the same H-NOX protein.
- mutations in equivalent positions in other mammalian or non-mammalian H-NOX proteins are also encompassed by this invention.
- residues other than the ones mentioned in Table 2 can also be mutated.
- Exemplary mutant H-NOX proteins comprise one or more mutations that impart altered NO or O 2 ligand-binding relative to the corresponding wild-type H- NOX domain and are operative as a physiologically compatible mammalian NO blood gas carrier.
- T. tengcongensis 15A refers to the replacement of isoleucine by alanine at the fifth position in T. tengcongensis H-NOX.
- the same isoleucine to alanine mutation can be made in the corresponding residue in any other H-NOX protein (this residue may or may not be the fifth residue in the sequence of other H-NOX proteins).
- the H-NOX protein has at least one mutation in which a residue that corresponds to Ile5, Trp9, Asn74, Prol 15, Argl 35, or Tyrl40 of T. tengcongensis H- NOX, 1145 of ⁇ l(l-385), or Phel42 of L. pneumophila 2 is replaced by any other amino acid.
- the H-NOX protein has at least two mutations, wherein at least one mutation is the replacement of a residue that corresponds to Ile5, Trp9, Asn74, Prol 15, Argl35, or Tyrl40 of T. tengcongensis H-NOX, 1145 of ⁇ l(l-385), or Phel42 of L.
- the mutation in the H-NOX protein corresponds to a I5A mutation, a 15L mutation, a W9F mutation, a Y 140F mutation, a Y 140L mutation, a Y 140H mutation, a W9F Y140H double mutation, or a F78Y Y140F double mutation of T. tengcongensis or a I145Y mutation of ⁇ l.
- the mutation in the H-NOX protein corresponds to a W9Y mutation, a W9H mutation, a W9N mutation, a N74H mutation, a N74E mutation, a N74A mutation, a P1 15A mutation, a R135Q mutation, a I5L Pl 15A double mutant, a N74A Y 140H double mutant, or a W9F N74A double mutant of T. tengcongensis.
- at least one C-terminal amino acid (such as at least about 50 contiguous C-terminal amino acids or between about 25 to about 200 contiguous C-terminal amino acids) in the H-NOX protein has been removed compared to the corresponding wild-type protein (such as R. norvegicus or H. sapiens ⁇ l).
- T. tengcongensis T. tengcongensis
- Lp L. pneumophila
- Dd D. desulfuricans
- Vc V. cholera
- Np N. punctiforme
- Cb C. botulinium
- Ca C. acetobutylicum
- rat human
- C. elegans Ce
- any of the wild-type or mutant H-NOX proteins can be modified and/or formulated using standard methods to enhance therapeutic or industrial applications.
- a variety of methods are known in the art for insulating such agents from immune surveillance, including crosslinking, PEGylation, carbohydrate decoration, etc. (e.g., Rohlfs, R. J. et al. (May 15, 1998). "Arterial Blood Pressure Responses to Cell-Free Hemoglobin Solutions And The Reaction With Nitric Oxide," J. Biol. Chem. 273(20): 12128-12134; Migita, R. et al. (June 1997).
- H-NOX protein Fusing an H- NOX protein with a human protein such as human serum albumin can increase the serum half-life, viscosity, and colloidal oncotic pressure.
- an H-NOX protein is modified during or after its synthesis to decrease its immunogenicity and/or to increase its plasma retention time.
- H-NOX proteins can also be encapsulated (such as encapsulation within liposomes or nanoparticles).
- an H-NOX protein has a similar or improved NO dissociation constant, O 2 dissociation constant, NO k of r, O 2 k off , NO reactivity, autoxidation rate, plasma retention time, or any combination of two or more of the foregoing compared to any currently used compound for delivering NO, such as any organic nitrate for bioconversion into NO.
- an H-NOX protein has a low affinity for O 2 (such as an O 2 dissociation constant of at least about 1 ⁇ M at 37 0 C) or no detectable affinity for O 2 . Since little, if any, O 2 is bound to the H-NOX protein, there is minimal oxidation by NO of O 2 bound to the heme of the H-NOX protein. Thus, minimal NO, O 2 , and H-NOX protein is inactivated by this NO oxidation.
- hemoglobin means a protein or a mutant thereof from the well- characterized family of hemoglobins, which are iron-containing O 2 -transport metalloproteins in red blood cells. Purified, stroma-free, human hemoglobin has a kinetic K D for O 2 of about 200-500 nM. This value is subunit dependent.
- a 6-coordinate Fe-NO complex is meant a 6-coordinate ferrous-nitrosyl that produces a UV-Vis Soret peak at approximately 416-422 nm, as described, e.g., by Boon, E. M. et al., (August 2006), "Nitric Oxide Binding to Prokaryotic Homologs of the Solube Guanylate Cyclase ⁇ l HONOX Domain,” J. Biol. Chem.
- a 5-coordinate Fe H -NO complex is meant a 5-coordinate ferrous-nitrosyl that produces a UV-Vis Soret peak at approximately 397-400 nm, as described, e.g., by Boon, E. M.
- a "k off” means a dissociation rate, such as the rate of release of NO or
- the k o ff, ki, or k 2 for NO for the H-NOX protein is between about 1 x 10"V 1 to about 10 s 1 at 37 °C, such as between about 1 x 10 "4 S "1 to about 0.012 s "1 , about 1 x 10 ⁇ "1 to about 0.007 s * ⁇ about 0.005 s "1 to about 0.01 1 s "1 , or about 1 x 10 "4 S 1 to about 1 x 10 ' V at 37 0 C.
- the k o f f for O 2 for an H-NOX protein is between about 1 to about 1,000 s "1 at 37 ⁇ C, such as about 1 to about 50 s ' ⁇ about 50 to about 100 about 100 to about 250 about 500 to about 750 s "1 , or about 750 to about 1 ,000 s "1 at 37 °C.
- a "Ic 0n " is meant an association rate, such as the rate of binding of NO or O 2 to a protein.
- a lower numerical lower Ic 0n indicates a slower rate of association.
- the Ic 0n for O 2 for an H-NOX protein is between about 0.14 to about 60 ⁇ WC's '1 at 20 0 C, such as about 6 to about 60 about 6 to 12 ⁇ M 's '1 , about 15 to about 60 ⁇ M " ⁇ s " ', about 5 to about 18 ⁇ M “ s “1 , or about 6 to about 15 ⁇ M 's "1 .
- dissociation constant is meant a “kinetic dissociation constant” or a “calculated dissociation constant.”
- a “kinetic dissociation constant” or “K D” means a ratio of kinetic off-rate (k off ) to kinetic on-rate (Ic 0n ), such as a K D value determined as an absolute value using standard methods (e.g., standard spectroscopic, stopped-flow, or flash-photolysis methods) including methods known to the skilled artisan and/or described herein.
- Calculated dissociation constant or “calculated K D” refers to an approximation of the kinetic dissociation constant based on a measured k off .
- the value for the Ic 0n for NO for an H-NOX protein is assumed to be 710 ⁇ M "1 s '1 , which is a reported Ic 0n for ⁇ l(l-385) that was measured at 4 0 C and does not increase significantly at 37 0 C (Zhao, et. ai, (1999). "A Molecular Basis for Nitric Oxide Sensing by Soluble Guanylate Cyclase," PNAS. 96:14753-14758, which is hereby incorporated by reference in its entirety, particularly with respect to the calculation of NO Ic 0n for H-NOX proteins).
- a value for the Ic 0n is derived via the correlation between kinetic K D and k of r as described herein.
- the kinetic or calculated K D for NO binding by an H-NOX protein is within about 0.01 to about 100-fold of that of hemoglobin under the same conditions (such as at 20 0 C), such as between about 0.1 to about 10-fold or between about 0.5 to about 2-fold of that of hemoglobin under the same conditions (such as at 20 0 C).
- the NO dissociation constant of the H-NOX protein is between about 0.1 to about 20 pM at 37 0 C, such as about 0.5 to about 15, about 0.5 to about 12, about 0.7 to about 4, or about 0.7 to about 3 at 37 0 C.
- the NO dissociation constant of the H-NOX protein is at least about 0.1 pM at 37 0 C, such as at least about any of 0.5, 1, 3, 5, 10, 12, 50, 100, 400, 500, 1000, 2000, 3000, or 4000 pM at 37 0 C. In some embodiments, the NO dissociation constant of the H-NOX protein is less than about 5000 pM at 37 0 C, such as less than about any of 4000 pM, 3000 pM, 2000 pM, 1000 pM, 500 pM, 400 pM, 100 pM, 50 pM, 12 pM, 10 pM, 5 pM, 3 pM, or 1 pM at 37 0 C.
- the kinetic or calculated KQ for O 2 binding by an H-NOX protein is within about 0.01 to about 100-fold of that of hemoglobin under the same conditions (such as at 20 0 C), such as between about 0.1 to about 10-fold or between about 0.5 to about 2- fold of that of hemoglobin under the same conditions (such as at 20 0 C).
- the O 2 dissociation constant of the H-NOX protein it at least about 1 ⁇ M at 37 °C, such as at least about any of 5 ⁇ M, 10 ⁇ M, 20 ⁇ M, 30 ⁇ M, 40 ⁇ M, 50 ⁇ M, 60 ⁇ M, 70 ⁇ M, 80 ⁇ M, 90 ⁇ M, or 100 ⁇ M at 37 0 C.
- NO affinity is a qualitative term that refers to the strength of NO binding to a protein (such as binding to a heme group or to an oxygen bound to a heme group associated with a protein).
- NO stability refers to the stability or resistance of a protein to oxidation by NO in the presence of oxygen. For example, the ability of the protein to not be oxidized when bound to NO in the presence of oxygen is indicative of the protein's NO stability.
- NO reactivity refers to the rate at which iron in the heme of a heme-binding protein is oxidized by NO in the presence of oxygen at a concentration of 2 ⁇ M protein. A lower numerical value for NO reactivity in units of s "1 indicates a lower NO reactivity.
- the NO reactivity of an H-NOX protein is less than about 700 s "1 at 20 "C, such as less than about 600 s "1 , 500 s “1 , 400 s ⁇ 300 s ', 200 s '1 , 100 s “1 , 75 s ', 50 25 10 s “1 , 50 s “1 , 3 s “1 , 2 s '1 , 1.8 s “1 , 1.5 s “1 , 1.2 s 1 , 1.0 s '1 , 0.8 s "1 , 0.7 or 0.6 s "1 at 20 0 C.
- the NO reactivity of an H-NOX protein is between about 0.1 to about 600 s "1 at 20 °C, such as between about 0.5 to about 400 s "1 , about 0.5 to about 100 about 0.5 to about 50 s "1 , about 0.5 to about 10 s “1 , about I to about 5 s “1 , or about 0.5 to about 2.1 s “1 at 20 “C.
- the reactivity of an H-NOX protein is at least about 10, 100, 1,000, or 10,000 fold lower than that of hemoglobin under the same conditions, such as at 20 "C.
- an "autoxidation rate” refers to the rate at which iron in the heme of a heme-binding protein is autoxidized.
- a lower numerical autoxidation rate in units of s ⁇ ' indicates a lower autoxidation rate.
- the rate of heme autoxidation of an H-NOX protein is less than about 1.0 h “ 'at 37 0 C, such as less than about any of 0.9 h “1 , 0.8 h “1 , 0.7 h “ ⁇ 0.6 h ' ⁇ 0.5 h “ ⁇ 0.4 h '1 , 0.3 h “1 , 0.2 h “1 , 0.1 h “1 , or 0.05 h "1 at 37 0 C.
- the rate of heme autoxidation of an H-NOX protein is between about 0.006 to about 5.0 h ⁇ ' at 37 °C, such as about 0.006 to about 1.0 h ⁇ l , 0.006 to about 0.9 h "1 , or about 0.06 to about 0.5 h "1 at 37 0 C.
- a mutant H-NOX protein has (a) an NO or O 2 dissociation constant, association rate (Ic 0n for NO or O 2 ), or dissociation rate (k off for NO or O 2 ) within 2 orders of magnitude of that of hemoglobin, (b) has an NO affinity weaker (e.g., at least about 10-fold, 100- fold, or 1000-fold weaker) than that of sGC ⁇ l (c) an NO reactivity with bound O 2 at least 1000- fold less than hemoglobin, (d) an in vivo plasma retention time at least 2, 10, 100, or 1000-fold higher than that of hemoglobin, or (e) any combination of two or more of the foregoing.
- NO affinity weaker e.g., at least about 10-fold, 100- fold, or 1000-fold weaker
- Exemplary suitable NO carriers provide dissociation constants within two orders of magnitude of that of hemoglobin, i.e. between about 0.01 and 100-fold, such as between about 0.1 and 10-fold, or between about 0.5 and 2-fold of that of hemoglobin.
- a variety of established techniques may be used to quantify dissociation constants, such as the techniques described herein (Boon, E. M. et al. (2005). "Molecular Basis For NO Selectivity in Soluble Guanylate Cyclase," Nature Chem. Biol. 1 :53-59; Boon, E. M. et al. (October 2005). "Ligand Discrimination in Soluble Guanylate Cyclase and the H-NOX Family of Heme Sensor Proteins," Curr.
- Exemplary NO carriers provide low or minimized NO reactivity of the H-NOX protein with bound O 2 , such as an NO reactivity lower than that of hemoglobin. In some embodiments, the NO reactivity is much lower, such as at least about 10, 100, 1,000, or 10,000-fold lower than that of hemoglobin.
- a variety of established techniques may be used to quantify NO reactivity (Boon, E. M. et al. (2005). "Molecular Basis For NO Selectivity in Soluble Guanylate Cyclase," Nature Chem. Biol. 1 :53-59; Boon, E. M. et al. (October 2005). "Ligand Discrimination in Soluble Guanylate Cyclase and the H-NOX Family of Heme Sensor Proteins," Curr.
- T. tengcongensis H-NOX Y 140H has an NO reactivity similar to that of wild-type T. tengcongensis H-NOX.
- Exemplary mutants for NO delivery have an NO affinity weaker, preferably at least
- sGC ⁇ l 10-fold, 100-fold, or 1000-fold weaker than that of sGC ⁇ l .
- a range of engineered H-NOX proteins with varying affinities are empirically tested for efficacy in particular disease states, with a range in some embodiments of NO affinities of 0.1 to 1000 nM.
- suitable NO carriers provide high or maximized stability, particularly in vivo stability.
- stability metrics may be used, such as oxidative stability (e.g., stability to autoxidation or oxidation by NO), temperature stability, and in vivo stability.
- oxidative stability e.g., stability to autoxidation or oxidation by NO
- temperature stability e.g., temperature stability
- in vivo stability e.g., temperature stability
- a variety of established techniques may be used to quantify stability, such as the techniques described herein (Boon, E. M. et al. (2005). "Molecular Basis For NO Selectivity in Soluble Guanylate Cyclase," Nature Chem. Biol. 1 :53-59; Boon, E. M. et al. (October 2005). "Ligand Discrimination in Soluble Guanylate Cyclase and the H-NOX Family of Heme Sensor Proteins,” Curr. Opin.
- H-NOX Domains From sGC to Bacterial NO Sensors
- J. Inorg. Biochem. 99(4): 892-902 exemplary metrics of stability include retention time, rate of clearance, and half-life.
- H-NOX proteins from thermophilic organisms are expected to be stable at high temperatures.
- the plasma retention times are at least about 2-, 10-, 100-, or 1000-fold greater than that of hemoglobin (e.g. Bobofchak, K. M. et al. (March 2003).
- hemoglobin-based carriers are limited by the rapid clearance of cell-free hemoglobin from plasma due the presence of receptors for hemoglobin that remove cell-free hemoglobin from plasma. Since there are no receptors for H- NOX proteins in plasma, wild-type and mutant H-NOX proteins are expected to have a longer plasma retention time than that of hemoglobin. If desired, the plasma retention time can be increased by PEGylating or crosslinking an H-NOX protein or fusing an H-NOX protein with another protein using standard methods (such as those described herein and those known to the skilled artisan).
- the NO dissociation constant of the H-NOX protein is within 2 orders of magnitude of that of hemoglobin, and the NO reactivity of the H-NOX protein is at least 10-fold lower than that of hemoglobin.
- the k of r, kj, or k 2 for NO of the H-NOX protein is between about 1 x 10 "4 S 1 to about 10 s "1 at 37 0 C, and the O 2 dissociation constant of the H-NOX protein is at least about 1 ⁇ M at 37 0 C.
- the k of r, ki, or k 2 for NO of the H-NOX protein is between about 1 x 10 "4 S 1 to about 10 s "1 at 37 °C, and the O 2 dissociation constant of the H-NOX protein is at least about 1 ⁇ M at 37 °C.
- the k o ff, ki, or k 2 for NO of the H-NOX protein is between about 1 x 10 "4 S 1 to about 10 s "1 at 37 0 C, and the NO reactivity of the H-NOX protein is less than about 700 s "1 at 20 °C (e.g., less than about 600 s "1 , 500 s "1 , 100 s 1 , 20 or 1.8 s 'at 20 “C).
- the O 2 dissociation constant of the H-NOX protein is at least about 1 ⁇ M at 37 "C, and the NO reactivity of the H-NOX protein is less than about 700 s "1 at 20 “C (e.g., less than about 600 s 1 , 500 s "1 , 100 s "1 , 20 or 1.8 s " 'at 20 “C).
- the k off , ki, or k 2 for NO of the H-NOX protein is between about 1 x 10 '4 S “1 to about 10 s "1 at 37 “C, and the rate of heme autoxidation of the H-NOX protein is less than about 1 h " 'at 37 “C.
- the O 2 dissociation constant of the H-NOX protein is at least about 1 ⁇ M at 37 "C, and the rate of heme autoxidation of the H-NOX protein is less than about 1 h " 'at 37 “C. In some embodiments, the rate of heme autoxidation of the H-NOX protein is less than about 1 h ' 'at 37 ⁇ C, and the NO reactivity of the H-NOX protein is less than about 700 s 1 at 20 0 C (e.g., less than about 600 s ⁇ 100 s "1 , 20 or 1.8 s 'at 20 “C). In some embodiments, the viscosity of the H-NOX protein solution is between 1 and 4 centipoise (cP). In some embodiments, the colloid oncotic pressure of the H-NOX protein solution is between 20 and 50 mm Hg.
- Table 3 lists exemplary sizes, oxygen affinities, autoxidation stabilities, NO reactivity rates, and modifications for wild-type and mutant H-NOX proteins.
- vehicle size refers to the molecular weight of a modified (e.g., PEGylated) or unmodified H-NOX protein.
- Tables 4-14 Exemplary data for particular mutants are reported in Tables 4-14.
- ⁇ l and B2 refer to proteins derived from rat H-NOX proteins. Since the amino acid sequences of mammalian ⁇ l H-NOX domains differ by at most two amino acids, similar results are expected for the corresponding mutations in other mammalian ⁇ l H-NOX proteins, such as human ⁇ l .
- Table 4 demonstrates that the dissociation constant for NO binding can be significantly changed by mutating one or more residues in H-NOX proteins.
- the ability of allosteric regulators to dramatically affect the dissociation constant and dissociation rate of NO for sGC supports the ability of mutations that alter the structure of sGC or other H-NOX proteins to alter the dissociation constant and dissociation rate of NO.
- the dissociation constant for NO binding can be further altered by combining any of the mutations listed in Table 4 or by introducing one or more additional mutations into an H-NOX protein, as described herein. Table 4. K D values for NO binding to H-NOX and other hemoproteins
- d k values bracket broadest ranges of fits from NO dissociation from 5-coordinate and 6-coordinate, each averaged from 2 — 4 dissociation experiments using saturating CO and 30 mM Na 2 S 2 O 4 as the NO trap and containing 0.88 - 2.2 ⁇ M protein. Data were best fit by a double exponential equation:
- the k o ff for these exemplary mutant H-NOX proteins range from 0.00013 to 0.011 s "1 at 37 0 C.
- NO dissociation rates from hemoproteins are derived using chemical traps as indicated in each cited reference.
- NO dissociation rates from organic nitrates and NONOates are measured using a NO electrode and confirmed using an oxyhemoglobin trap.
- values are adjusted to 37 0 C using the fact that rates double approximately for every 10 0 C.
- the k ⁇ , f r for NO binding can be further altered by combining any of the single or double mutations listed in Table 5 or by introducing one or more additional mutations into an H-NOX protein, as described herein.
- Hemoprotein or Nitrovasodilator kp ff (s ⁇ ) sGC 0.001 to 0.66 a ⁇ 1 ( 1 -385) 0.0023 to 0.0087 a ⁇ 2 (1-217) 0.0069 to 0.011 a ⁇ l (1-194) 0.0009 to 0.0041 a
- Table 7 illustrates the alteration of the O 2 association rate (k on ), O 2 dissociation rate
- any of the single or double mutations listed in Table 7 are combined with another mutation (such as another mutation in Table 7 or any other mutation described herein) to further alter the O 2 association rate, O 2 dissociation rate, O 2 dissociation constant, autoxidation rate, or combinations of two or more of the foregoing.
- Table 8 illustrates that the O 2 association rate, O 2 dissociation rate, O 2 , autoxidation rate, NO reactivity, and stability of Fe 11 O 2 complexes in H-NOX proteins may be altered by the introduction of one or more mutations.
- any of the single or double mutations listed in Table 8 are combined with another mutation (such as another mutation in Table 8 or any other mutation described herein) to further alter the O 2 association rate, O 2 dissociation rate, O 2 , autoxidation rate, NO reactivity, or stability Of Fe 11 O 2 complexes in an H-NOX protein.
- introduction of one or more additional mutations, such as those described herein, may be used to further alter these values. Table 8.
- Table 9 demonstrates that the dissociation constant for O 2 binding can be significantly changed by mutating one or more residues in H-NOX proteins.
- the kinetic K D values for these exemplary H-NOX proteins range from 21.20 nM to 1000000.00 nM at 20 0 C.
- the dissociation constant for O 2 binding can be further altered by combining any of the single or double mutations listed in Table 9 or by introducing one or more additional mutations into an H- NOX protein, as described herein.
- Table 10 demonstrates that the dissociation rates for O 2 binding can be significantly changed by mutating one or more residues in H-NOX proteins.
- the dissociation rates for these exemplary H-NOX proteins range from 0.21 s "1 to 23.4 s "1 at 20 0 C. If desired, the dissociation rate for O 2 binding can be further altered by combining any of the single or double mutations listed in Table 10 or by introducing one or more additional mutations into an H-NOX protein, as described herein.
- Table 11 demonstrates that the association rates for O 2 binding can be significantly changed by mutating one or more residues in H-NOX proteins.
- the association rates for these exemplary H-NOX proteins range from 60 ⁇ M ' V to 0.14 ⁇ M 's "1 at 20 0 C. If desired, the association rate for O 2 binding can be further altered by combining any of the single or double mutations listed in Table 11 or by introducing one or more additional mutations into an H-NOX protein, as described herein.
- Table 12 illustrates the effect of exemplary H-NOX mutations on NO
- Each number listed in Table 12 for the Fe-unligated form is for a single peak (which is listed in between the ⁇ and ⁇ columns). When NO or O2 binds, this single peak splits into two peaks, ⁇ and ⁇ (which are listed below the ⁇ and ⁇ columns, respectively). If desired, NO or O 2 -binding can be further altered by combining any of the single or double mutations listed in Table 12 or by introducing one or more additional mutations into an H-NOX protein, as described herein.
- Table 13 contains U V-visible peak positions for some Fe (II), Fe (III), Fe(I I)-NO, and Fe(II)-O 2 complexes.
- a hemoglobin or H-NOX protein When a hemoglobin or H-NOX protein is anaerobic, it has a Soret peak at —431 nm, and it is in an unligated state. If the H-NOX protein does not bind NO, then the Soret peak will not change when NO is added. If the H-NOX protein binds NO and forms a 6-coordinate ferrous-nitrosyl complex, then its Soret peak will shift to between 420 nm and 424 nm when NO is added.
- Soret peak will shift to -399 nm. If the H-NOX protein does not bind O 2 , then the Soret peak will not change when O 2 is added. If the H-NOX protein does bind O 2 , then its Soret peak will shift to between 414 nm and 418 nm when O 2 is added, which is the same shift that occurs in hemoglobin, indicative OfO 2 bound to the heme. Soret peaks for oxidized H-NOX (Fe(III)) may be relevant to the state of the H-NOX protein after storage or use.
- NO or O 2 -binding can be further altered by combining any of the single or double mutations listed in Table 13 or by introducing one or more additional mutations into an H-NOX protein, as described herein.
- Table 13 UV-visible peak positions for some Fe (II), Fe (III), Fe(II)-NO, and Fe(II)-O 2 complexes.
- Table 14 contains autoxidation rates for exemplary T. tengcongensis H-NOX proteins. If desired, the autoxidation rate can be further altered by combining any of the mutations listed in Table 14 or by introducing one or more additional mutations into an H-NOX protein, as described herein.
- the 2 nm and 3 nm values mean in Table 14 refer to a shift in the UV-Vis Soret peak by 2 to 3 nm over the time period of the observation; this extremely small change may be due to autoxidation.
- a Stable denotes lack of heme oxidation after at least 24 hours.
- RT denotes room temperature.
- nucleic acids encoding any of the mutant H-NOX proteins described herein.
- a "nucleic acid” refers to two or more deoxyribonucleotides and/or ribonucleotides in either single or double-stranded form, and unless otherwise limited, encompasses known analogs of naturally-occurring nucleotides that hybridize to nucleic acids in a manner similar to nucleotides occurring in nature.
- the nucleic acid is a recombinant nucleic acid.
- recombinant nucleic acid means a nucleic acid of interest that is free of one or more nucleic acids ⁇ e.g., genes) which, in the genome occurring in nature of the organism from which the nucleic acid of interest is derived, flank the nucleic acid of interest.
- an H-NOX nucleic acid is operably linked to another nucleic acid encoding all or a portion of another protein such that the recombinant nucleic acid encodes a fusion protein that includes an H-NOX protein (e.g., an H-NOX domain with or without another domain from an H-NOX protein) and all or part of another protein, such as human serum albumin.
- the term therefore includes, for example, a recombinant DNA which is incorporated into a vector, into an autonomously replicating plasmid or virus, or into the genomic DNA of a prokaryote or eukaryote, or which exists as a separate molecule (e.g., a cDNA, a genomic DNA fragment, or a cDNA fragment produced by PCR or restriction endonuclease digestion) independent of other sequences.
- a recombinant DNA which is incorporated into a vector, into an autonomously replicating plasmid or virus, or into the genomic DNA of a prokaryote or eukaryote, or which exists as a separate molecule (e.g., a cDNA, a genomic DNA fragment, or a cDNA fragment produced by PCR or restriction endonuclease digestion) independent of other sequences.
- the invention also features vectors with one more nucleic acids encoding any of the mutant H-NOX proteins that are described herein.
- a "vector” means a construct that is capable of delivering, and optionally expressing one or more nucleic acids of interest in a host cell. Examples of vectors include, but are not limited to, plasmids, viral vectors, DNA or RNA expression vectors, cosmids, and phage vectors.
- the vector contains a nucleic acid under the control of an expression control sequence.
- An "expression control sequence” means a nucleic acid sequence that directs transcription of a nucleic acid of interest.
- An expression control sequence can be a promoter, such as a constitutive or an inducible promoter, or an enhancer. The expression control sequence is operably linked to the nucleic acid segment to be transcribed.
- the nucleic acid includes a segment of or the entire nucleic acid sequence of any of the nucleic acids shown in FIGS. 2-4D or 8A-8DD.
- the nucleic acid includes at least about 50, 100, 150, 200, 300, 400, 500, 600, 700, 800, or more contiguous nucleotides from a H-NOX nucleic acid and contains one or more mutations (e.g., 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 mutations) compared to the H-NOX nucleic acid from which it was derived.
- a mutant H-NOX nucleic acid contains less than about 20, 15, 12, 10, 9, 8, 7, 6, 5, 4, 3, or 2 mutations compared to the H-NOX nucleic acid from which it was derived.
- the invention also features degenerate variants of any nucleic acid encoding a mutant H-NOX protein.
- the invention also includes a cell or population of cells containing at least one nucleic acid encoding a mutant H-NOX protein described herein.
- exemplary cells include insect, plant, yeast, bacterial, and mammalian cells. These cells are useful for the production of mutant H- NOX proteins using standard methods, such as those described herein.
- Any wild-type or mutant H-NOX protein described herein may be used for the formulation of pharmaceutical or non-pharmaceutical compositions. As discussed further below, these formulations are useful in a variety of therapeutic and industrial applications.
- the pharmaceutical composition includes one or more wild- type or mutant H-NOX proteins (such as any of the H-NOX wild-type or mutant proteins described herein) and a pharmaceutically acceptable carrier.
- the H-NOX protein is an isolated or purified protein.
- pharmaceutically acceptable carrier is meant any material which, when combined with an active ingredient, allows the ingredient to retain biological activity and does not provoke an unacceptable immune response (e.g., a severe allergy or anaphylactic shock) based on the knowledge of a skilled practitioner. Examples include, but are not limited to, any of the standard pharmaceutical carriers such as phosphate buffered saline solutions, water, emulsions such as oil/water emulsion, and various types of wetting agents.
- Exemplary diluents for aerosol or parenteral administration are phosphate buffered saline or normal (0.9%) saline.
- Compositions comprising such carriers are formulated by well known conventional methods (see, for example, Remington's Pharmaceutical Sciences, 18th edition, A. Gennaro, ed., Mack Publishing Co., Easton, PA, 1990; and Remington, The Science and Practice of Pharmacy 20th Ed. Mack Publishing, 2000, which are each hereby incorporated by reference in their entireties, particularly with respect to formulations).
- compositions of this invention can be formulated for any appropriate manner of administration, including, for example, intravenous, intra-arterial, intravesicular, inhalation, intraperitoneal, intrapulmonary, intramuscular, subcutaneous, intra-tracheal, transmucosal, intraocular, intrathecal, or transdermal administration.
- the carrier may include, e.g., water, saline, alcohol, a fat, a wax, or a buffer.
- any of the above carriers or a solid carrier such as mannitol, lactose, starch, magnesium stearate, sodium saccharine, talcum, cellulose, glucose, sucrose, or magnesium carbonate, may be employed.
- Biodegradable microspheres e.g., polylactate polyglycolate
- the pharmaceutical or non-pharmaceutical compositions include a buffer (e.g., neutral buffered saline, phosphate buffered saline, etc), a carbohydrate (e.g., glucose, mannose, sucrose, dextran, etc), an antioxidant, a chelating agent (e.g., EDTA, glutathione, etc.), a preservative, another compound useful for binding and/or transporting NO, an inactive ingredient (e.g., a stabilizer, filler, etc), or combinations of two or more of the foregoing.
- the composition is formulated as a lyophilizate.
- H-NOX proteins may also be encapsulated within liposomes or nanoparticles using well known technology.
- Other exemplary formulations that can be used for H-NOX proteins are described by, e.g., U.S. Pat. Nos. 6,974,795, and 6,432,918, which are each hereby incorporated by reference in their entireties, particularly with respect to formulations of proteins.
- compositions described herein may be administered as part of a sustained release formulation (e.g., a formulation such as a capsule or sponge that produces a slow release of compound following administration).
- a sustained release formulation e.g., a formulation such as a capsule or sponge that produces a slow release of compound following administration.
- Such formulations may generally be prepared using well known technology and administered by, for example, oral, rectal or subcutaneous implantation, or by implantation at the desired target site.
- Sustained-release formulations may contain an H-NOX protein dispersed in a carrier matrix and/or contained within a reservoir surrounded by a rate controlling membrane. Carriers for use within such formulations are biocompatible, and may also be biodegradable.
- the formulation provides a relatively constant level of H- NOX protein release. The amount of H-NOX protein contained within a sustained release formulation depends upon the site of implantation, the rate and expected duration of release, and the nature of the condition to be treated or prevented.
- the pharmaceutical composition contains an effective amount of a wild-type or mutant H-NOX protein.
- the term "effective amount" intends such amount of one or more proteins described herein which in combination with its parameters of efficacy and toxicity should be effective in a given therapeutic form based on the knowledge of the practicing specialist.
- an effective amount can be in one or more doses.
- an effective dosage of a pharmaceutical composition may or may not be achieved in conjunction with another drug, compound, or pharmaceutical composition.
- an effective amount can be considered in the context of administering one or more therapeutic agents, and a single agent can be considered to be given in an effective amount if, in conjunction with one or more other agents, a desirable or beneficial result can be or is achieved.
- An exemplary dose of hemoglobin as a blood substitute is from about 10 mg to about
- an effective amount of an H-NOX protein for administration to a human is between a few grams to over about 350 grams.
- Other exemplary doses of an H-NOX protein include about any of 4.4., 5, 10, or 13 G/DL (where G/DL is the concentration of the H-NOX protein solution prior to infusion into the circulation) at an appropriate infusion rate, such as about 0.5 ml/min (see, for example, Winslow, R. Chapter 12 In Blood Substitutes).
- a dose of less than 10 mg of H-NOX protein is used for temporary vasodilation.
- the unit content of active ingredients contained in an individual dose of each dosage form need not in itself constitute an effective amount since the necessary effective amount could be reached by the combined effect of a plurality of administrations.
- the selection of the amount of an H-NOX protein to include in a pharmaceutical composition depends upon the dosage form utilized, the condition being treated, and the particular purpose to be achieved according to the determination of the ordinarily skilled artisan in the field.
- compositions include genetically engineered, recombinant H-NOX proteins, which may be isolated or purified, comprising one or more mutations that collectively impart altered NO or O 2 ligand-binding relative to the corresponding wild-type H-NOX protein, and operative as a physiologically compatible mammalian blood gas NO carrier.
- mutant H-NOX proteins as described herein.
- the invention also provides NO carriers comprising or consisting essentially of one or more wild-type or mutant H-NOX proteins. Suitable buffers and other ingredients for formulating proteins (such as proteins delivered to the blood or gastrointestinal system) are known in the art.
- human H-NOX proteins either wild-type human proteins or human proteins into which one or more mutations have been introduced
- other non-antigenic H-NOX proteins ⁇ e.g., mammalian H-NOX proteins
- amino acids in an H- NOX protein can be mutated to the corresponding amino acids in a human H-NOX.
- amino acids on the surface of the tertiary structure of a non-human H-NOX protein can be mutated to the corresponding amino acid in a human H-NOX protein.
- H-NOX proteins e.g. , isolated or purified H-NOX proteins
- pharmaceutical compositions described herein may be used in therapeutic applications.
- H-NOX proteins can be selected for such applications based on the desired [0124] NO dissociation constant, O 2 dissociation constant, NO k off , O2 Kn, NO reactivity,
- H-NOX proteins can be used to treat any condition for which delivery of NO is beneficial.
- exemplary target indications include diseases of functional NO deficiency, such as where a vasodilator or an NO carrier is indicated, including conditions exacerbated by chronic hypertension, such as heart failure, renal failure, and stroke.
- the treated condition is a cardiovascular condition (e.g., myocardial infarction or heart surgery), hypertension, a vasoconstrictive condition (e.g., stroke), erectile dysfunction, constipation, or bowel obstruction.
- H-NOX proteins can be used to deliver NO to treat a sphincter control deficit, thereby relaxing the smooth muscle.
- H-NOX proteins that function in the digestive system can relax the smooth muscle of the ileum as the H-NOX proteins pass through the digestive system.
- the methods and compositions are applicable to both acute (providing rapid NO to tissues or a specific site, e.g., acute myocardial infarction or stroke) and chronic situations (e.g., chronic hypertension or post- acute recovery from cardiac infarction or stroke).
- the invention features a method of delivering NO to an individual (e.g., a mammal, such as a primate (e.g., a human, a monkey, a gorilla, an ape, a lemur, etc), a bovine, an equine, a porcine, a canine, or a feline) by administering to an individual in need thereof a wild-type or mutant H-NOX protein in an amount sufficient to deliver NO to the individual.
- a mammal such as a primate (e.g., a human, a monkey, a gorilla, an ape, a lemur, etc), a bovine, an equine, a porcine, a canine, or a feline) by administering to an individual in need thereof a wild-type or mutant H-NOX protein in an amount sufficient to deliver NO to the individual.
- a primate e.g., a human, a monkey, a gorilla, an ape,
- the invention provides methods of carrying or delivering blood gas to an individual such as a mammal, comprising the step of delivering to the blood of the individual (e.g., a mammal) one or more of H-NOX compositions.
- Methods for delivering proteins to the blood, digestive system, or tissues are known in the art.
- the H-NOX protein is an apoprotein that is capable of binding heme or is a holoprotein with heme bound.
- the H-NOX protein may or may not have heme bound prior to the administration of the H-NOX protein to the individual.
- NO is bound to the H-NOX protein before it is delivered to the individual.
- NO is not bound to the H-NOX protein prior to the administration of the protein to the individual, and the H-NOX protein transports NO from one location in the individual to another location in the individual.
- H-NOX proteins bind NO in the blood stream and only release it where NO concentrations are very low (such as sites of vasoconstriction). This targeted delivery of NO may produce fewer side-effects than conventional vasodilators that release NO independent of local NO concentration and thus function systemically, with side effects such as headaches and peripheral tingling.
- an individual intends a mammal, including but not limited to, a primate (e.g., a human, monkey, gorilla, ape, lemur, etc.), a bovine, an equine, a porcine, a canine, and a feline.
- a primate e.g., a human, monkey, gorilla, ape, lemur, etc.
- bovine e.g., a bovine
- an equine e.g., a porcine
- canine e.g., a canine
- feline e.g., a bovine, an equine, a porcine, a canine, and a feline.
- the individual may have been diagnosed with, is suspected of having, or is at risk of developing an indication, such as a cardiovascular condition (e.g., myocardial infarction or heart surgery), hypertension, a condition exacerbated by hypertension ⁇ e.g., heart failure, renal failure, or stroke), a vasoconstrictive condition ⁇ e.g., stroke), a functional NO deficiency, erectile dysfunction, constipation, or bowel obstruction.
- the individual may exhibit one or more symptoms associated with the indication.
- the individual can be genetically or otherwise predisposed to developing such a condition.
- a condition or disease ⁇ e.g., as a cardiovascular condition such as myocardial infarction or heart surgery, hypertension, a condition exacerbated by hypertension such as heart failure, renal failure, or stroke, a vasoconstrictive condition such as stroke, a functional NO deficiency, erectile dysfunction, constipation, or bowel obstruction) or are "at risk" for the condition or disease.
- a condition or disease such as myocardial infarction or heart surgery, hypertension, a condition exacerbated by hypertension such as heart failure, renal failure, or stroke, a vasoconstrictive condition such as stroke, a functional NO deficiency, erectile dysfunction, constipation, or bowel obstruction
- an "at risk” individual is an individual who is at risk of development of a condition, such as a cardiovascular condition ⁇ e.g., myocardial infarction or heart surgery), hypertension, a condition exacerbated by hypertension ⁇ e.g., heart failure, renal failure, or stroke), a vasoconstrictive condition ⁇ e.g., stroke), a functional NO deficiency, erectile dysfunction, constipation, or bowel obstruction.
- An individual “at risk” may or may not have a detectable disease or condition, and may or may not have displayed detectable disease prior to the treatment methods described herein.
- At risk denotes that an individual has one or more so-called risk factors, which are measurable parameters that correlate with development of a disease or condition and are known in the art. An individual having one or more of these risk factors has a higher probability of developing the disease or condition than an individual without these risk factor(s).
- risk factors include, but are not limited to, age, sex, race, diet, history of previous disease, presence of precursor disease, genetic ⁇ i.e., hereditary) considerations, and environmental exposure.
- beneficial or desired results include, but are not limited to, alleviation of symptoms associated with a condition (such as, but not limited to, a cardiovascular condition such as myocardial infarction or heart surgery, hypertension, a condition exacerbated by hypertension such as heart failure, renal failure, or stroke, a vasoconstrictive condition such as stroke, a functional NO deficiency, erectile dysfunction, constipation, or bowel obstruction) diminishment of the extent of the symptoms associated with a condition, or prevention of a worsening of the symptoms associated with a condition.
- a condition such as, but not limited to, a cardiovascular condition such as myocardial infarction or heart surgery, hypertension, a condition exacerbated by hypertension such as heart failure, renal failure, or stroke, a vasoconstrictive condition such as stroke, a functional NO deficiency, erectile dysfunction, constipation, or bowel obstruction
- treatment with a one or more proteins disclosed herein is accompanied by no or fewer side effects than are associated
- a disease or condition means to defer, hinder, slow, retard, stabilize and/or postpone development of the disease or condition, such as a cardiovascular condition (e.g., myocardial infarction or heart surgery), hypertension, a condition exacerbated by hypertension (e.g., heart failure, renal failure, or stroke), a vasoconstrictive condition (e.g., stroke), a functional NO deficiency, erectile dysfunction, constipation, or bowel obstruction.
- This delay can be of varying lengths of time, depending on the history of the disease and/or individual being treated.
- a sufficient or significant delay can, in effect, encompass prevention, in that the individual does not develop the disease or condition.
- the method may reduce the probability of disease development in a given time frame and/or reduce the extent of the disease in a given time frame, when compared to not using the method. In some embodiments, such comparisons are based on clinical studies using a statistically significant number of subjects.
- Disease development can be detectable using standard clinical techniques. Development may also refer to disease progression that can be initially undetectable and includes occurrence, recurrence, and onset.
- the kofr for NO is more important than the K D because NO is already bound to the protein (making the Ic 0n less important) and NO needs to be released at or near a particular site in the body (at a rate influenced by the k off ).
- the H-NOX protein has a relatively high k off , ki, or k 2 for NO (such as at least about any of 0.05 s " ', 0.1 s "1 , or 1.0 s "1 ) so that vasodilation occurs rapidly.
- the H-NOX protein has a relatively low k of r, ki, or k 2 for NO (such as at less than about any of 0.05 s "1 , 0.01 or 0.001 s 1 ) so that NO is not released until the H-NOX protein reaches a site of low NO concentration (e.g., a vasoconstricted site).
- H-NOX proteins can also be used for imaging. In particular, light imaging (e.g., optical coherence tomography; see, for example, Villard, J. W. (2002).
- H-NOX proteins and pharmaceutical compositions of the invention can be administered to an individual by any conventional means such as by oral, topical, intraocular, intrathecal, intrapulmonary, intra- tracheal, or aerosol administration; by transdermal or mucus membrane adsorption; or by injection (e.g., subcutaneous, intravenous, intra-arterial, intravesicular, or intramuscular injection).
- H-NOX proteins may also be included in large volume parenteral solutions for administration to the blood.
- the H-NOX protein is administered to the blood (e.g., administration to a blood vessel such as a vein, artery, or capillary) of the individual.
- a sustained continuous release formulation of the composition is used.
- Administration of an H-NOX protein can occur, e.g., for a period of seconds to hours depending on the purpose of the administration.
- an exemplary time course of administration is as rapid as possible.
- Other exemplary time courses include about 10, 20, 30, 40, 60, 90, or 120 minutes.
- Exemplary infusion rates for H-NOX solutions are from about 30 mL/hour to about 13,260 mL/hour, such as about 100 mL/hour to about 3,000 mL/hour.
- An exemplary total dose of H-NOX protein is about 900 mg/kg administered over 20 minutes at 13,260 mL/hour.
- An exemplary total dose of H-NOX protein for a swine is about 18.9 grams.
- Exemplary dosing frequencies include, but are not limited to, at least 1, 2, 3, 4, 5, 6, or 7 times (i.e., daily) a week.
- an H-NOX protein is administered at least 2, 3, 4, or 6 times a day.
- the H-NOX protein can be administered, e.g., over a period of a few days or weeks.
- the H-NOX protein is administrated for a longer period, such as a few months or years.
- the dosing frequency of the composition may be adjusted over the course of the treatment based on the judgment of the administering physician.
- an effective amount of an H-NOX protein for administration to human is between a few grams to over 350 grams.
- two or more different H-NOX proteins are administered simultaneously, sequentially, or concurrently.
- another compound or therapy useful for the delivery of NO is administered simultaneously, sequentially, or concurrently with the administration of one or more H-NOX proteins.
- H-NOX proteins and composition described herein can also be used for a number of in vitro or industrial applications (see, e.g., U.S. Pat. No. 6,455,676, which is hereby incorporated by reference in its entirety, particularly with respect to in vitro or industrial applications).
- Particular H-NOX proteins can be selected for such applications based on the desired NO dissociation constant, O2 dissociation constant, NO k ⁇ , O 2 k off , NO reactivity, NO stability, autoxidation rate, half-life, or any combination of two or more of the foregoing for the particular application.
- the H-NOX protein is an apoprotein that is capable of binding heme or is a holoprotein with heme bound.
- H-NOX proteins can be used to add NO to solutions for which NO is desirable.
- H-NOX proteins are used to deliver NO delivery to cells.
- the delivery of NO to mitochondria may limit oxidative phosphorylation and enhance metabolism through the lactate pathway.
- the H-NOX protein in Clostridium acetobutylicum which is cultured under anaerobic fermentation as a biofuel generator, may naturally serve this function.
- the H-NOX proteins can be used to remove NO from solutions requiring the removal of NO.
- H-NOX proteins may be used to absorb or remove NO in bioreactors where NO is an inhibitor of cellular proliferation and/or mitochondrial function. Removing NO may improve mitochondrial function, limit apoptosis, increase per-cell productivity, or any combination of two or more of the foregoing.
- kits that include any of the H-NOX proteins described herein and suitable packaging.
- the invention includes a kit with (i) an H-NOX protein (such as a wild-type or mutant H-NOX protein described herein or formulations thereof as described herein) and (ii) instructions for using the kit to deliver NO to an individual.
- the invention features a kit with (i) an H-NOX protein (such as a wild-type or mutant H-NOX protein described herein or formulations thereof as described herein) and (ii) instructions for using the kit for any of the industrial uses described herein (e.g., addition of NO to a solution or removal of NO from a solution).
- Suitable packaging for compositions described herein are known in the art, and include, for example, vials (e.g., sealed vials), vessels, ampules, bottles, jars, flexible packaging (e.g., sealed Mylar or plastic bags), and the like. These articles of manufacture may further be sterilized and/or sealed. Also provided are unit dosage forms comprising the compositions described herein. These unit dosage forms can be stored in a suitable packaging in single or multiple unit dosages and may also be further sterilized and sealed.
- kits of the invention are typically written instructions on a label or package insert (e.g., a paper sheet included in the kit), but machine-readable instructions (e.g., instructions carried on a magnetic or optical storage disk) are also acceptable.
- the instructions relating to the use of H-NOX proteins generally include information as to dosage, dosing schedule, and route of administration for the intended treatment or industrial use.
- the kit may further comprise a description of selecting an individual suitable or treatment.
- kits may be unit doses, bulk packages (e.g., multi-dose packages) or sub- unit doses.
- kits may also be provided that contain sufficient dosages of H-NOX proteins disclosed herein to provide effective treatment for an individual for an extended period, such as about any of a week, 2 weeks, 3 weeks, 4 weeks, 6 weeks, 8 weeks, 3 months, 4 months, 5 months, 6 months, 7 months, 8 months, 9 months, or more.
- Kits may also include multiple unit doses of H-NOX proteins and instructions for use and packaged in quantities sufficient for storage and use in pharmacies, for example, hospital pharmacies and compounding pharmacies.
- the kit includes a dry (e.g., lyophilized) composition that can be reconstituted, resuspended, or rehydrated to form generally a stable aqueous suspension of H-NOX protein.
- the present invention also provides methods for the production of any of the mutant
- the method involves culturing a cell that has a nucleic acid encoding a mutant H-NOX protein under conditions suitable for production of the mutant H-NOX protein.
- the mutant H-NOX is also purified (such as purification of the H-NOX protein from the cells or the culture medium).
- mutant H-NOX proteins described herein can be generated a number of methods that are known in the art. Mutation can occur at either the amino acid level by chemical modification of an amino acid or at the codon level by alteration of the nucleotide sequence that codes for a given amino acid. Substitution of an amino acid at any given position in a protein can be achieved by altering the codon that codes for that amino acid. This can be accomplished by site- directed mutagenesis using, for example: (i) the Amersham technique (Amersham mutagenesis kit, Amersham, Inc., Cleveland, Ohio) based on the methods of Taylor, J. W. et al. (December 20, 1985).
- Mutagenesis can also be accomplished by other commercially available or noncommercial means, such as those that utilize site-directed mutagenesis with mutant oligonucleotides.
- Site-directed mutagenesis can also be accomplished using PCR-based mutagenesis such as that described in Zhengbin et al. (1992). pages 205-207 in PCR Methods and Applications, Cold Spring Harbor Laboratory Press, New York; Jones, D. H. et al. (February 1990). "A Rapid Method For Site-Specific Mutagenesis And Directional Subcloning by Using the Polymerase Chain Reaction to Generate Recombinant Circles," Biotechniques 8(2): 178-183; Jones, D. H. et al. (January 1991).
- a mutant H-NOX nucleic acid can be incorporated into a vector, such as an expression vector, using standard techniques. For example, restriction enzymes can be used to cleave the mutant H-NOX nucleic acid and the vector.
- the compatible ends of the cleaved mutant H-NOX nucleic acid and the cleaved vector can be ligated.
- the resulting vector can be inserted into a cell (e.g., an insect cell, a plant cell, a yeast cell, or a bacterial cell) using standard techniques (e.g., electroporation) for expression of the encoded H-NOX protein.
- heterologous proteins have been expressed in a number of biological expression systems, such as insect cells, plant cells, yeast cells, and bacterial cells.
- any suitable biological protein expression system can be utilized to produce large quantities of recombinant H-NOX protein.
- the H-NOX protein e.g., a mutant or wild- type H-NOX protein
- an "isolated protein” means a protein separated from one or more components with which the protein is naturally associated in nature, including, for example, nucleic acids, lipids, and other proteins.
- An isolated protein also does not occur in a library of proteins, such as a library of 2, 5, 10, 20, 50 or more different proteins.
- An isolated protein can be obtained, for example, by expression of a recombinant nucleic acid encoding the protein or by chemical synthesis of the protein.
- H-NOX proteins can be purified using standard techniques.
- a "purified protein” means a protein (e.g., a mutant or wild-type H-NOX protein) that has been separated from one or more components that are present when the protein is produced.
- the protein is at least about 60%, by weight, free from other components that are present when the protein is produced.
- the protein is at least about 75%, 90%, or 99%, by weight, pure.
- a purified protein can be obtained, for example, by purification (e.g., extraction) from a natural source, a recombinant expression system, or a reaction mixture for chemical synthesis.
- Exemplary methods of purification include immunoprecipitation, column chromatography such as immunoaffinity chromatography, magnetic bead immunoaff ⁇ nity purification, and panning with a plate-bound antibody, as well as other techniques known to the skilled artisan. Purity can be assayed by any appropriate method, e.g., by column chromatography, polyacrylamide gel electrophoresis, or HPLC analysis.
- the purified protein is incorporated into a pharmaceutical composition of the invention or used in a method of the invention.
- the pharmaceutical composition of the invention may have additives, carriers, or other components in addition to the purified protein.
- Wild-type and mutant H-NOX proteins were produced, expressed, and purified using standard methods, essentially as described by Boon, E.M. et al. (2005). "Molecular Basis For NO Selectivity in Soluble Guanylate Cyclase," Nature Chemical Biology 1 :53-59 and Karow, D. S. et al. (August 10, 2004).
- the value for the Jc 0n for NO for an H-NOX protein is assumed to be 710 ⁇ M ' V 1 , and the dissociation rate for NO (k off for an H-NOX protein with a 6-coordinate Fe"-NO complex or k] or k 2 for an H-NOX protein with a 5-coordinate Fe"-NO complex) is determined as described below.
- Data were acquired by scanning periodically on a Cary 3E spectrophotometer equipped with a Neslab RTE- 100 constant temperature bath set to varying temperatures (0-70 0 C) using a quartz cuvette with a size of 100 ⁇ L to 1 mL and a path-length of 1 -cm (Cary 3E, Varian, Inc., Palo Alto, CA).
- the dissociation of NO from the heme was monitored as the formation of the Fe"-CO complex at 423 nm. Difference spectra were calculated by subtracting the first scan from each subsequent scan.
- a single exponential increase in the concentration of heme-CO due to CO binding from the NO trap can be described by equation 1 :
- ⁇ A t is the change in signal amplitude at time t
- ⁇ AT is the total change in signal amplitude
- ki is the observed reaction rate constant.
- k/ and 1c2 values for H-NOX proteins with a 5-coordinate Fe 1 -NO complex the k of r for NO is described by the ki for NO and the k 2 for NO, as described by Winger, J. A. et al, (January 2007) "Dissociation of Nitric Oxide from Soluble Guanylate Cyclase and Heme-Nitric Oxide/Oxygen Binding Domain Constructs" J. Biol. Chem.
- the trapping solution was prepared as follows: a solution of sodium dithionite (Na 2 S 2 O 4 ) in 50 mM HEPES, pH 7.4, 50 mM NaCl was prepared in a Teflon-sealed Reacti-Vial (Pierce) using an anaerobic chamber (Coy Laboratory Products). The solution was removed from the anaerobicchamber and saturated with CO by bubbling the gas through the solution for 10 minutes.
- H-NOX protein-NO complexes were formed by incubation with excess DEA/NO (in 10 mM NaOH) at 25 0 C in 50 tnM HEPES, pH 7.4, 50 mM NaCl for 10 min. Complete conversion to the nitrosyl species was verified by following the shift in the Soret maximum from 431 to 399 nm.
- H-NOX proteins were placed in a septum-sealed anaerobic cuvette (a quartz cuvette with a size of 100 ⁇ L to 1 mL and a path-length of 1-cm) and deoxygenated using an oxygen-scavenged gas train.
- a buffer base line was subtracted from each spectrum, and spectra were corrected for base-line drift by normalization to an isosbestic point at 410 nm. Difference spectra were obtained by subtraction of the time 0 spectrum from all subsequent spectra. Values for the change in absorbance at 423 nm ( ⁇ A 423 ; ⁇ 1(1 -194) and ⁇ l(l- 385)) or 424 nm (AA 424 ; sGC and ⁇ 2(l-217)) were extracted from the difference spectra and plotted versus time to obtain dissociation time courses for each experiment. Dissociation time courses were obtained in duplicate or triplicate, and each experiment was repeated 2—5 times over several days. Generally, because of the relative difficulty in obtaining large amounts of purified sGC, ⁇ A4 2 4 values for full-length sGC, which are proportional to the experimental protein concentrations, were smaller than for the heme domain constructs.
- Equation 2 Kaleidagraph 3.X (Synergy Software Reading, PA). The data from each dissociation experiment were fit to a double exponentials as shown in Equation 2 below to obtain observed rate constants. In particular, equation 2 describes a two-exponential increase in the concentration of heme-CO (due to CO binding from the NO trap):
- ⁇ A t is the change in signal amplitude at time t
- ⁇ Ai and ⁇ A 2 are the contributions of each exponential process to the total change in signal amplitude
- ki and ki are the observed rate constants for each process.
- the k o fr for NO is described by the ki for NO and the k 2 for NO.
- the ki and the k 2 for NO are measured as described above for H-NOX proteins with a 5-coordinate Fe"-NO complex, as described by Winger, J. A. et al, (January 2007) "Dissociation of Nitric Oxide from Soluble
- the value for the Ic 0n for NO for an H-NOX protein is assumed to be 710 which is a reported Ic 0n for ⁇ l(l-385) that was measured at 4 0 C and does not increase significantly at 37 0 C (Zhao, et. al., (1999). "A Molecular Basis for Nitric Oxide Sensing by Soluble Guanylate Cyclase," PNAS. 96: 14753-14758, which is hereby incorporated by reference in its entirety, particularly with respect to the calculation of NO Ic 0n for H-NOX proteins).
- the calculated K D for NO is determined by calculating the ratio of either k off , kj, or k 2 (measured as described above) to Ic 0n (assumed to be 710 ⁇ M ' V).
- the samples were then diluted to 20 ⁇ M heme in 50 mM TEA, 50 mM NaCl, pH 7.5 buffer in a controlled-atmosphere quartz cuvette, with a size of 100 ⁇ L to 1 mL and a path- length of 1-cm.
- CO gas was flowed over the headspace of this cuvette for 10 minutes to form the Fe 1 '- CO complex, the formation of which was verified by UV-visible spectroscopy (Soret maximum 423 nm).
- This sample was then either used to measure CO-rebinding kinetics after flash photolysis while still under 1 atmosphere of CO gas, or it was opened and stirred in air for 30 minutes to fully oxygenate the buffer before flash photolysis to watch O 2 -rebinding events.
- the dissociation of O 2 from the heme was monitored as an increase in the absorbance at 437 nm, a maximum in the Fe" - Fe u -O 2 difference spectrum, or 425 nm, a maximum in the Fe 11 — Fe"-CO difference spectrum.
- the final traces were fit to a single exponential using the software that is part of the instrument. Each experiment was done a minimum of six times, and the resulting rates were averaged.
- the dissociation rates measured are independent of dithionite concentration (100, 50, 25, 10, 5 and 2.5 mM dithionite were tested) and independent of saturating CO as a trap for the reduced species, both with and without 10 mM dithionite present.
- the kinetic K D for O 2 is determined by calculating the ratio of ko ff to Ic 0n using the measurements of k off and Ic 0n described above.
- HNOX Homo sapiens hemoglobin
- Hs Hb Homo sapiens hemoglobin
- Buffer A 50 mM Hepes, pH 7.5, 50 mM NaCl
- Data were acquired on a HITECH Scientific SF-61 stopped-flow spectrophotometer equipped with a Neslab RTE- 100 constant- temperature bath set to 20 0 C (TGK Scientific LTD., Bradford On Avon, United Kingdom).
- the protein was rapidly mixed with NO in a 1 : 1 ratio with an integration time of 0.00125 sec.
- the wavelengths of maximum change were fit to a single exponential using the software that is part of the spectrometer, essentially measuring the rate-limiting step of oxidation by NO.
- the end products of the reaction were ferric-NO for the HNOX proteins and ferric-aquo for Hs Hb.
- the p50 value for mutant or wild-type H-NOX proteins can be measured as described by Guarnone, R. et al. (September/October 1995). "Performance Characteristics of Hemox-Analyzer For Assessment of The Hemoglobin Dissociation Curve," Haemalologica 80(5):426-430, which is hereby incorporated by reference in its entirety, particularly with respect to the measurement of p50 values.
- the p50 value is determined using a HemOx analyzer. The measurement chamber starts at 0% oxygen and slowly is raised, incrementally, towards 100% oxygen. An oxygen probe in the chamber measures the oxygen saturation %.
- a second probe measures two wavelengths of absorption, tuned to the alpha and beta peaks of the hemoprotein's ⁇ e.g., a protein such as H-NOX complexed with heme) UV-Vis spectra. These absorption peaks increase linearly as hemoprotein binds oxygen. The percent change from unbound to 100% bound is then plotted against the % oxygen values to generate a curve. The p50 is the point on the curve where 50% of the hemoprotein is bound to oxygen.
- the Hemox-Analyzer determines the oxyhemoprotein dissociation curve (ODC) by exposing 50 ⁇ L of blood or hemoprotein to an increasing partial pressure of oxygen and deoxygenating it with nitrogen gas.
- ODC oxyhemoprotein dissociation curve
- a Clark oxygen electrode detects the change in oxygen tension, which is recorded on the x-axis of an x-y recorder.
- the resulting increase in oxyhemoprotein fraction is simultaneously monitored by dual-wavelength spectrophotometry at 560 nm and 576 nm and displayed on the y-axis. Blood samples are taken from the antemedial vein, anticoagulated with heparin, and kept at 4°C on wet ice until the assay.
- the p50 values for any of the H-NOX proteins can be compared to that of hemoglobin as an indication of the relative affinity of the H-NOX protein for O 2 compared to that of hemoglobin.
- Table 15 lists previously reported p50 values for hemoglobin.
- the viscosity of the H-NOX solutions can be measured using a cone/plate rheometer (model DV-III, Brookfield; Middleboro, MA) with the CPE-40 cone spindle at a shear rate of 200/s. Solutions with viscosities between 1 and 4 centipoise (cP) administered in hemodilution oxygen delivery experiments are reported as safe (Winslow, R. M. et al. (October 2004). "Comparison of PEG-Modified Albumin And Hemoglobin in Extreme Hemodilution in the Rat," JAppl Physiol. 97(4): 1527-1534, U.S. Pat. Nos.
- the viscosity of the H-NOX protein solution is between 1 and 4 cP. Colloid oncotic pressure measurements
- the colloid oncotic pressure can be measured using a colloid osmometer according to the manufacturer's instructions (model 4420, Wescor; Logan, UT). Exemplary methods to measure colloid oncotic pressure are described in Vandegriff, K. D. et al. (November 1997). "Colloid Osmotic Properties of Modified Hemoglobins: Chemically Cross-Linked Versus Polyethylene Glycol Surface-Conjugated," Biophys. Chem. 69(l):23-30, in the world-wide web at "anaesthesiamcq.com/FluidBook/fl2_4.php;" U.S. Pat. Nos.
- colloid oncotic pressure of the H-NOX protein solution is between 20 and 50 mm Hg.
- Two animal models can be used to compare the efficacy of H-NOX proteins as NO carriers to standard nitrate therapy.
- the abdominal aorta is constricted (abdominal aorta constriction or "AC" model) via ventral abdominal laparotomy and application of a constriction tie over an inserted 21 -gauge needle, which is then removed to permit uniform vessel constriction.
- Sham-operated rats undergo similar surgery, but without creation of AC. After surgery, the rats are randomly divided into treatment groups of 14 animals per group (7 on drug and 7 on placebo), and allowed to recover for 7 days.
- Treatments per group are: (1) AC rats administered oral sr-ISDN or placebo; (2) AC rats administered IV one or more wild-type or mutant H-NOX proteins described herein or placebo ⁇ e.g., an inactivated H-NOX protein); (3) and (4) sham- operated rats treated as in (1) or (2), respectively. Treatments are once a day for 12 weeks, after which the animals are sacrificed, and the hearts are excised for standard histopathological analyses for the determination of cardiomyocyte morphology, fibrosis, collage deposition, ventricular diameter, aortic morphology, and other standard analyses for assessing disease progression or prevention.
- Polyethylene catheters are inserted in the external jugular vein, internal carotid artery, and left atrium, filled with heparinized saline, and their ends exteriorized behind the neck.
- a silk ligature is placed around the mid left anterior descending coronary artery, between the first and second diagonal branches, and tied.
- Metal beads are sutured on the anterior, lateral, and posterior epicardial surfaces in the short- axis plane at the mid left ventricular level for consistent echocardiographic orientation for serial topography.
- the pericardium and chest are then closed. Penicillin (1 million units) and streptomycin (1 g) are given intramuscularly, and the dogs are returned to their cages.
- nitrate therapy e.g., one or more wild-type or mutant H-NOX proteins that optionally have been characterized in vitro using any of the assays described herein and optionally have undergone optimization for toxicity and/or pharmacokinetics
- H-NOX protein therapy e.g., one or more wild-type or mutant H-NOX proteins that optionally have been characterized in vitro using any of the assays described herein and optionally have undergone optimization for toxicity and/or pharmacokinetics
- subgroup 1 10 control, 10 nitrate
- subgroup 2 10 control, 10 H-NOX protein
- the surviving dogs are anesthetized, and the hearts are arrested in diastole with an overdose of intravenous potassium chloride, excised, washed in normal saline solution, and weighed. Blood samples are taken for monitoring blood gases, hemograms, and electrolytes. Using standard procedures, the measurements during healing are made (such as ECG's, hemodynamics, etc.), and post-mortem analyses include those measures described above for chronic heart failure (e.g., collagen accumulation, myocyte morphology, etc.).
- H-NOX proteins that are as effective or more effective than ISDN (the standard of care for nitrate-based therapies for acute and chronic heart failure) in the myocardial infarction and/or the chronic AC model experiments are particularly useful for the treatment of myocardial infarction and/or chronic AC. Such H-NOX proteins are expected to also be useful to treat other indications for which delivery of NO is beneficial.
- Reference to "about” a value or parameter herein includes (and describes) embodiments that are directed to that value or parameter per se. For example, description referring to "about X” includes description of "X.”
Landscapes
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Organic Chemistry (AREA)
- General Health & Medical Sciences (AREA)
- Medicinal Chemistry (AREA)
- Engineering & Computer Science (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Veterinary Medicine (AREA)
- Public Health (AREA)
- Animal Behavior & Ethology (AREA)
- Pharmacology & Pharmacy (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- General Chemical & Material Sciences (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Nuclear Medicine, Radiotherapy & Molecular Imaging (AREA)
- Genetics & Genomics (AREA)
- Biochemistry (AREA)
- Gastroenterology & Hepatology (AREA)
- Biophysics (AREA)
- Molecular Biology (AREA)
- Zoology (AREA)
- Toxicology (AREA)
- Cardiology (AREA)
- Insects & Arthropods (AREA)
- Tropical Medicine & Parasitology (AREA)
- Diabetes (AREA)
- Heart & Thoracic Surgery (AREA)
- Hematology (AREA)
- Urology & Nephrology (AREA)
- Vascular Medicine (AREA)
- Epidemiology (AREA)
- Neurology (AREA)
- Biomedical Technology (AREA)
- Dermatology (AREA)
- Hospice & Palliative Care (AREA)
- Immunology (AREA)
- Neurosurgery (AREA)
- Peptides Or Proteins (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
Abstract
Description
Claims
Priority Applications (14)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| NZ573003A NZ573003A (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for the delivery of nitric oxide |
| JP2009512098A JP5698905B2 (en) | 2006-05-22 | 2007-05-21 | Methods for delivery of nitric oxide |
| ES07795153.1T ES2525226T3 (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for the administration of nitric oxide |
| US12/302,004 US8404632B2 (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for the delivery of nitric oxide |
| DK07795153.1T DK2040740T3 (en) | 2006-05-22 | 2007-05-21 | COMPOSITIONS AND PROCEDURES FOR DELIVERING NITROGEN MONOXIDE |
| BRPI0711767A BRPI0711767B8 (en) | 2006-05-22 | 2007-05-21 | pharmaceutical composition of blood gas carrier, use of an h-nox protein, kit, recombinant hnox protein and recombinant nucleic acid |
| AU2007268028A AU2007268028B2 (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for the delivery of nitric oxide |
| CA2653101A CA2653101C (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for the delivery of nitric oxide |
| EP07795153.1A EP2040740B1 (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for the delivery of nitric oxide |
| CN200780027797.2A CN101495134B (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for delivering nitric oxide |
| NZ594456A NZ594456A (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for the delivery of nitric oxide |
| IN600MUN2013 IN2013MN00600A (en) | 2006-05-22 | 2007-05-21 | |
| US13/772,283 US20130289252A1 (en) | 2006-05-22 | 2013-02-20 | Compositions and methods for the delivery of nitric oxide |
| US14/490,597 US9493527B2 (en) | 2006-05-22 | 2014-09-18 | Compositions and methods for the delivery of nitric oxide |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US92150506P | 2006-05-22 | 2006-05-22 | |
| US60/921,505 | 2006-05-22 |
Related Child Applications (2)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| US12/302,004 A-371-Of-International US8404632B2 (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for the delivery of nitric oxide |
| US13/772,283 Continuation US20130289252A1 (en) | 2006-05-22 | 2013-02-20 | Compositions and methods for the delivery of nitric oxide |
Publications (2)
| Publication Number | Publication Date |
|---|---|
| WO2007139767A2 true WO2007139767A2 (en) | 2007-12-06 |
| WO2007139767A3 WO2007139767A3 (en) | 2008-02-07 |
Family
ID=38779160
Family Applications (2)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2007/012133 Ceased WO2007139767A2 (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for the delivery of nitric oxide |
| PCT/US2007/012184 Ceased WO2007139791A2 (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for the delivery of oxygen |
Family Applications After (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2007/012184 Ceased WO2007139791A2 (en) | 2006-05-22 | 2007-05-21 | Compositions and methods for the delivery of oxygen |
Country Status (12)
| Country | Link |
|---|---|
| US (7) | US8404631B2 (en) |
| EP (7) | EP2474554A3 (en) |
| JP (5) | JP5698905B2 (en) |
| CN (5) | CN101495506B (en) |
| AU (2) | AU2007268052B2 (en) |
| BR (2) | BRPI0711901A2 (en) |
| CA (2) | CA2653101C (en) |
| DK (4) | DK2463297T3 (en) |
| ES (4) | ES2611157T3 (en) |
| IN (2) | IN2013MN00600A (en) |
| NZ (4) | NZ573003A (en) |
| WO (2) | WO2007139767A2 (en) |
Cited By (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20110243849A1 (en) * | 2010-04-02 | 2011-10-06 | Marletta Michael A | Heme-binding photoactive polypeptides and methods of use thereof |
| US8404631B2 (en) | 2006-05-22 | 2013-03-26 | The Regents Of The University Of California | Compositions and methods for the delivery of oxygen |
| EP2941440A4 (en) * | 2013-01-07 | 2016-06-15 | Omniox Inc | POLYMERIC FORMS OF H-NOX PROTEINS |
| WO2017143104A1 (en) | 2016-02-16 | 2017-08-24 | Omniox, Inc. | Modulation hypoxia associated with stroke |
| WO2020037052A1 (en) | 2018-08-15 | 2020-02-20 | Omniox, Inc. | H-nox proteins for treating cardiovascular and pulmonary conditions |
| US12285463B2 (en) | 2015-03-17 | 2025-04-29 | Omniox, Inc. | Modulation of tumor immunity by protein-mediated O2 delivery |
Families Citing this family (38)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP6467073B2 (en) * | 2018-01-04 | 2019-02-06 | オムニオクス, インコーポレイテッド | Polymer form of H-NOX protein |
| MY204234A (en) | 2018-08-27 | 2024-08-16 | Regeneron Pharma | Use of raman spectroscopy in downstream purification |
| CR20210687A (en) | 2019-06-25 | 2022-03-03 | Gilead Sciences Inc | FLT3L-Fc FUSION PROTEINS AND METHODS OF USE |
| CN114555123B (en) | 2019-10-18 | 2024-04-02 | 四十七公司 | Combination therapy for the treatment of myelodysplastic syndrome and acute myeloid leukemia |
| KR20220091576A (en) | 2019-10-31 | 2022-06-30 | 포티 세븐, 인코포레이티드 | Anti-CD47 and Anti-CD20 Based Treatment of Hematological Cancers |
| TWI778443B (en) | 2019-11-12 | 2022-09-21 | 美商基利科學股份有限公司 | Mcl1 inhibitors |
| LT4081305T (en) | 2019-12-24 | 2024-11-25 | Carna Biosciences, Inc. | Diacylglycerol kinase modulating compounds |
| KR20260017503A (en) | 2020-02-14 | 2026-02-05 | 길리애드 사이언시즈, 인코포레이티드 | Antibodies and fusion proteins that bind to ccr8 and uses thereof |
| JP7564888B2 (en) | 2020-05-01 | 2024-10-09 | ギリアード サイエンシーズ, インコーポレイテッド | CD73-inhibitory 2,4-dioxopyrimidine compounds |
| TW202302145A (en) | 2021-04-14 | 2023-01-16 | 美商基利科學股份有限公司 | Co-inhibition of cd47/sirpα binding and nedd8-activating enzyme e1 regulatory subunit for the treatment of cancer |
| TW202313094A (en) | 2021-05-18 | 2023-04-01 | 美商基利科學股份有限公司 | Methods of using flt3l-fc fusion proteins |
| CA3222277A1 (en) | 2021-06-23 | 2022-12-29 | Gilead Sciences, Inc. | Diacylglyercol kinase modulating compounds |
| WO2022271659A1 (en) | 2021-06-23 | 2022-12-29 | Gilead Sciences, Inc. | Diacylglyercol kinase modulating compounds |
| KR20240005901A (en) | 2021-06-23 | 2024-01-12 | 길리애드 사이언시즈, 인코포레이티드 | Diacylglycerol Kinase Modulating Compounds |
| WO2022271677A1 (en) | 2021-06-23 | 2022-12-29 | Gilead Sciences, Inc. | Diacylglyercol kinase modulating compounds |
| US20230183216A1 (en) | 2021-10-28 | 2023-06-15 | Gilead Sciences, Inc. | Pyridizin-3(2h)-one derivatives |
| EP4422756A1 (en) | 2021-10-29 | 2024-09-04 | Gilead Sciences, Inc. | Cd73 compounds |
| CN118488946A (en) | 2021-12-22 | 2024-08-13 | 吉利德科学公司 | IKAROS zinc finger family degradation agent and application thereof |
| EP4452415B1 (en) | 2021-12-22 | 2026-02-25 | Gilead Sciences, Inc. | Ikaros zinc finger family degraders and uses thereof |
| TW202340168A (en) | 2022-01-28 | 2023-10-16 | 美商基利科學股份有限公司 | Parp7 inhibitors |
| CA3253296A1 (en) | 2022-03-17 | 2023-09-21 | Gilead Sciences, Inc. | Ikaros zinc finger family degraders and uses thereof |
| AU2023240346A1 (en) | 2022-03-24 | 2024-09-19 | Gilead Sciences, Inc. | Combination therapy for treating trop-2 expressing cancers |
| TWI876305B (en) | 2022-04-05 | 2025-03-11 | 美商基利科學股份有限公司 | Combination therapy for treating colorectal cancer |
| WO2023205719A1 (en) | 2022-04-21 | 2023-10-26 | Gilead Sciences, Inc. | Kras g12d modulating compounds |
| US20240116928A1 (en) | 2022-07-01 | 2024-04-11 | Gilead Sciences, Inc. | Cd73 compounds |
| WO2024064668A1 (en) | 2022-09-21 | 2024-03-28 | Gilead Sciences, Inc. | FOCAL IONIZING RADIATION AND CD47/SIRPα DISRUPTION ANTICANCER COMBINATION THERAPY |
| EP4638436A1 (en) | 2022-12-22 | 2025-10-29 | Gilead Sciences, Inc. | Prmt5 inhibitors and uses thereof |
| CN120882725A (en) | 2023-04-11 | 2025-10-31 | 吉利德科学公司 | KRAS-regulated compounds |
| WO2024220917A1 (en) | 2023-04-21 | 2024-10-24 | Gilead Sciences, Inc. | Prmt5 inhibitors and uses thereof |
| US20250042922A1 (en) | 2023-06-30 | 2025-02-06 | Gilead Sciences, Inc. | Kras modulating compounds |
| US20250066328A1 (en) | 2023-07-26 | 2025-02-27 | Gilead Sciences, Inc. | Parp7 inhibitors |
| AU2024297978A1 (en) | 2023-07-26 | 2026-02-05 | Gilead Sciences, Inc. | Parp7 inhibitors |
| WO2025054347A1 (en) | 2023-09-08 | 2025-03-13 | Gilead Sciences, Inc. | Kras g12d modulating compounds |
| AU2024337913A1 (en) | 2023-09-08 | 2026-03-26 | Gilead Sciences, Inc. | Pyrimidine-containing polycyclic derivatives as kras g12d modulating compounds |
| US20250154172A1 (en) | 2023-11-03 | 2025-05-15 | Gilead Sciences, Inc. | Prmt5 inhibitors and uses thereof |
| WO2025137640A1 (en) | 2023-12-22 | 2025-06-26 | Gilead Sciences, Inc. | Azaspiro wrn inhibitors |
| US20250376484A1 (en) | 2024-05-21 | 2025-12-11 | Gilead Sciences, Inc. | Prmt5 inhibitors and uses thereof |
| WO2026039365A1 (en) | 2024-08-12 | 2026-02-19 | Gilead Sciences, Inc. | Kras modulating compounds |
Family Cites Families (16)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4873192A (en) | 1987-02-17 | 1989-10-10 | The United States Of America As Represented By The Department Of Health And Human Services | Process for site specific mutagenesis without phenotypic selection |
| US6022849A (en) | 1987-05-16 | 2000-02-08 | Baxter Biotech Technology Saarl | Mutant recombinant hemoglobins containing heme pocket mutations |
| US5731454A (en) * | 1990-02-12 | 1998-03-24 | Virginia Commonwealth University | Allosteric modifiers of hemoglobin useful for decreasing oxygen affinity and preserving oxygen carrying capability of stored blood |
| WO1995027040A1 (en) | 1994-04-01 | 1995-10-12 | Unisyn Technologies, Inc. | Culture media additives for hollow fiber bioreactors |
| AU735799C (en) * | 1997-02-28 | 2005-04-28 | Regents Of The University Of California, The | Methods and compositions for optimisation of oxygen transport by cell-free systems |
| AU750295B2 (en) | 1997-05-02 | 2002-07-11 | Baxter Biotech Technology S.A.R.L. | Hemoglobin mutants with increased soluble expression and/or reduced nitric oxide scavenging |
| DE19837015C2 (en) * | 1998-08-14 | 2003-03-20 | Vasopharm Biotech Gmbh | Isolated and purified human soluble guanylyl cyclase alpha1 / beta1 (hsGCalpha1 / beta1) |
| US6773613B1 (en) * | 1998-10-15 | 2004-08-10 | Sangart, Inc. | Method for production of stroma-free hemoglobin |
| CA2395136A1 (en) * | 1999-12-22 | 2001-06-28 | The Scripps Research Institute | Angiogenesis and vascular permeability modulators and inhibitors |
| WO2002002757A2 (en) | 2000-06-29 | 2002-01-10 | Incyte Genomics, Inc. | Adenylyl and guanylyl cyclases |
| US20030096240A1 (en) * | 2000-09-19 | 2003-05-22 | Ferid Murad | Genomic organization of mouse and human sGC |
| US6780849B2 (en) * | 2000-12-21 | 2004-08-24 | Scimed Life Systems, Inc. | Lipid-based nitric oxide donors |
| US20030153491A1 (en) | 2002-01-11 | 2003-08-14 | Winslow Robert M. | Methods and compositions for oxygen transport comprising a high oxygen affinity modified hemoglobin |
| EP1545595B1 (en) * | 2002-08-30 | 2010-07-07 | Biorexis Pharmaceutical Corporation | Modified transferrin fusion proteins comprising duplicate transferrin amino or carboxy terminal domains |
| CN1660100A (en) * | 2002-12-09 | 2005-08-31 | 上海倍艺医药科技有限公司 | Medication for treating diabetes and complication |
| DK2463297T3 (en) | 2006-05-22 | 2017-01-09 | Univ California | Oxygen Compositions and Methods |
-
2007
- 2007-05-21 DK DK11189814.4T patent/DK2463297T3/en active
- 2007-05-21 NZ NZ573003A patent/NZ573003A/en unknown
- 2007-05-21 ES ES11189814.4T patent/ES2611157T3/en active Active
- 2007-05-21 CA CA2653101A patent/CA2653101C/en active Active
- 2007-05-21 EP EP11189818A patent/EP2474554A3/en not_active Withdrawn
- 2007-05-21 CN CN200780027798.7A patent/CN101495506B/en active Active
- 2007-05-21 NZ NZ594457A patent/NZ594457A/en unknown
- 2007-05-21 US US12/302,002 patent/US8404631B2/en active Active
- 2007-05-21 EP EP11189808A patent/EP2463295A1/en not_active Withdrawn
- 2007-05-21 EP EP11189814.4A patent/EP2463297B1/en active Active
- 2007-05-21 NZ NZ594456A patent/NZ594456A/en unknown
- 2007-05-21 IN IN600MUN2013 patent/IN2013MN00600A/en unknown
- 2007-05-21 AU AU2007268052A patent/AU2007268052B2/en active Active
- 2007-05-21 NZ NZ573004A patent/NZ573004A/en unknown
- 2007-05-21 ES ES07795174T patent/ES2399662T3/en active Active
- 2007-05-21 CN CN201410766335.4A patent/CN104689300B/en active Active
- 2007-05-21 AU AU2007268028A patent/AU2007268028B2/en active Active
- 2007-05-21 CN CN201610487363.1A patent/CN106075388B/en active Active
- 2007-05-21 EP EP11189807.8A patent/EP2463294B1/en active Active
- 2007-05-21 EP EP07795174A patent/EP2032156B1/en active Active
- 2007-05-21 US US12/302,004 patent/US8404632B2/en active Active
- 2007-05-21 ES ES07795153.1T patent/ES2525226T3/en active Active
- 2007-05-21 WO PCT/US2007/012133 patent/WO2007139767A2/en not_active Ceased
- 2007-05-21 DK DK07795153.1T patent/DK2040740T3/en active
- 2007-05-21 CN CN201810357346.5A patent/CN108273040A/en active Pending
- 2007-05-21 BR BRPI0711901-1A patent/BRPI0711901A2/en not_active Application Discontinuation
- 2007-05-21 IN IN601MUN2013 patent/IN2013MN00601A/en unknown
- 2007-05-21 DK DK07795174.7T patent/DK2032156T3/en active
- 2007-05-21 JP JP2009512098A patent/JP5698905B2/en active Active
- 2007-05-21 WO PCT/US2007/012184 patent/WO2007139791A2/en not_active Ceased
- 2007-05-21 BR BRPI0711767A patent/BRPI0711767B8/en active IP Right Grant
- 2007-05-21 ES ES11189807.8T patent/ES2610653T3/en active Active
- 2007-05-21 EP EP11189812A patent/EP2463296A1/en not_active Withdrawn
- 2007-05-21 EP EP07795153.1A patent/EP2040740B1/en active Active
- 2007-05-21 JP JP2009512107A patent/JP5698906B2/en active Active
- 2007-05-21 CN CN200780027797.2A patent/CN101495134B/en active Active
- 2007-05-21 DK DK11189807.8T patent/DK2463294T3/en active
- 2007-05-21 CA CA2653080A patent/CA2653080C/en active Active
-
2013
- 2013-02-20 US US13/772,283 patent/US20130289252A1/en not_active Abandoned
- 2013-02-20 US US13/772,281 patent/US20130288970A1/en not_active Abandoned
-
2014
- 2014-09-04 JP JP2014180132A patent/JP6118296B2/en active Active
- 2014-09-04 JP JP2014180140A patent/JP6118772B2/en active Active
- 2014-09-17 US US14/489,395 patent/US9493526B2/en active Active
- 2014-09-18 US US14/490,597 patent/US9493527B2/en active Active
-
2016
- 2016-10-12 US US15/292,012 patent/US10202428B2/en active Active
-
2017
- 2017-03-27 JP JP2017061455A patent/JP6458070B2/en active Active
Non-Patent Citations (10)
| Title |
|---|
| "Blood Substitutes", 2006, ACADEMIC PRESS |
| BOBOFCHAK, K. M. ET AL.: "A Recombinant Polymeric Hemoglobin With Conformational, Functional, And Physiological characteristics of an in vivo 02 transporter", AM. J PHYSIOL. HEART CIRC. PHYSIOL., vol. 285, no. 2, August 2003 (2003-08-01), pages H549 - H561 |
| BOON, E. M. ET AL.: "Nitric Oxide Binding to Prokaryotic Homologs of the Solube Guanylate Cyclase ? 1 HONOX Domain", J BIOL. CHEM., vol. 281, no. 31, August 2006 (2006-08-01), pages 21892 - 21902 |
| BOON, E.M. ET AL.: "Molecular Basis For NO Selectivity in Soluble Guanylate Cyclase", NATURE CHEMICAL BIOLOGY, vol. 1, 2005, pages 53 - 59 |
| BOON; MARLETTA, CURRENT OPINION IN CHEMICAL BIOLOGY, vol. 9, 2005, pages 441 - 446 |
| KAROW, D. S. ET AL.: "Spectroscopic Characterization of The Soluble Guanylate Cyclase-Like Heme Domains From Vibrio Cholerae And Thermoanaerobacter Tengcongensis", BIOCHEMISTRY, vol. 43, no. 31, 10 August 2004 (2004-08-10), pages 10203 - 10211, XP055053530, DOI: doi:10.1021/bi049374l |
| LAKSHMINARAYAN ET AL.: "Ancient conserved domains shared by animal soluble guanylyl cyclases and bacterial signaling proteins", BMG GENOMICS, vol. 4, 2003, pages 5 - 13 |
| PELLICENA, P. ET AL.: "Crystal Structure of An Oxygen-Binding Heme Domain Related to Soluble Guanylate Cyclases", PROC NATL. ACAD SCI USA, vol. 101, no. 35, 31 August 2004 (2004-08-31), pages 12854 - 12859, XP055018589, DOI: doi:10.1073/pnas.0405188101 |
| See also references of EP2040740A4 |
| VILLARD, J. W.: "Use of a Blood Substitute to Determine Instantaneous Murine Right Ventricular Thickening with Optical Coherence Tomography", CIRCULATION, vol. 105, 2002, pages 1843 - 1849 |
Cited By (15)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US8404631B2 (en) | 2006-05-22 | 2013-03-26 | The Regents Of The University Of California | Compositions and methods for the delivery of oxygen |
| US9493527B2 (en) | 2006-05-22 | 2016-11-15 | The Regents Of The University Of California | Compositions and methods for the delivery of nitric oxide |
| US9493526B2 (en) | 2006-05-22 | 2016-11-15 | The Regents Of The University Of California | Compositions and methods for the delivery of oxygen |
| US10202428B2 (en) | 2006-05-22 | 2019-02-12 | The Regents Of The University Of California | Compositions and methods for the delivery of oxygen |
| US20110243849A1 (en) * | 2010-04-02 | 2011-10-06 | Marletta Michael A | Heme-binding photoactive polypeptides and methods of use thereof |
| EP3922643A1 (en) | 2013-01-07 | 2021-12-15 | Omniox, Inc. | Polymeric forms of h-nox proteins |
| EP2941440A4 (en) * | 2013-01-07 | 2016-06-15 | Omniox Inc | POLYMERIC FORMS OF H-NOX PROTEINS |
| US10385116B2 (en) | 2013-01-07 | 2019-08-20 | Omniox, Inc. | Polymeric forms of H-NOX proteins |
| US11987614B2 (en) | 2013-01-07 | 2024-05-21 | Omniox, Inc. | Polymeric forms of H-NOX proteins |
| US10766947B2 (en) | 2013-01-07 | 2020-09-08 | Omniox, Inc. | Polymeric forms of H-NOX proteins |
| US11117952B2 (en) | 2013-01-07 | 2021-09-14 | Omniox, Inc. | Polymeric forms of H-NOX proteins |
| US12285463B2 (en) | 2015-03-17 | 2025-04-29 | Omniox, Inc. | Modulation of tumor immunity by protein-mediated O2 delivery |
| CN109069582A (en) * | 2016-02-16 | 2018-12-21 | 欧姆尼奥克斯公司 | The adjusting of anoxic related with apoplexy |
| WO2017143104A1 (en) | 2016-02-16 | 2017-08-24 | Omniox, Inc. | Modulation hypoxia associated with stroke |
| WO2020037052A1 (en) | 2018-08-15 | 2020-02-20 | Omniox, Inc. | H-nox proteins for treating cardiovascular and pulmonary conditions |
Also Published As
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US9493527B2 (en) | Compositions and methods for the delivery of nitric oxide | |
| AU2013201114B2 (en) | Compositions and Methods for the Delivery of Nitric Oxide | |
| HK1172038A (en) | Compositions and methods for the delivery of nitric oxide | |
| HK1130265B (en) | Compositions and methods for the delivery of nitric oxide | |
| HK1130265A (en) | Compositions and methods for the delivery of nitric oxide | |
| AU2013201117A1 (en) | Compositions and Methods for the Delivery of Oxygen | |
| HK1137332A (en) | Compositions and methods for the delivery of nitric oxide | |
| HK1129688B (en) | Compositions and methods for the delivery of oxygen |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| WWE | Wipo information: entry into national phase |
Ref document number: 200780027797.2 Country of ref document: CN |
|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 07795153 Country of ref document: EP Kind code of ref document: A2 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2007268028 Country of ref document: AU |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 573003 Country of ref document: NZ Ref document number: 2496/MUMNP/2008 Country of ref document: IN |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2009512098 Country of ref document: JP Ref document number: 2653101 Country of ref document: CA |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 2007268028 Country of ref document: AU Date of ref document: 20070521 Kind code of ref document: A |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2007795153 Country of ref document: EP |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 12302004 Country of ref document: US |
|
| ENP | Entry into the national phase |
Ref document number: PI0711767 Country of ref document: BR Kind code of ref document: A2 Effective date: 20081119 |