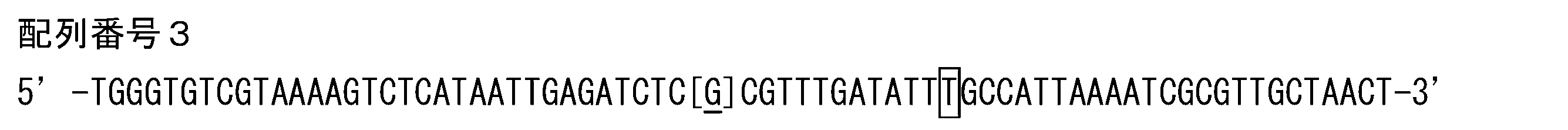WO2016174795A1 - トマト植物の葉かび病抵抗性マーカー、葉かび病抵抗性トマト植物、葉かび病抵抗性トマト植物の製造方法、および葉かび病抵抗性トマト植物のスクリーニング方法 - Google Patents
トマト植物の葉かび病抵抗性マーカー、葉かび病抵抗性トマト植物、葉かび病抵抗性トマト植物の製造方法、および葉かび病抵抗性トマト植物のスクリーニング方法 Download PDFInfo
- Publication number
- WO2016174795A1 WO2016174795A1 PCT/JP2015/084750 JP2015084750W WO2016174795A1 WO 2016174795 A1 WO2016174795 A1 WO 2016174795A1 JP 2015084750 W JP2015084750 W JP 2015084750W WO 2016174795 A1 WO2016174795 A1 WO 2016174795A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- snp
- leaf mold
- solcap
- chromosome
- tomato plant
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Images
Classifications
-
- A—HUMAN NECESSITIES
- A01—AGRICULTURE; FORESTRY; ANIMAL HUSBANDRY; HUNTING; TRAPPING; FISHING
- A01H—NEW PLANTS OR NON-TRANSGENIC PROCESSES FOR OBTAINING THEM; PLANT REPRODUCTION BY TISSUE CULTURE TECHNIQUES
- A01H6/00—Angiosperms, i.e. flowering plants, characterised by their botanic taxonomy
- A01H6/82—Solanaceae, e.g. pepper, tobacco, potato, tomato or eggplant
- A01H6/825—Solanum lycopersicum [tomato]
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6888—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for detection or identification of organisms
- C12Q1/6895—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for detection or identification of organisms for plants, fungi or algae
-
- A—HUMAN NECESSITIES
- A01—AGRICULTURE; FORESTRY; ANIMAL HUSBANDRY; HUNTING; TRAPPING; FISHING
- A01H—NEW PLANTS OR NON-TRANSGENIC PROCESSES FOR OBTAINING THEM; PLANT REPRODUCTION BY TISSUE CULTURE TECHNIQUES
- A01H1/00—Processes for modifying genotypes ; Plants characterised by associated natural traits
- A01H1/02—Methods or apparatus for hybridisation; Artificial pollination ; Fertility
- A01H1/021—Methods of breeding using interspecific crosses, i.e. interspecies crosses
-
- A—HUMAN NECESSITIES
- A01—AGRICULTURE; FORESTRY; ANIMAL HUSBANDRY; HUNTING; TRAPPING; FISHING
- A01H—NEW PLANTS OR NON-TRANSGENIC PROCESSES FOR OBTAINING THEM; PLANT REPRODUCTION BY TISSUE CULTURE TECHNIQUES
- A01H1/00—Processes for modifying genotypes ; Plants characterised by associated natural traits
- A01H1/04—Processes of selection involving genotypic or phenotypic markers; Methods of using phenotypic markers for selection
- A01H1/045—Processes of selection involving genotypic or phenotypic markers; Methods of using phenotypic markers for selection using molecular markers
-
- A—HUMAN NECESSITIES
- A01—AGRICULTURE; FORESTRY; ANIMAL HUSBANDRY; HUNTING; TRAPPING; FISHING
- A01H—NEW PLANTS OR NON-TRANSGENIC PROCESSES FOR OBTAINING THEM; PLANT REPRODUCTION BY TISSUE CULTURE TECHNIQUES
- A01H1/00—Processes for modifying genotypes ; Plants characterised by associated natural traits
- A01H1/12—Processes for modifying agronomic input traits, e.g. crop yield
- A01H1/122—Processes for modifying agronomic input traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance
- A01H1/1245—Processes for modifying agronomic input traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance for biotic stress resistance, e.g. pathogen, pest or disease resistance
- A01H1/1255—Processes for modifying agronomic input traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance for biotic stress resistance, e.g. pathogen, pest or disease resistance for fungal resistance
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/82—Vectors or expression systems specially adapted for eukaryotic hosts for plant cells, e.g. plant artificial chromosomes (PACs)
- C12N15/8241—Phenotypically and genetically modified plants via recombinant DNA technology
- C12N15/8261—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield
- C12N15/8271—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance
- C12N15/8279—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance for biotic stress resistance, pathogen resistance, disease resistance
- C12N15/8282—Phenotypically and genetically modified plants via recombinant DNA technology with agronomic (input) traits, e.g. crop yield for stress resistance, e.g. heavy metal resistance for biotic stress resistance, pathogen resistance, disease resistance for fungal resistance
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/13—Plant traits
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/156—Polymorphic or mutational markers
Definitions
- the present invention relates to a leaf mold resistance marker for tomato plants, a leaf mold resistant tomato plant, a method for producing a leaf mold resistant tomato plant, and a method for screening for a leaf mold resistant tomato plant.
- Non-patent Document 1 Non-patent Document 1
- Non-Patent Documents 2 and 3 An attempt has been made to cultivate varieties that are resistant to leaf mold by using a resistance gene against leaf mold. However, a fungal fungus capable of infecting tomato plants containing these resistance genes has emerged and has become a problem (Non-Patent Documents 2 and 3).
- the present invention is a novel leaf mold resistance marker for tomato plants exhibiting a new dominant leaf mold resistance, a leaf mold resistant tomato plant containing a dominant leaf mold resistance locus, and the same. It aims at providing the manufacturing method of a leaf-mold-resistant tomato plant, and the screening method of a leaf-mold-resistant tomato plant.
- the leaf mold resistance marker of the tomato plant of the present invention is at least one of the leaf mold resistance locus on chromosome 1 and the leaf mold resistance locus on chromosome 6.
- Including a leaf mold resistance locus of The leaf mold resistance locus on the first chromosome is identified by at least one SNP marker selected from the group consisting of solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′;
- the leaf mold resistance locus on the sixth chromosome is identified by at least one SNP marker selected from the group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_3521, SG_70, and SG_70 ′.
- the leaf mold resistant tomato plant of the present invention comprises at least one leaf mold resistant locus on chromosome 1 and a leaf mold resistant locus on chromosome 6.
- the leaf mold resistance locus on the first chromosome is identified by at least one SNP marker selected from the group consisting of solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′;
- the leaf mold resistance locus on the sixth chromosome is identified by at least one SNP marker selected from the group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_3521, SG_70, and SG_70 ′.
- the method for producing a leaf mold resistant tomato plant of the present invention comprises the following steps (a) and (b).
- the screening method for leaf mold resistant tomato plants of the present invention is the first as a parent for producing leaf mold resistant tomato plants by crossing from a test tomato plant as a leaf mold resistance marker for tomato plants. Selecting a leafy mildew resistant tomato plant comprising at least one of a leafy mildew resistance locus on a chromosome and a leafy mildew resistance locus on chromosome 6.
- the leaf mold resistance locus on the first chromosome is identified by at least one SNP marker selected from the group consisting of solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′;
- the leaf mold resistance locus on the sixth chromosome is identified by at least one SNP marker selected from the group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_3521, SG_70, and SG_70 ′.
- first resistance locus sits on the first chromosome and consists of solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′. It was found that it is identified (hereinafter also referred to as “regulation”) by at least one SNP marker selected from the group.
- a second new leaf mold resistance locus (hereinafter, also referred to as “second resistance locus”) sits on the sixth chromosome, and solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_3521, SG_70, SG_70 And at least one SNP marker selected from the group consisting of SG_70 ′. Furthermore, the tomato plant containing the leaf mold resistance marker exhibits a dominant leaf mold resistance. For this reason, according to the leaf mold resistance marker of the tomato plant of the present invention, for example, a leaf mold resistant tomato plant can be easily screened.
- the leaf mold resistant tomato plant of the present invention includes, for example, at least one of the first resistance locus and the second resistance locus, and therefore exhibits, for example, leaf mold resistance. Is possible.
- the tomato plant resistant to leaf mold of the present invention contains, for example, a dominant resistance locus, it is possible to obtain a progeny showing dominant resistance to leaf mold by crossing with other tomato plants. Can do.
- the tomato plant containing the above-mentioned leaf mold resistance marker also exhibits resistance to, for example, a leaf mold fungus capable of infecting a tomato plant containing the above-mentioned prior art literature leaf mold resistance gene. For this reason, the leaf mold resistant tomato plant of the present invention does not require conventional pesticide control, and thus, for example, the labor and cost problems of the pesticide spraying can be avoided.
- FIG. 1 is a schematic diagram showing the relative locus positions of SNP (single nucleotide polymorphism) markers on the first chromosome.
- FIG. 2 is a schematic diagram showing the relative sitting position of the SNP marker in the sixth chromosome.
- Tomato plant leaf mold resistance marker The tomato plant leaf mold resistance marker of the present invention (hereinafter also referred to as "resistance marker”) is, as described above, the leaf mold resistance on the first chromosome. And a fungal resistance locus on at least one of the loci and chromosome 6 on the sixth chromosome, wherein the fungal resistance loci on the first chromosome are solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658.
- the resistance marker of the present invention includes at least one of the leaf mold resistance loci on the first chromosome and the leaf mold resistance locus on the sixth chromosome.
- a leaf mold resistance locus on the chromosome is identified with at least one SNP marker selected from the group consisting of solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′, It is characterized by being identified by at least one SNP marker selected from the group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ', solcap_snp_sl_3521, SG_70, and SG_70', and others Configuration and conditions are not particularly limited.
- tomato plant is a plant that is classified as Section Lycopersicon in Subgenus Solmum sensu stricto of Solanum Solanum, as a specific example, S. lycopersicum , S. peruvianum , S. arcumum Peralta , S. chilense , S. corneliomulleri , S. huaylasense Peralta , S. cheesmaniae (L. Riley) Fosberg, S. chmielewskii , S. galapagense S. et al . C. Darwin & Peralta, S. habrochaites , S. neorikki , S. pennelli , S. pimpinellifolium and the like, preferably, cross is easy S. Lycopersicum .
- examples of the mold fungus include filamentous fungi.
- examples of the filamentous fungus include Fulvia fulva .
- the fulvia fulva is also referred to as, for example, Pasalora fulva or Cladosporium fulvum .
- leaf mold resistance is also referred to as “leaf mold resistance”, for example.
- the resistance means, for example, the ability to inhibit or suppress the occurrence and progression of diseases caused by infection with fungal pathogens, specifically, for example, the absence of disease, the progression of the disease that has occurred, and Any of suppression of the progression of the disease that has occurred (also referred to as “inhibition”) may be used.
- the leaf mold resistance marker of the present invention includes at least one of the leaf mold resistance locus on the first chromosome and the leaf mold resistance locus on the sixth chromosome.
- the tomato plant containing the leaf mold resistance marker contains the first resistance locus
- the tomato plant having leaf mold resistance is, for example, any one other than the first chromosome instead of the first chromosome.
- the first resistance locus may be included on the chromosome.
- the tomato plant containing the first resistance locus has chromosome 2, chromosome 3, chromosome 4, chromosome 5, chromosome 6, chromosome 7, chromosome 8, chromosome 9,
- the first resistance locus may be included on any chromosome of chromosome, chromosome 11 and chromosome 12.
- the tomato plant containing the leaf mold resistance marker contains the second resistance locus
- the tomato plant having leaf mold resistance is, for example, in place of the sixth chromosome, except for the sixth chromosome.
- the second resistance locus may be included on the chromosome of the throat.
- the tomato plant containing the second resistance locus has chromosome 1, chromosome 2, chromosome 3, chromosome 4, chromosome 5, chromosome 7, chromosome 8, chromosome 9,
- the second resistance locus may be contained on any chromosome of chromosome, chromosome 11 and chromosome 12.
- the tomato plant having leaf mold resistance includes the first resistance locus and the second resistance locus
- the tomato plant including the leaf resistance plant locus is on the same chromosome.
- the first resistance locus and the second resistance locus may be included, or the first resistance locus and the second resistance locus may be included on different chromosomes.
- the leaf mold resistance locus means a quantitative trait locus or gene region that provides leaf mold resistance.
- the quantitative trait locus generally means a chromosomal region involved in the expression of quantitative traits.
- QTL can be defined using molecular markers that indicate specific loci on the chromosome. Techniques for defining QTL using the molecular markers are well known in the art.
- the molecular marker used for defining the leaf mold resistance locus is not particularly limited.
- the molecular markers include SNP markers, AFLP (amplified fragment length polymorphism) markers, RFLP (restriction fragment length polymorphism) markers, microsatellite markers, SCAR (sequence-characterized amplified region) markers, and CAPS. (Cleaved amplified polymorphic sequence) markers and the like.
- the SNP marker for example, one SNP may be used as the SNP marker, or a combination of two or more SNPs may be used as the SNP marker.
- the resistance marker of the present invention may include only the first resistance locus or only the second resistance locus.
- the tomato plant containing the resistance marker for example, includes both the first resistance locus and the second resistance locus, whereby leaf mold resistance is further improved.
- the tomato plant in which each of the first resistance locus and the second resistance locus is a heterozygous type of resistance and susceptibility is, for example, the first resistance locus. Shows higher leaf mold resistance than tomato plants that are resistant homozygous, and the second resistance locus is resistant to leaf mold equivalent to tomato plants that are resistant homozygous. Indicates.
- the resistance marker of the present invention includes a leaf mold resistance locus on the first chromosome and a leaf mold resistance locus on the sixth chromosome.
- each resistance locus will be described.
- the SNP marker identifies the first resistance locus.
- the first resistance locus may be further defined by (1-2) a nucleotide sequence including the SNP marker, or (1-3) a nucleotide sequence in a region between two SNP marker sites. Or may be defined by a combination thereof. When prescribed
- the first resistance locus is specified by (1-1), but the first resistance locus is not limited to this.
- the first resistance locus is described in (1-1). Instead, it may be specified by (1-2) or (1-3), or may be specified by a combination of (1-2) and (1-3). Combination of (1-1) and (1-2) Combination of (1-1) and (1-3) (1-1), (1-2) and (1-3) Combination
- the first resistance locus is defined by the SNP marker as shown in (1-1).
- the SNP marker is not particularly limited, and examples thereof include solcap_snp_sl_60160, solcap_snp_sl_8658, solcap_snp_sl_8658 ′, and the like.
- the notation of the SNP marker represented by “solcap_snp_sl_number” excluding solcap_snp_sl_8658 ′ can be understood by those skilled in the art from the technical common sense at the time of filing of the present application. It can be viewed on the consortium website (http://solgenomics.net/).
- solcap_snp_sl_8658 ′ is a SNP marker newly identified by the present inventors, and if it is a person skilled in the art in the art, based on the base sequence containing these SNP markers described later, the sitting position of the SNP marker Can be identified.
- Reference 1 Hamilton JP, Sim SC, Stoffel K, Van Deynze A, Buell CR, et al. (2012) “Single Nucleotide Polymorphism Discovery in Cultivated Tomato via Sequencing by Synthesis.”, The Plant Genome 5.
- the solcap_snp_sl_60160 (hereinafter also referred to as “SNP (a)”) indicates a polymorphism in which the underlined base surrounded by parentheses in SEQ ID NO: 1 is T, for example. That is, for example, when the underlined base is T, the tomato plant is resistant to leaf mold, and when it is a base other than T (for example, C), it indicates that it is susceptible to leaf mold.
- the base sequence of SEQ ID NO: 1 can be obtained, for example, from a tomato plant deposited under accession number FERM BP-22282 described later.
- SEQ ID NO: 1 5'-TGATGGAGGAAACATTACATTCTAATATTTTCGCAGCAAACATCTACAC [ T ] GTTTGATGCTTTTAATGTATCAGCCTGTTATTCAGAA-3 '
- the SNP (a) can also be specified from known information on a database such as the above-mentioned web site, for example.
- the base sequence of SEQ ID NO: 6, for example, is the base sequence of Heinz (variety name, Heinz 1706) except for the SNP (a), and the underlined base surrounded by parentheses corresponds to the SNP (a) It is polymorphic. That is, for example, when the underlined base is T, it is resistant to leaf mold, and when it is a base other than T (for example, C), it indicates that it is susceptible to leaf mold.
- the position of the SNP (a) can be identified from information on the known base sequence of Heinz.
- the solcap_snp_sl_8658 (hereinafter also referred to as “SNP (b)”) indicates, for example, a polymorphism in which the underlined base surrounded by parentheses in SEQ ID NO: 2 is T. That is, for example, when the underlined base is T, the tomato plant is resistant to leaf mold, and when it is a base other than T (for example, C), it indicates that it is susceptible to leaf mold.
- the base sequence of SEQ ID NO: 2 can be obtained from, for example, a tomato plant deposited under the accession number FERM BP-22282 described later.
- the SNP (b) can also be specified from known information on a database such as the above-mentioned web site, for example.
- the base sequence of SEQ ID NO: 7 is the base sequence of Heinz except for, for example, the SNP (b) and solcap_snp_sl_8658 ′ described later, and the underlined base surrounded by parentheses is a polymorphism corresponding to the SNP (b) It is. That is, for example, when the underlined base is T, the tomato plant is resistant to leaf mold, and when it is a base other than T (for example, C), it indicates that it is susceptible to leaf mold.
- the position of the SNP (b) can be identified from information on the known Heinz base sequence, for example.
- the solcap_snp_sl_8658 ′ (hereinafter also referred to as “SNP (b ′)”) indicates, for example, a polymorphism in which the base surrounded by the square in SEQ ID NO: 2 is C. That is, for example, when the base surrounded by the square is C, the tomato plant is resistant to leaf mold, and when it is a base other than C (for example, T), it indicates that it is susceptible to leaf mold.
- the SNP (b ') can also be specified from known information on a database such as the above-mentioned website.
- the base sequence of SEQ ID NO: 7 is, for example, a base sequence of Heinz except for the SNP (b) and SNP (b ′), and the base surrounded by a square is a polymorphism corresponding to the SNP (b ′) It is. That is, for example, when the base surrounded by the square is C, the tomato plant is resistant to leaf mold, and when it is a base other than C (for example, T), it indicates that it is susceptible to leaf mold.
- the position of the SNP (b ′) can be identified from information on the known Heinz base sequence, for example.
- the sitting position of the SNP marker on the chromosome is not particularly limited.
- solcap_snp_sl_60250 and solcap_snp_sl_600089 indicate the use positions of the solcap_snp_sl_60160, solcap_snp_sl_8658 ', and solcap_snp_sl_8658 in the first chromosome of the tomato plant. Solcap_snp_sl_60250 and solcap_snp_sl_600089 will be described later.
- the number of the SNP markers included in the first resistance locus is not particularly limited, and may be any one, two, or three of the SNP markers, for example.
- the relationship between these three types of polymorphisms (SNP markers) and leaf mold resistance has not been reported so far, and it was first discovered by the present inventors in leaf mold resistance. It is a new polymorphism involved.
- the combination of the SNP markers is not particularly limited, and examples thereof include the following combinations.
- Combination of two combinations SNP (a) and SNP (b) Combination of SNP (a) and SNP (b ′) Combination of SNP (b) and SNP (b ′)
- Three combinations SNP (a), SNP ( b) and a combination of SNP (b ′) are preferable, for example.
- the first resistance locus may be defined by, for example, a base sequence including the SNP marker, as shown in (1-2).
- the first resistance locus may be composed of, for example, the base sequence or may include the base sequence.
- the base sequence including the SNP marker is not particularly limited, and examples thereof include the following polynucleotides (a) and (b).
- the polynucleotide of (a) is a nucleotide sequence including the SNP marker of the SNP (a), and the polynucleotide of (b) is at least one of the SNP marker of the SNP (b) and the SNP (b ′). Corresponds to the nucleotide sequence to be included.
- the polynucleotide (a) is a base sequence including the SNP (a), that is, solcap_snp_sl_60160, and is, for example, the following polynucleotide (a1), (a2), or (a3).
- the (a2) and the (a3) are polynucleotides having the same function as the (a1) with respect to the leaf mold resistance at the first resistance locus, respectively.
- (A1) a polynucleotide comprising the base sequence of SEQ ID NO: 1 (a2) in the base sequence other than the 50th base (T) of (a1), one or several bases are deleted, substituted, inserted and / or Polynucleotide comprising the added base sequence (a3) Polynucleotide comprising the base sequence having 80% or more identity to the base sequence other than the 50th base (T) of (a1)
- the underlined 50th base (T) in brackets in SEQ ID NO: 1 is a base corresponding to the polymorphism of the solcap_snp_sl_60160.
- the polynucleotide (a1) can be obtained, for example, from a tomato plant deposited under the deposit number FERM BP-22282 described later.
- the “one or several” is, for example, 1 to 18, 1 to 13, 1 to 10, 1 to 9, 1, 2, 3, or There are four.
- the numerical range of the number of bases and the like discloses, for example, all positive integers belonging to the range. That is, for example, the description “1 to 5” means all disclosures of “1, 2, 3, 4, 5” (hereinafter the same).
- the “identity” is, for example, 80% or more, 85% or more, 89% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more.
- the “identity” can be determined by aligning two base sequences (hereinafter the same).
- the polynucleotide (b) is a nucleotide sequence including at least one of the SNP (b) and the SNP (b ′), that is, at least one of solcap_snp_sl_8658 and solcap_snp_sl_8658 ′.
- the polynucleotide (b) has a base sequence containing the SNP (b) as the SNP marker
- the polynucleotide (b) is, for example, the following (b1), (b2), or (b3) Of the polynucleotide.
- the (b2) and the (b3) are polynucleotides having functions equivalent to the (b1) with respect to the leaf blight resistance at the first resistance locus, respectively.
- (B1) a polynucleotide comprising the base sequence of SEQ ID NO: 2 (b2) in the base sequence other than the 40th base (T) of (b1), one or several bases are deleted, substituted, inserted and / or Polynucleotide comprising the added base sequence (b3) Polynucleotide comprising the base sequence having 80% or more identity to the base sequence other than the 40th base (T) of (b1)
- the underlined 40th base (T) in parentheses in SEQ ID NO: 2 is a base corresponding to the polymorph of solcap_snp_sl_8658.
- the polynucleotide (b1) can be obtained from, for example, a tomato plant deposited under the deposit number FERM BP-22282 described later.
- the “one or several” is, for example, 1 to 15, 1 to 12, 1 to 9, 1 to 8, 1, 2, 3, or 4. It is a piece.
- the “identity” is, for example, 80% or more, 85% or more, 89% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more.
- the polynucleotide (b) has a base sequence containing the SNP (b ′) as the SNP marker
- the polynucleotide (b) is, for example, the following (b1 ′), (b2 ′), or It is a polynucleotide of (b3 ′).
- the (b2 ′) and the (b3 ′) are polynucleotides having functions equivalent to the (b1 ′) with respect to the leaf mold resistance at the first resistance locus, respectively.
- the 22nd base (C) surrounded by a square in SEQ ID NO: 2 is a base corresponding to the polymorphism of the solcap_snp_sl_8658 ′.
- the polynucleotide (b1 ′) can be obtained, for example, from a tomato plant deposited under the deposit number FERM BP-22282 described later.
- polynucleotide (b2 ′) for example, the description of “1 or several” of the polynucleotide (b2) can be used as the “1 or several”.
- the description of “identity” of the polynucleotide (b3) can be cited as the “identity”.
- the polynucleotide (b) has a base sequence containing the SNP (b) and SNP (b ′) as the SNP marker
- the polynucleotide (b) is, for example, the following (b1 ′′)
- the polynucleotide is (b2 ′′) or (b3 ′′).
- the (b2 ′′) and the (b3 ′′) are polynucleotides having functions equivalent to the (b1 ′′) with respect to the leaf mold resistance at the first resistance locus, respectively. .
- (B1 ′′) a polynucleotide comprising the base sequence of SEQ ID NO: 2 (b2 ′′) In the base sequence other than the 22nd base (C) and the 40th base (T) of the (b1 ′′), 1 or Polynucleotide (b3 ′′) consisting of a base sequence in which several bases are deleted, substituted, inserted and / or added (22) base (C) and 40th base (T) of (b1 ′′)
- a polynucleotide comprising a base sequence having 80% or more identity to a base sequence other than
- the underlined 40th base (T) in brackets and the 22nd base (C) in squares in SEQ ID NO: 2 are the solcap_snp_sl_8658 and the solcap_snp_sl_8658, respectively. It is a base corresponding to the polymorphism of '.
- the polynucleotide (b1 ′′) can be obtained, for example, from a tomato plant deposited under the deposit number FERM BP-22282 described later.
- polynucleotide (b2 ′′) for example, the description of “1 or several” of the polynucleotide (b2) can be cited as the “1 or several”.
- the description of “identity” of the polynucleotide (b3) can be cited as the “identity”.
- the number of base sequences including the SNP marker included in the first resistance locus is not particularly limited, and for example, any one or two of the polynucleotides (a) and (b) above It may be.
- a combination of base sequences including the SNP marker is not particularly limited, and examples thereof include the following combinations.
- the first resistance locus is, for example, between the two SNP marker sites. It may be defined by the base sequence of this region.
- the base sequence of the region between the two SNP marker sites is not particularly limited.
- solcap_snp_sl_60250, solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_600089 the base sequence of the region between the two SNP markers selected from the group consisting of solcap_snp_sl_60250, solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_600089.
- the solcap_snp_sl_60250 indicates, for example, the polymorphism of the base in the underlined part enclosed in parentheses in SEQ ID NO: 11.
- the solcap_snp_sl_60250 indicates a polymorphism of G, but the solcap_snp_sl_60250 may be, for example, a base other than G, that is, A, T, or C.
- the base sequence of SEQ ID NO: 11 can be obtained from, for example, a tomato plant deposited under the accession number FERM BP-22282 described later.
- SEQ ID NO: 11 5'-GATGAACTATGGAGAAATACTGTAGTGAGTAAATATTTGAGAATAAGCA [ G ] ATAGATGTAAGATAAAATGAATACATGACAAGA-3 '
- the solcap_snp_sl_60250 can be specified from known information on a database such as the above-mentioned website, for example.
- the base sequence of SEQ ID NO: 12 is, for example, the base sequence of Heinz except for the solcap_snp_sl_60250, and the underlined base surrounded by parentheses is a polymorphism corresponding to the solcap_snp_sl_60250.
- the position of the solcap_snp_sl_60250 can be identified from information on the known Heinz base sequence, for example.
- SEQ ID NO: 12 5'-GATGAACTATGGAGAAATACTGTAGTGAGTAAATATTTGAGAATAAGCA [ G ] ATAGATGTAAGATAAAATGAATACATGACAAGA-3 '
- the solcap_snp_sl — 60089 indicates, for example, the polymorphism of the underlined base surrounded by the parentheses in SEQ ID NO: 13.
- the solcap_snp_sl_60089 indicates a polymorphism that is C, but the solcap_snp_sl_60089 may be a base other than C, that is, A, T, or G, for example.
- the base sequence of SEQ ID NO: 13 can be obtained, for example, from a tomato plant deposited under accession number FERM BP-22282 described later.
- SEQ ID NO: 13 5'-TTTCGTGTTTCGTATTTAGAACTTCTTAATGAAAAT [ C ] CTGGCTCAGCAGCTGGTAGTTAGTAGAGTGATTTGTGGGTGTATAT-3 '
- the solcap_snp_sl — 60089 can be identified from known information on a database such as the aforementioned website.
- the base sequence of SEQ ID NO: 14 is, for example, the base sequence of Heinz except for the solcap_snp_sl_60089, and the underlined base surrounded by parentheses is a polymorphism corresponding to the solcap_snp_sl_60095.
- the position of the solcap_snp_sl_60089 can be specified from, for example, information on the known base sequence of Heinz.
- SEQ ID NO: 14 5'-TTTCGTGTTTCGTATTTAGAACTTCTTAATGAAAAT [ C ] CTGGCTCCGCCGCTGGTAGTTAGTAGAGTGATTTGTGGGTGTATAT-3 '
- the upstream end portion and the downstream end portion can be identified by the two SNP marker sites in the region.
- the region may be, for example, between the two SNP marker sites, and may or may not include both or one of the two SNP marker sites, for example.
- the upstream end and the downstream end of the region become the SNP marker site, but the upstream end and the downstream side.
- the base with the end may be, for example, the underlined base in the above-described base sequence or other bases.
- the two SNP markers that define the region are not particularly limited, and examples thereof include the following combinations.
- the first resistance locus is defined by the base sequence of the region between the two SNP marker sites, the first resistance locus is further added to the region in the base sequence of the region. It is preferable that the SNP marker to be seated is included. Specifically, the first resistance locus preferably includes at least one SNP marker selected from the group consisting of solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 'in the base sequence of the region.
- the SNP marker seated on the region may be, for example, one or both of the two SNP marker sites that define the region on the chromosome, or two SNP markers that define the region. SNP markers that sit between the two sites may also be used.
- the former is also referred to as an SNP marker at the end of the region, and the latter is also referred to as an SNP marker inside the region.
- the SNP marker seated in the region may be, for example, both the SNP marker at the end of the region and the SNP marker inside the region.
- the SNP marker in the region includes, for example, an SNP marker seated between a site of the upstream SNP marker that defines the region and a site of the downstream SNP marker, for example, This can be determined as appropriate based on the sitting position of the SNP marker shown in FIG.
- the number of the SNP markers between the two SNP marker sites may be, for example, one or more. As a specific example, all the SNP markers seated between the SNP marker sites defining the region are used. is there.
- the combination of the base sequence of the region between the sites of the two SNP markers and the SNP marker in the base sequence of the region is not particularly limited, and examples thereof include the following condition (i).
- Condition (i) The nucleotide sequence of the region between the sites of solcap_snp_sl_60250 and solcap_snp_sl_60089 in the chromosome, and
- the base sequence of the region includes at least one SNP marker selected from the group consisting of solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′
- the number of SNP markers in the region is not particularly limited. Three may be sufficient.
- the combination is not particularly limited, and examples thereof include the following combinations. Combination of two combinations SNP (a) and SNP (b) Combination of SNP (a) and SNP (b ′) Combination of SNP (b) and SNP (b ′) Three combinations SNP (a), SNP ( b) and a combination of SNP (b ′)
- the second resistance locus is defined by (2-1) the SNP marker as described above.
- the second resistance locus may be further defined by (2-2) a nucleotide sequence including the SNP marker, or (2-3) a nucleotide sequence of a region between the two SNP marker sites Or may be defined by a combination thereof.
- regulated by the said combination the said combination in particular is not restrict
- the second resistance locus is specified by (2-1), but the second resistance locus is not limited to this.
- the second resistance locus is described in (2-1). Instead, it may be specified by the above (2-2) or (2-3), or may be specified by a combination thereof.
- the second resistance locus may be defined by the SNP marker, for example, as shown in (2-1).
- the SNP marker is not particularly limited, and examples thereof include solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_35221, SG_70, SG_70 ′, and the like. Note that the SNP marker represented by “solcap_snp_sl_number” excluding solcap_snp_sl_25252 ′ can be viewed on the aforementioned website.
- solcap_snp_sl_25252 ′, SG_70, and SG_70 ′ are SNP markers newly identified by the present inventors, and those skilled in the art can use the above-described base sequences containing these SNP markers based on the above-described base sequences.
- the sitting position of the SNP marker can be specified.
- the solcap_snp_sl_25252 (hereinafter also referred to as “SNP (c)”) indicates, for example, a polymorphism in which the underlined base in brackets in SEQ ID NO: 3 is G. That is, for example, when the underlined base is G, the tomato plant is resistant to leaf mold, and when it is a base other than G (for example, A), it indicates that it is susceptible to leaf mold.
- the base sequence of SEQ ID NO: 3 can be obtained from, for example, a tomato plant deposited under the accession number FERM BP-22282 described later.
- the SNP (c) can also be identified from known information on a database such as the above-mentioned website.
- the base sequence of SEQ ID NO: 8 is the base sequence of Heinz except for, for example, the SNP (c) and solcap_snp_sl_25252 ′ described later, and the underlined base surrounded by parentheses is a polymorphism corresponding to the SNP (c) It is. That is, for example, when the underlined base is G, the tomato plant is resistant to leaf mold, and when it is a base other than G (for example, A), it indicates that it is susceptible to leaf mold.
- the position of the SNP (c) can be identified from information on the known Heinz base sequence, for example.
- the solcap_snp_sl_25252 ′ (hereinafter also referred to as “SNP (c ′)”) indicates, for example, a polymorphism in which the base surrounded by a square in SEQ ID NO: 3 is T. That is, for example, if the base surrounded by the square is T, the tomato plant is resistant to leaf mold, and if it is a base other than T (for example, C), it indicates that it is susceptible to leaf mold.
- the SNP (c ′) can also be specified from known information on a database such as the aforementioned website.
- the base sequence of SEQ ID NO: 8 is, for example, a base sequence of Heinz except for the SNP (c) and the SNP (c ′), and the base surrounded by a square is a multiple corresponding to the SNP (c ′). It is a type. That is, for example, if the base surrounded by the square is T, the tomato plant is resistant to leaf mold, and if it is a base other than T (for example, C), it indicates that it is susceptible to leaf mold. In this way, the position of the SNP (c ′) can be specified from, for example, information on the known base sequence of Heinz.
- the solcap_snp_sl_35221 (hereinafter also referred to as “SNP (d)”) indicates, for example, a polymorphism in which the underlined base in brackets in SEQ ID NO: 4 is A. That is, for example, when the underlined base is A, the tomato plant is resistant to leaf mold, and when it is a base other than A (for example, G), it indicates that it is susceptible to leaf mold.
- the base sequence of SEQ ID NO: 4 can be obtained from, for example, a tomato plant deposited under accession number FERM BP-22282, which will be described later.
- SEQ ID NO: 4 5'-TCTAATACCATCTGCAAGTTTCTGAGCCTC [ A ] TCTGACTTCAGCGGACATCTACTTAAG-3 '
- the SNP (d) can also be specified from known information on a database such as the above-mentioned website.
- the base sequence of SEQ ID NO: 9 is, for example, the base sequence of Heinz except for the SNP (d), and the underlined base surrounded by parentheses is a polymorphism corresponding to the SNP (d). That is, for example, when the underlined base is A, the tomato plant is resistant to leaf mold, and when it is a base other than A (for example, G), it indicates that it is susceptible to leaf mold.
- the position of the SNP (d) can be identified from information on the known Heinz base sequence, for example.
- SEQ ID NO: 9 5'-CTCGAAATTGATAAGTTTTCTCTAATACCATCTGCAAGTTTCTGAGCCTC [ A ] TCTGACTTCAGCGGACATCTACTTAAGGTCTGTAAAACAATCAACTTTAT-3 '
- SNP (e) represents a polymorphism in which the underlined base surrounded by parentheses in SEQ ID NO: 5 is A, for example. That is, for example, when the underlined base is A, the tomato plant is resistant to leaf mold, and when it is a base other than A (for example, G), it indicates that it is susceptible to leaf mold.
- the base sequence of SEQ ID NO: 5 can be obtained from, for example, a tomato plant deposited under accession number FERM BP-22282, which will be described later.
- the SNP (e) can also be specified from known information on a database such as the above-mentioned web site, for example.
- the base sequence of SEQ ID NO: 10 is, for example, the base sequence of Heinz except for the SNP (e) and SG_70 ′ described later, and the underlined base surrounded by parentheses is a polymorphism corresponding to the SNP (e) It is. That is, for example, when the underlined base is A, the tomato plant is resistant to leaf mold, and when it is a base other than A (for example, G), it indicates that it is susceptible to leaf mold.
- the position of the SNP (e) can be specified from, for example, information on the known base sequence of Heinz.
- SNP (e') indicates a polymorphism in which the base surrounded by a square in SEQ ID NO: 5 is G, for example. That is, for example, when the base surrounded by the square is G, the tomato plant is resistant to leaf mold, and when it is a base other than G (for example, A), it indicates that it is susceptible to leaf mold.
- the SNP (e ′) can also be specified from known information on a database such as the above-described website.
- the base sequence of SEQ ID NO: 10 is, for example, a base sequence of Heinz except for the SNP (e) and the SNP (e ′), and the base surrounded by a square corresponds to the SNP (e ′). It is a type. That is, for example, when the base surrounded by the square is G, the tomato plant is resistant to leaf mold, and when it is a base other than G (for example, A), it indicates that it is susceptible to leaf mold.
- the position of the SNP (e ′) can be identified from information on the known Heinz base sequence, for example.
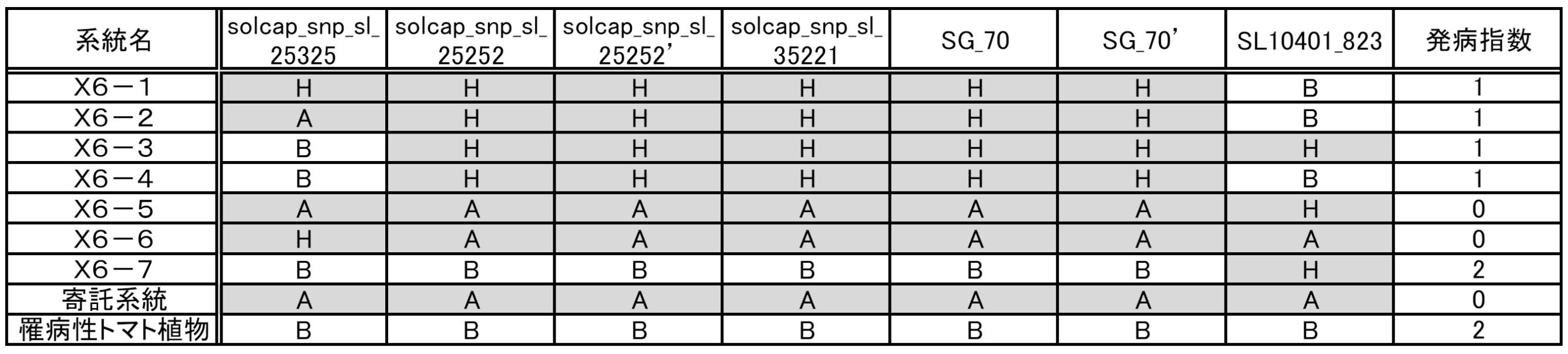
- the sitting position of the SNP marker on the chromosome is not particularly limited. As shown in FIG. 2, for example, on the chromosome 6 of the tomato plant, the SNP marker is solcap_snp_sl_35_25_25_25, solcap_sn_25_25_25, solcap_snp_sl_25_25_, solp_sl_25_25_, solp_sl_25_25_p 'And SL10401_823 are seated in this order.
- solcap_snp_sl_25325 and SL10401_823 are located at the positions of solcap_snp_sl_25252, solcap_snp_sl_25252 ', solcap_snp_sl_3521, SG_70, and SG_70' in the sixth chromosome of the tomato plant. solcap_snp_sl_25325 and SL10401_823 will be described later.
- the number of the SNP markers included in the second resistance locus is not particularly limited. For example, any one or two or more of the SNP markers, that is, 2, 3, 4 Or all five.
- the relationship between these five types of polymorphisms (SNP markers) and leaf mold resistance has not been reported so far, and it was first discovered by the present inventors in leaf mold resistance. It is a new polymorphism involved.
- the combination of the SNP markers is not particularly limited, and examples thereof include the following combinations. Combination of two combinations SNP (c) and SNP (c ′) Combination of SNP (c) and SNP (d) Combination of SNP (c) and SNP (e) Combination of SNP (c) and SNP (e ′) Combination of SNP (c ') and SNP (d) Combination of SNP (c') and SNP (e) Combination of SNP (c ') and SNP (e') Combination of SNP (d) and SNP (e) SNP ( d) Combination of SNP (e ') Combination of SNP (e) and SNP (e') Three combinations SNP (c), SNP (c '), and combination of SNP (d) SNP (c), SNP (C ′) and a combination of SNP (e) SNP (c), a combination of SNP (c ′) and SNP (e ′) SNP (c), a combination of SNP
- the second resistance locus may be defined by a base sequence including the SNP marker, for example, as shown in (2-2).
- the second resistance locus may be composed of, for example, the base sequence or may include the base sequence.
- the base sequence including the SNP marker is not particularly limited, and examples thereof include the following polynucleotides (c), (d) and (e).
- the polynucleotide of (c) is a nucleotide sequence including at least one of the SNP marker of the SNP (c) and the SNP (c ′), and the polynucleotide of (d) is the SNP marker of the SNP (d).
- the nucleotide sequence including the polynucleotide (e) corresponds to a nucleotide sequence including at least one of the SNP marker (e) and the SNP marker (e ′).
- the polynucleotide (c) is a nucleotide sequence including at least one of the SNP (c) and the SNP (c ′), that is, at least one of solcap_snp_sl_25252 and solcap_snp_sl_25252 ′.
- the polynucleotide (c) has a base sequence containing the SNP (c) as the SNP marker, the polynucleotide (c) is, for example, the following (c1), (c2), or (c3) Of the polynucleotide.
- the (c2) and the (c3) are polynucleotides having the same function as the (c1) with respect to the leaf mold resistance at the second resistance locus, respectively.
- (C1) a polynucleotide comprising the base sequence of SEQ ID NO: 3 (c2) In the base sequence other than the 34th base (G) of (c1), one or several bases are deleted, substituted, inserted and / or Polynucleotide comprising the added base sequence (c3) Polynucleotide comprising the base sequence having 80% or more identity to the base sequence other than the 34th base (G) of the above (c1)
- the underlined 34th base (G) in brackets in SEQ ID NO: 3 is a base corresponding to the polymorphism of the solcap_snp_sl_25252.
- the polynucleotide (c1) can be obtained from, for example, a tomato plant deposited under the deposit number FERM BP-22282 described later.
- the “one or several” is, for example, 1 to 14, 1 to 11, 1 to 8, 1 to 4, 1, 2, or 3. is there.
- the “identity” is, for example, 80% or more, 85% or more, 89% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more.
- the polynucleotide (c) has a base sequence containing the SNP (c ′) as the SNP marker
- the polynucleotide (c) is, for example, the following (c1 ′), (c2 ′), or (C3 ′) is a polynucleotide.
- the (c2 ′) and the (c3 ′) are polynucleotides having functions equivalent to the (c1 ′) with respect to the leaf mold resistance at the second resistance locus, respectively.
- the 46th base (T) surrounded by a square in SEQ ID NO: 3 is a base corresponding to the polymorphism of the solcap_snp_sl — 25252 ′.
- the polynucleotide (c1 ′) can be obtained from, for example, a tomato plant deposited under the deposit number FERM BP-22282 described later.
- polynucleotide (c2 ′) for example, the description of “1 or several” of the polynucleotide (c2) can be used as the “1 or several”.
- the description of “identity” of the polynucleotide (c3) can be cited as the “identity”.
- the polynucleotide (c) has a base sequence containing the SNP (c) and SNP (c ′) as the SNP marker
- the polynucleotide (c) is, for example, the following (c1 ′′), (C2 ′′) or (c3 ′′) polynucleotide.
- the (c2 ′′) and the (c3 ′′) are polynucleotides having functions equivalent to the (c1 ′′) with respect to the leaf mold resistance at the second resistance locus, respectively. .
- (C1 ′′) a polynucleotide comprising the base sequence of SEQ ID NO: 3 (c2 ′′) In the base sequence other than the 34th base (G) and the 46th base (T) of the above (c1 ′′), 1 or Polynucleotide (c3 ′′) consisting of a base sequence in which several bases are deleted, substituted, inserted and / or added.
- 34th base (G) and 46th base (T) of (c1 ′′) A polynucleotide comprising a base sequence having 80% or more identity to a base sequence other than
- the underlined 34th base (G) and the 46th base (T) enclosed in squares in SEQ ID NO: 3 are the solcap_snp_sl_25252 and the solcap_snp_sl_25252, respectively. It is a base corresponding to the polymorphism of '.
- the polynucleotide (c1 ′′) can be obtained from, for example, a tomato plant deposited under accession number FERM BP-22282, which will be described later.
- polynucleotide (c2 ′′) for example, the description of “1 or several” of the polynucleotide (c2) can be used as the “1 or several”.
- the description of “identity” of the polynucleotide (c3) can be cited as the “identity”.
- the polynucleotide (d) is a nucleotide sequence including the SNP (d), that is, solcap_snp_sl_35221, and is, for example, the following polynucleotide (d1), (d2), or (d3).
- the (d2) and the (d3) are polynucleotides having functions equivalent to the (d1) with respect to the leaf mold resistance at the second resistance locus, respectively.
- (D1) a polynucleotide comprising the base sequence of SEQ ID NO: 4 (d2) In the base sequence other than the 31st base (A) of (d1), one or several bases are deleted, substituted, inserted and / or Polynucleotide consisting of an added base sequence (d3) Polynucleotide consisting of a base sequence having 80% or more identity to the base sequence other than the 31st base (A) of (d1)
- the underlined 31st base (A) in brackets in SEQ ID NO: 4 is a base corresponding to the polymorphism of the solcap_snp_sl_35221.
- the polynucleotide (d1) can be obtained from, for example, a tomato plant deposited under the deposit number FERM BP-22282 described later.
- the “one or several” is, for example, 1 to 12, 1 to 9, 1 to 8, 1 to 4, 1, 2, or 3. is there.
- the “identity” is, for example, 80% or more, 85% or more, 89% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more.
- the polynucleotide (e) has a base sequence containing at least one of the SNP (e) and the SNP (e ′), that is, at least one of SG_70 and SG_70 ′.
- the polynucleotide (e) is, for example, the following (e1), (e2), or (e3) Of the polynucleotide.
- the (e2) and the (e3) are polynucleotides having the same function as the (e1) with respect to the leaf mold resistance at the second resistance locus, respectively.
- (E1) a polynucleotide comprising the base sequence of SEQ ID NO: 5 (e2) In the base sequence other than the 40th base (A) of (e1), one or several bases are deleted, substituted, inserted and / or Polynucleotide comprising the added base sequence (e3) Polynucleotide comprising the base sequence having 80% or more identity to the base sequence other than the 40th base (A) of (e1)
- the underlined 40th base (A) in parentheses in SEQ ID NO: 5 is the base corresponding to the polymorphism of SG_70.
- the polynucleotide (e1) can be obtained from, for example, a tomato plant deposited under the deposit number FERM BP-22282 described later.
- the “one or several” is, for example, 1 to 16, 1 to 12, 1 to 9, 1 to 8, 1, 2, 3, or There are four.
- the “identity” is, for example, 80% or more, 85% or more, 89% or more, 90% or more, 95% or more, 96% or more, 97% or more, 98% or more, 99% or more.
- the polynucleotide (e) has a base sequence containing the SNP (e ′) as the SNP marker
- the polynucleotide (e) is, for example, the following (e1 ′), (e2 ′), or It is a polynucleotide of (e3 ′).
- the (e2 ′) and the (e3 ′) are polynucleotides having functions equivalent to the (e1 ′) with respect to the leaf mold resistance at the second resistance locus, respectively.
- the 75th base (G) surrounded by a square in SEQ ID NO: 5 is a base corresponding to the polymorphism of SG_70 ′.
- the polynucleotide (e1 ′) can be obtained from, for example, a tomato plant deposited under the accession number FERM BP-22282 described later.
- polynucleotide (e2 ′) for example, the description of “1 or several” of the polynucleotide (e2) can be cited as the “1 or several”.
- the description of “identity” of the polynucleotide (e3) can be cited as the “identity”.
- the polynucleotide (e) has a base sequence containing the SNP (e) and SNP (e ′) as the SNP marker
- the polynucleotide (e) is, for example, the following (e1 ′′), (E2 ′′) or (e3 ′′) polynucleotide.
- the (e2 ′′) and the (e3 ′′) are polynucleotides having functions equivalent to the (e1 ′′) with respect to the leaf mold resistance at the second resistance locus, respectively. .
- (E1 ′′) a polynucleotide comprising the base sequence of SEQ ID NO: 5 (e2 ′′) In the base sequence other than the 40th base (A) and the 75th base (G) of the (e1 ′′), 1 or Polynucleotide (e3 ′′) consisting of a base sequence in which several bases are deleted, substituted, inserted and / or added.
- 40th base (A) and 75th base (G) of (e1 ′′) A polynucleotide comprising a base sequence having 80% or more identity to a base sequence other than
- the underlined 40th base (A) in brackets and the 75th base (G) in squares in SEQ ID NO: 5 are the SG_70 and SG_70, respectively. It is a base corresponding to the polymorphism of '.
- the polynucleotide (e1 ′′) can be obtained, for example, from a tomato plant deposited under the accession number FERM BP-22282 described later.
- polynucleotide (e2 ′′) for example, the description of “1 or several” of the polynucleotide (e2) can be used as the “1 or several”.
- the description of “identity” of the polynucleotide (e3) can be cited as the “identity”.
- the number of base sequences including the SNP marker included in the second resistance locus is not particularly limited.
- any one of the polynucleotides (c), (d), and (e) is used. Or two or more, ie two or all three.
- a combination of base sequences including the SNP marker is not particularly limited, and examples thereof include the following combinations. Two combinations of polynucleotide (c) and (d) polynucleotide combination (c) and (e) polynucleotide combination (d) and (e) polynucleotide combination 3 Combinations of (c) polynucleotides, (d) polynucleotides, and (e) polynucleotide combinations
- the second resistance locus is, for example, between the two SNP marker sites. It may be defined by the base sequence of this region.
- the base sequence of the region between the two SNP marker sites is not particularly limited. Examples include a base sequence of a region between two SNP marker sites, a base sequence of a region between two SNP marker sites selected from the group consisting of solcap_snp_sl_25325, solcap_snp_sl_25252, solcap_snp_sl_35221, SG_70, and SL10401_823.
- the solcap_snp_sl_25325 indicates, for example, the polymorphism of the underlined base surrounded by the parentheses in SEQ ID NO: 15.
- the solcap_snp_sl_25325 indicates a polymorphism of G, but the solcap_snp_sl_25325 may be, for example, a base other than G, that is, A, T, or C.
- the base sequence of SEQ ID NO: 15 can be obtained from, for example, a tomato plant deposited under the accession number FERM BP-22282 described later.
- SEQ ID NO: 15 5'-TCACGAGCAGCAAGAAAGCTATTATCGCTAGTATCCAATA [ G ] TACGAAAGAGAACACTCCATCCAACATGT-3 '
- the solcap_snp_sl_25325 can be specified from, for example, known information on a database such as the aforementioned website.
- the base sequence of SEQ ID NO: 16 is, for example, the base sequence of Heinz except for the solcap_snp_sl_25325, and the underlined base surrounded by parentheses is a polymorphism corresponding to the solcap_snp_sl_25325.
- the position of the solcap_snp_sl_25325 can be identified from, for example, information on the known base sequence of Heinz.
- the SL10401_823 indicates, for example, the polymorphism of the base in the underlined part enclosed in parentheses in SEQ ID NO: 17.
- SL10401_823 indicates a polymorphism that is T, but SL10401_823 may be, for example, a base other than T, that is, A, G, or C.
- the base sequence of SEQ ID NO: 17 can be obtained from, for example, a tomato plant deposited under the accession number FERM BP-22282 described later.
- SEQ ID NO: 17 5'-TACATTCTGTTTTATTTGTTCCTTACTTGTCT [ T ] AGTTGGGCTCATATCTCATTTTGAATGTAAATTACAGGTGCAAGACAA-3 '
- the SL 10401_823 can also be specified from known information on a database such as the above-described website.
- the base sequence of SEQ ID NO: 18 is, for example, the base sequence of Heinz except for the SL10401_823, and the underlined base surrounded by parentheses is a polymorphism corresponding to the SL10401_823.
- the position of the SL 10401_823 can be specified from, for example, information on a known Heinz base sequence.
- SEQ ID NO: 18 5'-TACATTCTGTTTTATTTGTTCCTTACTTGTCT [ T ] AGTTGGGCTCATATCTCATTTTGAATGTAAATTACAGGTGCAAGACAA-3 '
- the upstream end portion and the downstream end portion can be identified by the two SNP marker sites in the region.
- the region may be, for example, between the two SNP marker sites, and may or may not include both or one of the two SNP marker sites, for example.
- the upstream end and the downstream end of the region become the SNP marker site, but the upstream end and the downstream side.
- the base with the end may be, for example, the underlined base in the above-described base sequence or other bases.
- the two SNP markers that define the region are not particularly limited, and examples thereof include the following combinations.
- the second resistance locus is defined by the base sequence of the region between the two SNP marker sites
- the second resistance locus is further added to the region in the base sequence of the region. It is preferable that the SNP marker to be seated is included.
- the second resistance locus includes, for example, at least one SNP selected from the group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_35221, SG_70, and SG_70 ′ in the base sequence of the region. It is preferable.
- the SNP marker seated on the region may be, for example, one or both of the two SNP marker sites that define the region on the chromosome, or two SNP markers that define the region. SNP markers that sit between the two sites may also be used.
- the former is also referred to as an SNP marker at the end of the region, and the latter is also referred to as an SNP marker inside the region.
- the SNP marker seated in the region may be, for example, both the SNP marker at the end of the region and the SNP marker inside the region.
- the SNP marker in the region includes, for example, an SNP marker seated between a site of the upstream SNP marker that defines the region and a site of the downstream SNP marker, for example, This can be determined as appropriate based on the sitting position of the SNP marker shown in FIG.
- the number of the SNP markers between the two SNP marker sites may be, for example, one or more. As a specific example, all the SNP markers seated between the SNP marker sites defining the region are used. is there.
- the combination of the base sequence of the region between the sites of the two SNP markers and the SNP marker in the base sequence of the region is not particularly limited, and examples thereof include the following condition (ii).
- Condition (ii) The base sequence of the region between the sites of solcap_snp_sl_25325 and SL10401_823 in the chromosome, and The base sequence of the region includes at least one SNP marker selected from the group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_35221, SG_70, and SG_70 ′.
- the number of SNP markers in the region is not particularly limited. That is, it may be 2, 3, 4, or 5.
- the combination is not particularly limited, and examples thereof include the following combinations. Combination of two combinations SNP (c) and SNP (c ′) Combination of SNP (c) and SNP (d) Combination of SNP (c) and SNP (e) Combination of SNP (c) and SNP (e ′) Combination of SNP (c ') and SNP (d) Combination of SNP (c') and SNP (e) Combination of SNP (c ') and SNP (e') Combination of SNP (d) and SNP (e) SNP ( d) Combination of SNP (e ') Combination of SNP (e) and SNP (e') Three combinations SNP (c), SNP (c '), and combination of SNP (d) SNP (c), SNP (C ′) and a combination of SNP (e
- leaf mold resistance can be imparted to a tomato plant.
- the degree of leaf mold resistance of a tomato plant can be represented by, for example, a disease index by referring to the method described in Reference Information 4 below.
- a disease index For the calculation of the disease index by this method, the explanation of Example 1 described later can be used.
- the disease index 1 or less can be set as disease resistance
- the disease index 2 or more can be set as susceptibility.
- Reference information 4 Nanshin trial disease pest soil fertilizer department, “Eco-shot is effective for tomato leaf mold control of tomatoes”, [online], Nagano Prefecture, [March 3, 2015 search], Internet ⁇ URL : http://www.pref.nagano.lg.jp/nogi/sangyo/nogyo/gijutsu/fukyugijutsu/200802/200802fukyu.html>
- the resistance marker of the present invention may further include another resistance marker, for example.
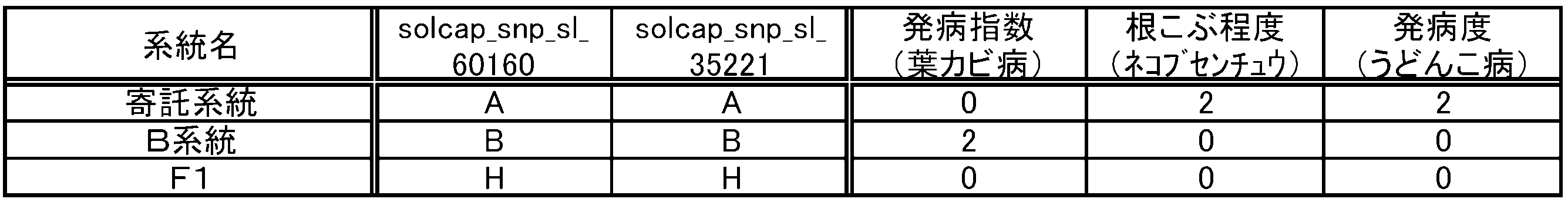
- Examples of the other resistance include root-knot nematode resistance and powdery mildew resistance.
- the leaf mold resistant tomato plant of the present invention comprises the leaf mold resistance locus on chromosome 1 and the leaf mold resistance locus on chromosome 6. At least one leaf mold resistance locus, wherein the leaf mold resistance locus on the first chromosome is identified by at least one SNP marker selected from the group consisting of solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′ , Wherein the fungal resistance locus on chromosome 6 is at least one selected from the group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_35221, SG_70, and SG_70 ′.
- the leaf mold resistant tomato plant of the present invention comprises at least one leaf mold resistant locus on chromosome 1 and a leaf mold resistant locus on chromosome 6.
- the leaf mold resistance locus on the first chromosome is identified by at least one SNP marker selected from the group consisting of solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′, and the leaf mold resistance on the sixth chromosome
- the locus is identified by at least one SNP marker selected from the group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_35221, SG_70, and SG_70 ′.
- the leaf mold resistant tomato plant of the present invention includes the resistance marker of the present invention including at least one of the first resistance locus and the second resistance locus.
- the description of the resistance marker of the invention can be incorporated.
- the leaf mold resistance locus on the first chromosome and the leaf mold resistance locus on the sixth chromosome are, for example, the first resistance gene in the resistance marker of the present invention, respectively. It can be read as a locus and a second resistance locus.
- the description of the resistance marker of the present invention can be used for the leaf mold resistant tomato plant of the present invention.
- the leaf mold resistant tomato plant of the present invention exhibits resistance to leaf mold.
- the leaf mold resistance is at least one of the first resistance locus on the first chromosome and the second resistance locus on the sixth chromosome. Brought about by.
- the leaf mold resistant tomato plant of the present invention comprises at least one of the first resistance locus and the second resistance locus, and the first resistance locus is located on the first chromosome, A second resistance locus is contained on chromosome 6.
- the leaf mold resistant tomato plant of the present invention contains the first resistance locus
- the leaf mold resistant tomato plant of the present invention is, for example, in place of the first chromosome, except for the first chromosome.
- the first resistance locus may be included on any chromosome.
- the leaf mold resistant tomato plant of the present invention has, for example, the second chromosome, the third chromosome, the fourth chromosome, the fifth chromosome, the sixth chromosome, the seventh chromosome, the eighth chromosome, the ninth chromosome, the tenth chromosome.
- the first resistance locus may be included on any chromosome of chromosome, chromosome 11 and chromosome 12.
- the leaf mold resistant tomato plant of the present invention contains the second resistance locus
- the leaf mold resistant tomato plant of the present invention is replaced with, for example, a chromosome 6 other than the 6th chromosome.
- the second leaf mold resistance locus may be included on any chromosome.
- the leaf mold resistant tomato plant of the present invention has, for example, the first chromosome, the second chromosome, the third chromosome, the fourth chromosome, the fifth chromosome, the seventh chromosome, the eighth chromosome, the ninth chromosome, the tenth chromosome.
- the second resistance locus may be included on any chromosome of chromosome, chromosome 11 and chromosome 12.
- the leaf mold resistant tomato plant of the present invention includes the first resistance locus and the second resistance locus
- the tomato plant of the present invention has the first resistance on the same chromosome.
- a sex locus and the second resistance locus may be included, or the first resistance locus and the second resistance locus may be included on different chromosomes.
- each of the first resistance locus and the second resistance locus is, for example, the first resistance gene in the resistance marker of the present invention.
- the description of the locus and the second resistance locus can be incorporated.
- the leaf mold resistant tomato plant of the present invention may contain only the first resistance locus or only the second resistance locus.
- the tomato plant containing the resistance marker is improved in leaf mold resistance by including both the first resistance locus and the second resistance locus, for example. .
- the tomato plant in which each of the first resistance locus and the second resistance locus is a heterozygous type of resistance and susceptibility is, for example, the first resistance locus. Shows higher leaf mold resistance than tomato plants that are resistant homozygous, and the second resistance locus is resistant to leaf mold equivalent to tomato plants that are resistant homozygous. Indicates.
- the leaf mold resistant tomato plant of the present invention preferably comprises the leaf mold resistant locus and the sixth leaf mold resistant locus on the first chromosome.
- a tomato plant deposited under the deposit number FERM BP-22282 S. lycopersicum ) or its progeny line can be mentioned.
- the deposited tomato plant includes, for example, the first resistance locus on the first chromosome and the second resistance locus on the sixth chromosome.
- the deposit information is shown below.
- Deposit Type International Depositary Agency Name: National Institute of Technology and Evaluation Patent Biological Depositary Center Address: Japan 2-5-8 Kazusa Kamashichi, Kisarazu City, Chiba Prefecture 292-0818 Japan Accession Number: FERM BP- 22282 Display for identification: Takii7 Date of receipt: January 28, 2015
- the leaf mold resistant tomato plant of the present invention can also be produced, for example, by introducing at least one of the first resistance locus and the second resistance locus into a tomato plant.
- a method for introducing the leaf mold resistance locus into the tomato plant is not particularly limited, and examples thereof include conventionally known genetic engineering techniques.
- the leaf mold resistance locus to be introduced can be exemplified by the above-mentioned leaf mold resistance locus.
- the leaf mold resistant tomato plant of the present invention is not particularly limited in characteristics other than leaf mold resistance, such as trait and ecological characteristics.
- the leaf mold resistant tomato plant of the present invention may further have other resistance.
- examples of the other resistance include root-knot nematode resistance and powdery mildew resistance.
- the “plant body” may mean either a plant individual indicating the whole plant or a part of the plant individual.
- the plant individual part include organs, tissues, cells, vegetative propagation bodies, and the like.
- the organ include petals, corolla, flowers, leaves, seeds, fruits, stems, roots and the like.
- the tissue is, for example, a part of the organ.
- the plant part may be, for example, one kind of organ, tissue and / or cell, or two or more kinds of organ, tissue and / or cell.
- production method a method for producing a leaf mold resistant tomato plant of the present invention.
- the following methods are examples, and the present invention is not limited to these methods.
- the production method can also be referred to as a growth method, for example.
- the leaf mold resistance locus can be rephrased as the resistance marker of the present invention.
- the method for producing a leaf mold resistant tomato plant of the present invention includes the following steps (a) and (b).
- (b) A leaf from (one or more) tomato plant or progeny line obtained from step (a) Process of selecting tomato plants with mold resistance
- the production method of the present invention is characterized by using the leaf mold tomato plant of the present invention as a parent, and other processes and conditions are not limited at all.
- the description of the resistance marker of the present invention can be used in the production method of the present invention.
- the leaf mold resistant tomato plant used as the first parent may be the leaf mold resistant tomato plant of the present invention.
- the leaf mold resistant tomato plant is preferably, for example, a tomato plant deposited under the accession number FERM BP-22282 as described above or its progeny line.
- the leaf mold resistant tomato plant used as the first parent can be obtained, for example, by the screening method of the present invention described later. Therefore, for example, prior to the step (a), the leaf mold resistant tomato plant is obtained from, for example, a test tomato plant (hereinafter also referred to as “candidate tomato plant”) by the following step (x). You may select and prepare.
- (X) A step of selecting the leaf mold resistant tomato plant of the present invention from (one or more) test tomato plants
- step (x) selection of the leaf mold resistant tomato plant can be said to be selection of a tomato plant containing the leaf mold resistant locus.
- the said (x) process can be performed by the following (x1) process and (x2) process, for example.
- (X1) a detection step of detecting the presence or absence of the leaf mold resistance locus on the chromosome of the (one or more) test tomato plants
- (x2) Selection process for selecting the tomato plant as a mold resistant tomato plant
- the selection in the step (x) is, for example, selection of a tomato plant including the leaf mold resistance locus, and specifically, the leaf mold disease of the tomato plant to be tested is selected. By detecting the resistance locus, the leaf mold resistant tomato plant can be selected.
- the leaf mold resistance locus is detected by, for example, defining the first resistance locus as described in the resistance marker of the present invention, (1-1) the SNP marker, (1 -2) defining the base sequence containing the SNP marker, (1-3) the base sequence of the region between the two SNP marker sites, and combinations thereof, and the second resistance locus (2) It can be detected using: -1) the SNP marker, (2-2) the base sequence containing the SNP marker, (2-3) the base sequence of the region between the two SNP marker sites, and combinations thereof.
- the selection in the step (x) includes, for example, a leaf mold resistant tomato containing at least one of the leaf mold resistance locus on the first chromosome and the leaf mold resistance locus on the sixth chromosome.
- the leaf mold resistance locus is at least one SNP selected from the group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ', solcap_snp_sl_35221, SG_70, and SG_70'. It is specified by the car.
- the selection in the step (x) may be, for example, selection of a leaf mold resistant tomato plant containing only the leaf mold resistance locus on the first chromosome, or leaf mold resistance on the sixth chromosome. It may be a selection of a leaf mold resistant tomato plant containing only the sex locus.
- the tomato plant containing the resistance marker is improved in leaf mold resistance by including both the first resistance locus and the second resistance locus, for example. .
- the tomato plant in which each of the first resistance locus and the second resistance locus is a heterozygous type of resistance and susceptibility is, for example, the first resistance locus.
- the selection in the step (x) is selection of a leaf mold resistant tomato plant comprising the leaf mold resistance gene on the first chromosome and the leaf mold resistance gene on the sixth chromosome. It is preferable.
- the first resistance locus is (1 -1) Selected by the SNP marker.
- the first resistance locus may be further selected by (1-2) a nucleotide sequence including the SNP marker, or (1-3) a nucleotide sequence in a region between two SNP marker sites. Or may be selected by a combination thereof.
- the said combination is not restrict
- the first resistance locus is selected by (1-1), but the first resistance locus is not limited to this. For example, in (1-1), Instead, it may be selected in (1-2) or (1-3), or may be selected in combination of (1-2) and (1-3). Combination of (1-1) and (1-2) Combination of (1-1) and (1-3) (1-1), (1-2) and (1-3) Combination
- the SNP marker to be selected in the first resistance locus is not particularly limited.
- “(1-1) SNP marker in the resistance marker of the present invention” The description of “specification” can be used.
- the selection in the step (x) is performed by, for example, a tomato plant including a leaf mold resistance locus on chromosome 1 specified by solcap_snp_sl_60160 and solcap_snp_sl_8658, preferably solcap_snp_sl_86_, solcap_sp86_ Selection of tomato plants containing the leaf mold resistance locus on chromosome 1 specified in
- the selection in the step (x) is, for example, selection of a leaf mold resistant tomato plant containing a leaf mold resistance locus on the first chromosome.
- the leaf mold resistance gene on the first chromosome is identified by at least one of the polynucleotides (a) and (b).
- the description of “(1-2) Identification by nucleotide sequence including SNP marker” in the resistance marker of the present invention can be used.
- the selection in the step (x) is, for example, selection of a tomato plant containing a leaf mold resistance locus on the first chromosome specified by the polynucleotides (a) and (b). can give.
- the selection in the step (x) is, for example, leaf mold resistance containing a leaf mold resistance locus on the first chromosome
- the fungal resistance locus on the first chromosome is selected from a group of solcap_snp_sl_60250, solcap_snp_sl_60160, solcap_snp_sl_8658, solcap_snp_sl_8658 ′, Includes the base sequence of the region between the sites.
- solcap_snp_sl_60250, solcap_snp_sl_60160, solcap_snp_sl_8658, solcap_snp_sl_8658 ′ Includes the base sequence of the region between the sites.
- the description of “(1-3) Identification by the base sequence of the region between the two SNP marker sites” in the resistance marker of the present invention is used. it can.
- the selection in the step (x) further includes, for example, on the first chromosome including at least one SNP marker selected from the group consisting of solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′ in the base sequence of the region. Selection of tomato plants having a leaf mold resistance locus.
- the selection in the step (x) may be, for example, selection of a tomato plant containing a leaf mold resistance locus on the first chromosome that satisfies the condition (i).
- the chromosome for detecting the presence or absence of the first resistance locus is preferably the first chromosome.
- the second resistance locus is (2-1) the SNP as described above. Selected by marker.
- the second resistance locus may be further selected from (2-2) a base sequence containing the SNP marker, or (2-3) a base sequence in a region between two SNP marker sites. Or may be selected by a combination thereof.
- the said combination is not restrict
- the second resistance locus is selected by (2-1), but the second resistance locus is not limited to this.
- the second resistance locus is described in (2-1). Instead, it may be selected in the above (2-2) or (2-3), or may be selected in combination thereof.
- the SNP marker to be selected in the second resistance locus is not particularly limited.
- “(2-1) SNP marker in the resistance marker of the present invention” The description of “specification” can be used.
- the selection in the step (x) includes, for example, solcap_snp_sl_25252, solcap_snp_sl_3521, and a tomato plant including the second resistance locus identified by SG_70, preferably solcap_snp_sl_25_25, solcap_sl_25_25, And selection of tomato plants containing the second resistance locus identified by SG_70 ′.
- the selection in the step (x) is, for example, selection of a leaf mold resistant tomato plant containing a leaf mold resistance locus on the chromosome 6. And the leaf mold resistance locus on the sixth chromosome is identified by at least one selected from the group consisting of (c), (d) and (e).
- the description of “(2-2) Identification by nucleotide sequence containing SNP marker” in the resistance marker of the present invention can be used.
- the selection in the step (x) includes, for example, a leaf mold resistance locus on the chromosome 6 specified by the polynucleotides (c), (d) and (e). Selection of tomato plants.
- the selection in the step (x) is, for example, leaf mold resistance containing a leaf mold resistance locus on the chromosome 6
- the tomato plant resistance gene on the sixth chromosome is selected from solcap_snp_sl_25325, solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_Sl_3521SG, s________________________________ 104_ It includes the base sequence of the region between the sites of one SNP marker.
- the description of “(2-3) Identification by the base sequence of the region between the two SNP marker sites” in the resistance marker of the present invention is used. it can.
- the selection in the step (x) further includes, for example, at least one SN marker selected from the group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_35221, SG_70, and SG_70 ′ in the base sequence of the region.
- SN marker selected from the group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_35221, SG_70, and SG_70 ′ in the base sequence of the region.
- examples include selection of tomato plants containing the fungal resistance locus on the sixth chromosome.
- the selection in the step (x) may be, for example, selection of a tomato plant containing a leaf mold resistance locus on the sixth chromosome that satisfies the condition (ii).
- the chromosome for detecting the presence or absence of the second resistance locus is preferably the sixth chromosome.
- the tomato plant used as the other parent is not particularly limited, and may be, for example, a tomato plant containing a known leaf mold resistance gene or other tomato plant having resistance.
- the leaf mold resistant tomato plant of the present invention may be used.
- the other tomato plants having resistance include root-knot nematode resistant tomato plants, powdery mildew resistant tomato plants, and the like.
- a method for crossing the leaf mold resistant tomato plant with the other tomato plant is not particularly limited, and a known method can be adopted.
- the target for selecting the leaf mold resistant tomato plant may be, for example, the tomato plant obtained from the step (a) or a progeny line obtained from the tomato plant. .
- the target may be, for example, an F1 tomato plant obtained by crossing in step (a) or a progeny line.
- the progeny line may be, for example, an inbred progeny or backcross progeny of the F1 tomato plant obtained by crossing in the step (a), or crossing the tomato plant of F1 with another tomato plant. It may be a tomato plant obtained by
- selection of a leaf mold resistant tomato plant can be performed, for example, by confirming the leaf mold resistance directly or indirectly.
- the direct confirmation can be carried out by evaluating, for example, the leaf mold resistance of the obtained F1 tomato plant or its progeny line by the disease index as described above.
- the F1 tomato plant or its progeny can be confirmed by inoculating, for example, a leaf mold and evaluating the leaf mold resistance using the disease index.
- the F1 tomato plant or its progeny showing a disease severity of 1 or less can be selected as a leaf mold resistant tomato plant.
- the selection by the indirect confirmation can be performed by, for example, the following steps (b1) and (b2).
- B1) A detection step for detecting the presence or absence of a leaf mold resistance locus on the chromosome of (one or more) tomato plants or progeny lines obtained from the step (a)
- (b2) the leaf mold disease A selection step of selecting the (one or more) tomato plant or its progeny line obtained by the step (a) as a leaf mold resistant tomato plant due to the presence of the resistance locus.
- Selection of the leaf mold resistant tomato plant in the step (b) is, for example, the same as the method described in the step (x), and more specific by detecting the presence or absence of the leaf mold resistant locus. Can be performed by detecting the presence or absence of the leaf mold resistance locus using the molecular marker.
- the tomato plant or its progeny line in which the leaf mold resistance was confirmed can be selected as a leaf mold resistant tomato plant.
- the production method of the present invention may further include a seeding step of collecting seeds from the progeny line obtained by crossing.
- Screening method for leaf mold resistant tomato plant The method for screening leaf mold resistant tomato plant of the present invention (hereinafter also referred to as “screening method”) is for producing a leaf mold resistant tomato plant by crossing. As a parent, from one or more test tomato plants, a leaf mold resistance locus on chromosome 1 and a leaf mold resistance locus on chromosome 6 as a leaf mold resistance marker for tomato plants.
- At least one selected from the group consisting of solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′ comprising selecting a leafy mildew resistant tomato plant comprising at least one Chromosome 6 identified by SNP marker Leaf blight resistance locus of, solcap_snp_sl_25252, solcap_snp_sl_25252 characterized in that it is identified by at least one SNP marker selected from the group consisting of ', solcap_snp_sl_35221, SG_70, and SG_70'.
- the screening method of the present invention comprises at least one of a leaf mold resistance locus on chromosome 1 and a leaf mold resistance locus on chromosome 6 as a leaf mold resistance marker from a tomato plant to be tested.
- the fungal resistance locus on chromosome 6 is a group consisting of solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl — 35221, SG_70, and SG_70 ′.
- Characterized in that it is identified by at least one SNP marker selected other steps and conditions are not particularly limited.
- a parent with resistance to leaf mold can be obtained by the resistance marker of the present invention.
- the description of the resistance marker of the present invention can be used.
- step (x) in the method for producing a leaf mold resistant tomato plant of the present invention can be cited.
- Example 1 Analysis of the inheritance pattern of the leaf mold resistance locus, identification of the leaf mold resistance locus, and the leaf mold resistance locus and leaf mold resistance of a novel leaf mold resistant tomato plant
- the leaf mold resistance locus is a marker for leaf mold resistance in tomato plants
- the tomato plant containing the resistance locus is a leaf mold resistant tomato plant.
- the tomato plant resistant to tomato plants can be screened using the tomato plant resistance marker.
- Leaf mold resistance Tomato plant (Accession No. FERM BP-22282) of the deposited line and leaf mold disease-susceptible tomato plant “Momotaro (Takii Seedling Co., Ltd., S. lycopersicum )” (hereinafter referred to as “susceptibility”)
- 114 F2 segregated populations (hereinafter also referred to as “114 lines”) were produced by crossing with “tomato plants”.
- the inbred progeny F3 was produced by self-breeding each of the 114 lines. 9 to 15 individuals of F3 of each line were used, and an inoculation test for leaf mold was performed as shown below.
- the deposited line includes solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8_s_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25
- the diseased tomato plant includes solcap_snp_sl_60_60, solcap_snp_sl_86_, and solcap_snp_sl_25_25_25_25_25_25_25_25_25_25_25 in the first chromosome, and solcap_s1_s_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25_25
- the strain distributed from Wageningen University (leaf mold 1) was used.
- the fungal strain 1 was inoculated on a V8 agar medium containing V8 (registered trademark) vegetable juice (manufactured by Campbell Soup Company). After the inoculation, the cells were cultured at 20 ° C. for 2 weeks.
- the conidia of the above-mentioned leaf mold fungus strain 1 were recovered from the culture medium using sterilized water. Further, the conidia were diluted with sterilized water so as to be 1 ⁇ 10 5 cells / mL, and a conidia spore suspension was prepared and used as an inoculation source.
- the inoculation test a tomato plant in which two true leaves were developed was used.
- the tomato plant was inoculated with the fungal strain 1 by spraying 1 mL of the conidial spore suspension per individual tomato plant.
- the tomato plants after inoculation were grown for 2 weeks under the conditions of 20 ° C. and 12 hours of day length.
- the disease investigation was conducted as follows.
- the leaf mold fungus strain 1 is a leaf mold fungus capable of infecting tomato plants containing Cf (resistance gene to Cladosporium fulvum) 2, 4, 5, 9, and 11. It is known that
- Disease index 0 No sporulation observed (high resistance)
- Disease index 1 Sporulation is observed, but the lesion area is less than 50% of the leaf area (resistance)
- Disease index 2 Sporulation is observed, and the lesion area is 50% or more of the leaf area (susceptibility)
- Disease index [(0 ⁇ n 0 ) + (1 ⁇ n 1 ) + (2 ⁇ n 2 )] / investigated number of individuals
- “0, 1, 2” indicates the disease index
- “n” “0 , n 1 , n 2 ” indicates the number of individuals with disease index 0, disease index 1, and disease index 2, respectively.
- the region on the first chromosome was a region containing SNP specified by solcap_snp_sl_60160.
- the region on the sixth chromosome was a region containing SNP specified by SG_70. From this result, it was clarified that the novel leaf mold resistance gene locus sits on chromosome 1 and chromosome 6.
- solcap_snp_sl_60160 on the first chromosome is a resistant homozygous type, heterozygous type, or susceptible homozygous type.
- 5 individuals were selected, each containing SG_70 on chromosome 6 in a resistant homozygous, heterozygous, or diseased homozygous type.
- the disease index was determined. Asked. The disease index was similarly determined except that the deposited line or the susceptible tomato plant was used in place of the selected tomato plant.
- the leaf mold fungus strain 2 is a leaf mold fungus collected from a tomato plant that spontaneously developed leaf mold disease in a tomato cultivation field in Iwate Prefecture.
- the fungal strain 2 of the leaf mold can be obtained by conducting an inoculation test similar to the above (2) on tomato plants containing Cf-2, 4, 5, 9, and 11, respectively. It has been confirmed that tomato plants containing 9 can be infected.
- stock 2 can be obtained from the Kumamoto Prefecture pest control station, for example.
- the leaf mold fungus strain 3 is a leaf mold fungus collected from a tomato plant that spontaneously developed leaf mold in a tomato cultivation field in Fukushima Prefecture.
- the fungal strain 3 of the leaf mold can be obtained by carrying out an inoculation test similar to the above (2) on tomato plants containing Cf-2, 4, 5, 9, and 11, respectively. It has been confirmed that tomato plants containing 9 can be infected.
- the leaf mold fungus exhibiting the same infectivity as the leaf mold strain 3 can be obtained from, for example, the Vegetable Tea Research Institute.
- the leaf mold fungus strain 4 is a leaf mold fungus obtained from Wageningen University, and is also known to be a leaf mold fungus capable of infecting tomato plants containing Cf-2, 4 and 5. .
- the results of the deposited line and the diseased tomato plant are shown in Table 1.
- A indicates that the SNP marker is included in a resistant homozygous form
- B indicates that the SNP marker is included in a susceptible homozygous form (hereinafter the same).
- the susceptible tomato plant comprises solcap_snp_sl_60160 on the first chromosome in a susceptible homozygous form and SG_70 on the sixth chromosome in a susceptible homozygous form. It was.
- the susceptible tomato plant had a disease index of 2 and was susceptible to leaf mold against any leaf mold.
- the deposited line contained solcap_snp_sl_60160 on the first chromosome in a resistant homozygous form and SG_70 on the sixth chromosome in a resistant homozygous form.
- the deposited strain had a disease index of 0 and showed high resistance to leaf mold against any leaf mold. From these results, it is found that the resistance locus specified by solcap_snp_sl_60160 on the first chromosome and the resistance locus specified by SG_70 on the sixth chromosome are responsible for leaf mold resistance. It could be confirmed.
- the resistance locus specified by solcap_snp_sl_60160 on the first chromosome and the resistance locus specified by SG_70 on the sixth chromosome are responsible for leaf mold resistance, It was found that the resistance locus serves as a leaf mold resistance marker for tomato plants, and that a leaf mold resistant tomato plant can be screened using the tomato plant leaf mold resistance marker. Furthermore, since each of the fungal strains 1 to 4 is a fungal fungus capable of infecting a tomato plant containing a known fungal resistance gene, the first resistance locus and the The second resistance locus has also been shown to be effective against fungal fungi that are capable of infecting tomato plants that contain a known fungal resistance gene.
- Table 2 shows the results of inoculating the tomato plants of different genotypes with the mold fungus strain 1.
- H indicates that the SNP marker is included in a heterozygous form (the same applies hereinafter).
- solcap_snp_sl_60160 is contained in a resistant homozygous, heterozygous, or susceptible homozygous type and SG_70 in a susceptible homozygous type for the fungal strain 1
- solcap_snp_sl_60160 was resistant to homozygous and heterozygous forms
- solcap_snp_sl_60160 was resistant to homozygous and heterozygous forms
- the tomato plants included in the mold only some grayish white spots were observed on the leaves.
- the disease index is 1, 1, 2 respectively.
- the disease index is 1, 1, 2 respectively.
- the disease index is 1, 1, 2 respectively.
- a tomato plant containing solcap_snp_sl_60160 in a susceptible homozygous form and SG_70 in a resistant homozygous, heterozygous form, or a susceptible homozygous form has a disease index of 0 and is high against leaf mold Resistant.
- Leaf Blight Resistance Locus on Chromosome 1 From the 114 strain, solcap_snp_sl_60160 on the first chromosome is heterozygous, and SG_70 on the sixth chromosome is susceptible homozygous. Individuals to be included were selected (first F2 selection line). Next, the first progeny progeny F3 was produced by selfing each of the first F2 selected lines.
- the for first inbred posterity F3 the same method as the (3), the SNP assay, Solcap_snp_sl_60160, and Solcap_snp_sl_60160 a SNP marker near solcap_snp_sl_60303, solcap_snp_sl_60250, solcap_snp_sl_8658, solcap_snp_sl_8658 ', solcap_snp_sl_60089, and solcap_snp_sl_60043 A polymorphic base corresponding to was identified. About these individual
- the SNP assay and the inoculation test were similarly performed on the deposited line and the diseased tomato plant.
- Table 6 shows the results.
- a and H are shaded.
- the disease index is 1 or less. It was.
- solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′ are SNP markers highly correlated with leaf mold resistance at the first resistance locus.
- solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ′ are highly correlated with each other, solcap_snp_sl_60250 and solcap disease region between the colcap_snp_sl_60250 and solcap_sl_s_60250 ′ It was found that there was a correlation.
- the resistance loci identified by the SNP markers solcap_snp_sl_60160, solcap_snp_sl_8658, and solcap_snp_sl_8658 ', and the leaf region of the solcap_snp_sl_60250 and the solcap_snp_sl_site are identified in It turned out that it becomes a resistance marker, and it can be screened for a leaf-mold-resistant tomato plant using the leaf-mold-resistance marker of the said tomato plant.
- Leaf mold resistance locus on chromosome 6 From the 114 strain, solcap_snp_sl_60160 on chromosome 1 is included in a susceptible homozygous form, and SG_70 on chromosome 6 is in a heterozygous form Individuals to be included were selected (second F2 selection line). Next, the second progeny progeny F3 was produced by self-propagating each of the second F2 selected lines.
- the SNP assay is performed on the second progeny progeny F3 in the same manner as in (3) above, and SG_70, and solcap_snp_sl_25325, solcap_snp_sl_25252, solcap_snp_s_25_s1_25_1_25_s1
- a polymorphic base corresponding to was identified. About these individual
- Table 7 shows the results.
- a and H are shaded.
- solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_35221, SG_70, and SG_70 ′ are resistant homozygous (A) or heterozygous (H), The index was 1 or less.
- solcap_snp_sl_25252 solcap_snp_sl_25252 ′, solcap_snp_sl_3521, SG_70, and SG_70 ′ are SNP markers highly correlated with leaf mold resistance.
- solcap_snp_sl_25252 solcap_snp_sl_25252 ′, solcap_snp_sl_3521, SG_70, and SG_70 ′ are highly correlated with each other
- solsap_sn23_sl23_sl23_s25_s25_s25_s25_s25_s25_s25_s25_s25_s25_s25_s25_s25 region that includes the SNP marker on the sixth chromosome It was found to be highly correlated with disease resistance.
- the SNP markers solcap_snp_sl_25252, solcap_snp_sl_25252 ′, solcap_snp_sl_35221, SG_70, and the resistance loci identified by SG_70 ′, and the scapap_snp_sl_253_SL_0125_specific region are identified. It turned out that it becomes a leaf mold resistance marker of a plant, and it can be screened for a leaf mold resistant tomato plant using the leaf mold resistance marker of the tomato plant.
- the deposited strain instead of the 114 strain, the deposited strain, the susceptible tomato plant, and a tomato plant containing the Cf gene shown in Table 8 below in a homozygous form were used.
- the disease index was determined in the same manner as in Example 1 (2) except that 1 to 6 were used.
- Vetomold is Cf-2
- Purdue 135 is Cf-4
- Ontario7717 is Cf-5
- Ontario7719 is Cf-9
- Ontario7716 is Cf-4
- 11 are included as leaf mold resistance genes, respectively.
- each tomato plant containing a known leaf mold resistance gene was obtained from Plant Gene Resources of Canada (PGRC).
- the leaf mold fungus strain 5 is a leaf mold fungus collected from a tomato plant that spontaneously developed leaf mold disease in a tomato cultivation field in Gifu Prefecture.
- the fungal strain 5 is obtained by conducting an inoculation test similar to Example 1 (2) on tomato plants containing Cf-2, 4, 5, 9, and 11, respectively. It has been confirmed that tomato plants containing the gene cannot be infected.
- the leaf mold fungus registered by MAFF number 712002 etc. in the agricultural biological resource gene bank can be utilized, for example.
- the leaf mold fungus strain 6 is a leaf mold fungus collected from a tomato plant that naturally developed leaf mold disease in a tomato cultivation field in Shiga Prefecture.
- the leaf mold fungus strain 6 is obtained by conducting the same inoculation test as in Example 1 (2) on tomato plants containing Cf-2, 4, 5, 9, and 11, respectively. It has been confirmed that tomato plants containing -2 and 4 can be infected.
- the fungal fungus exhibiting the same infectivity as that of the fungal strain 6 described above for example, the fungal fungus registered under MAFF No. 726530 or the like in the agricultural biological resource gene bank can be used.
- the susceptible tomato plant had a disease index of 2 against leaf mold fungus lines 1 to 6 and was susceptible to leaf mold.
- the deposited strain had a disease index of 0 and was resistant to leaf mold against leaf mold fungi strains 1-6.
- Vetomold was resistant to leaf mold fungi lines 3 and 5, but was susceptible to leaf mold fungi lines 1, 2, 4, and 6.
- Purdue 135 was resistant to leaf mold fungal strains 2 and 5, but was susceptible to leaf mold fungal strains 1, 3, 4, and 6.
- Ontario7717 was resistant to leaf mold strains 2, 3, 5 and 6, but was susceptible to leaf mold strains 1 and 4. Ontario7719 was resistant to leaf mold fungi lines 4-6, but was susceptible to leaf mold fungi lines 1-3.
- Ontario7716 was resistant to leaf mold fungal lines 2-6, but was susceptible to leaf mold fungus line 1. Therefore, the first resistance locus and the second resistance locus included in the deposited line are different from Cf-2, 4, 5, 9, and 11 which are leaf mold resistance genes. I found out that
- Example 3 By crossing the tomato plant of the present invention containing the first resistance locus and the second resistance locus with a tomato plant having other resistance, high leaf mold resistance and other resistance It was confirmed that a tomato plant having sex could be easily produced.
- LA2172 obtained from Tomato Genetics Resource Center (TGRC)
- TGRC Tomato Genetics Resource Center
- the deposited line was crossed with the B line to produce F1.
- the disease index was determined in the same manner as in Example 1 (2).
- polymorphic bases corresponding to solcap_snp_sl_60160 and solcap_snp_sl_35221 were identified in the same manner as in Example 1 (3).
- the root-knot degree which is a disease severity by the following root-knot nematode inoculation test, and the disease severity of powdery mildew were calculated
- the root-knot nematode used was derived from a single egg mass of sweet potato root-knot nematode collected from a tomato cultivation house in Yamanashi Prefecture. First, sterilized soil was put in a cultivation cup, and 2 tomato seedlings without root-knot nematode were grown for 30 days in a thermostat set at 30 ° C. ⁇ 1 ° C. Three holes for inoculation having a diameter of 10 mm and a depth of 20 mm were made with a glass rod in the soil around the grown tomato 30-day seedling stock.
- the root-knot nematode is suspended in clear water so as to be 200 heads / mL, and 1 mL of the suspension of root-knot nematode is sucked up with a Komagome pipette, and 1 mL of the suspension per hole is poured into the hole for inoculation. And further grown under the same conditions. Forty days after the inoculation, roots were collected from the grown individuals, washed with water, and the disease was investigated as follows.
- the powdery mildew was collected from a tomato plant that naturally developed powdery mildew at a tomato cultivation house in Shiga Prefecture. Then, the collected powdery mildew fungus was diluted so that the conidia concentration was 2.5 ⁇ 10 4 cells / mL, and a conidia spore suspension was prepared and used as an inoculation source. Then, to the tomato seedlings inoculated with the root-knot nematode, 1 mL of the conidial spore suspension per one tomato plant was uniformly sprayed by hand spray. On the 20th day after inoculation, the disease was investigated as follows.
- polymorphic bases corresponding to solcap_snp_sl_60160 and solcap_snp_sl_35221 are specified in the same manner except that the deposited strain and the B strain are used in place of the F1, and the disease index, the degree of rooting, and udon are identified. The severity of this disease was determined.
- F1 produced by crossing a deposited strain having resistance to leaf mold and a B strain having resistance to root-knot nematode and powdery mildew is solcap_snp_sl_60160 and solcap_snp_sl_35221 are both heterozygous. Met.
- the F1 was highly resistant to leaf mold, root-knot nematode and powdery mildew. From these results, by crossing a tomato plant containing the first resistance locus and the second resistance locus with a tomato plant having other resistance, high leaf mold resistance and other It was found that tomato plants having resistance can be easily produced.
- the leaf mold resistant tomato plant of the present invention for example, a leaf mold resistant tomato plant can be easily screened.
- the leaf mold resistant tomato plant of the present invention contains, for example, at least one of the leaf mold resistance locus on the first chromosome and the leaf mold resistance locus on the sixth chromosome, for example, leaf mold resistance can be exhibited.
- the tomato plant resistant to leaf mold of the present invention contains, for example, a dominant resistance locus, it is possible to obtain a progeny showing dominant resistance to leaf mold by crossing with other tomato plants. Can do.
- the leaf mold resistant tomato plant of the present invention also exhibits resistance to a leaf mold fungus capable of infecting a tomato plant containing the leaf mold resistance gene of the above-mentioned prior art document. For this reason, the leaf mold resistant tomato plant of the present invention does not require conventional pesticide control, and thus, for example, the labor and cost problems of the pesticide spraying can be avoided.
Landscapes
- Life Sciences & Earth Sciences (AREA)
- Health & Medical Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Genetics & Genomics (AREA)
- Engineering & Computer Science (AREA)
- Organic Chemistry (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Botany (AREA)
- Wood Science & Technology (AREA)
- Zoology (AREA)
- Analytical Chemistry (AREA)
- Biotechnology (AREA)
- General Health & Medical Sciences (AREA)
- Environmental Sciences (AREA)
- Developmental Biology & Embryology (AREA)
- Physics & Mathematics (AREA)
- General Engineering & Computer Science (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Molecular Biology (AREA)
- Biophysics (AREA)
- Microbiology (AREA)
- Biochemistry (AREA)
- Immunology (AREA)
- Physiology (AREA)
- Natural Medicines & Medicinal Plants (AREA)
- Mycology (AREA)
- Biomedical Technology (AREA)
- Spectroscopy & Molecular Physics (AREA)
- Animal Husbandry (AREA)
- Cell Biology (AREA)
- Plant Pathology (AREA)
- Breeding Of Plants And Reproduction By Means Of Culturing (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
Abstract
Description
前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、
前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定されることを特徴とする。
前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、
前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定されることを特徴とする。
(a)前記本発明の葉かび病抵抗性トマト植物と、他のトマト植物とを交雑する工程
(b)前記(a)工程より得られたトマト植物またはその後代系統から、葉かび病抵抗性トマト植物を選抜する工程
前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、
前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定されることを特徴とする。
本発明のトマト植物の葉かび病抵抗性マーカー(以下、「抵抗性マーカー」ともいう。)は、前述のように、第1染色体上の葉かび病抵抗性遺伝子座および第6染色体上の葉かび病抵抗性遺伝子座の少なくとも一方の葉かび病抵抗性遺伝子座を含み、前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定されることを特徴とする。本発明の抵抗性マーカーは、第1染色体上の葉かび病抵抗性遺伝子座および第6染色体上の葉かび病抵抗性遺伝子座の少なくとも一方の葉かび病抵抗性遺伝子座を含み、前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定されることが特徴であり、その他の構成および条件は、特に制限されない。
本発明において、前記第1の抵抗性遺伝子座は、前述のように、(1-1)前記SNPマーカーによって特定される。前記第1の抵抗性遺伝子座は、さらに、(1-2)前記SNPマーカーを含む塩基配列によって規定されてもよいし、(1-3)2つの前記SNPマーカーの部位間の領域の塩基配列によって規定されてもよいし、これらの組合せにより規定されてもよい。前記組合せによって規定する場合、前記組合せは、特に制限されず、例えば、以下の組合せが例示できる。また、前記第1の抵抗性遺伝子座は、前記(1-1)により特定されているが、前記第1の抵抗性遺伝子座は、これに限定されず、例えば、前記(1-1)に代えて、前記(1-2)または前記(1-3)で特定されてもよいし、前記(1-2)および前記(1-3)の組合せで特定されてもよい。
前記(1-1)および前記(1-2)の組合せ
前記(1-1)および前記(1-3)の組合せ
前記(1-1)、前記(1-2)および前記(1-3)の組合せ
前記第1の抵抗性遺伝子座は、前記(1-1)に示すように、前記SNPマーカーによって規定される。前記SNPマーカーは、特に制限されず、例えば、solcap_snp_sl_60160、solcap_snp_sl_8658、solcap_snp_sl_8658’等があげられる。なお、solcap_snp_sl_8658’を除く、「solcap_snp_sl_番号」で表されるSNPマーカーの表記は、当該技術分野における当業者であれば、本願の出願時の技術常識から理解可能であり、ナス科植物ゲノム研究国際コンソーシアムのwebサイト(http://solgenomics.net/)で閲覧できる。なお、これらのSNP解析は、例えば、下記参考文献1~3を参照できる(以下、同様)。また、solcap_snp_sl_8658’は、本発明者らが新たに同定したSNPマーカーであり、当該技術分野における当業者であれば、後述するこれらのSNPマーカーを含む塩基配列に基づき、前記SNPマーカーの座乗位置を特定できる。
参考文献1:Hamilton JP, Sim SC, Stoffel K, Van Deynze A, Buell CR, et al. (2012) “Single Nucleotide Polymorphism Discovery in Cultivated Tomato via Sequencing by Synthesis.”, The Plant Genome 5.
参考文献2:Sim S-C, Durstewitz G, Plieske J, Wieseke R, Ganal MW, et al., (2012) “Development of a Large SNP Genotyping Array and Generation of High-Density.”, Genetic Maps in Tomato. PLoS ONE 7(7)
参考文献3:Blanca J, Canizares J, Cordero L, Pascual L, Diez MJ, et al., (2012) “Variation Revealed by SNP Genotyping and Morphology Provides Insight into the Origin of the Tomato.”, PLoS ONE 7(10)
配列番号1
5’-TGATGGAGGAAACATTACATTCTAATATTTTCGCAGCAAACATCTACAC[T]GTTTGATGCTTTTAATGTATCAGCCTGTTATTCAGAA-3’
配列番号6
5’-GTGATGGAGGAAACATTACATTCTAATATTTTCGCAGCAAACATCTACAC[T]GTTTGATGCTTTTAATGTATCAGCCTGTTATTCAGAAGCTGTCTTCGTTT-3’
2個の組合せ
SNP(a)およびSNP(b)の組合せ
SNP(a)およびSNP(b’)の組合せ
SNP(b)およびSNP(b’)の組合せ
3個の組合せ
SNP(a)、SNP(b)、およびSNP(b’)の組合せ
前記組合せのうち、葉かび病抵抗性との相関性がより高いことから、好ましくは、例えば、以下の組合せである。
SNP(a)およびSNP(b)の組合せ
SNP(b)およびSNP(b’)の組合せ
SNP(a)、SNP(b)、およびSNP(b’)の組合せ
前記第1の抵抗性遺伝子座は、前記(1-2)に示すように、例えば、前記SNPマーカーを含む塩基配列によって規定されてもよい。前記第1の抵抗性遺伝子座は、例えば、前記塩基配列からなるものでもよいし、前記塩基配列を含むものでもよい。
(a1)配列番号1の塩基配列からなるポリヌクレオチド
(a2)前記(a1)の50番目の塩基(T)以外の塩基配列において、1もしくは数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなるポリヌクレオチド
(a3)前記(a1)の50番目の塩基(T)以外の塩基配列に対して、80%以上の同一性を有する塩基配列からなるポリヌクレオチド
(b1)配列番号2の塩基配列からなるポリヌクレオチド
(b2)前記(b1)の40番目の塩基(T)以外の塩基配列において、1もしくは数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなるポリヌクレオチド
(b3)前記(b1)の40番目の塩基(T)以外の塩基配列に対して、80%以上の同一性を有する塩基配列からなるポリヌクレオチド
(b1’)配列番号2の塩基配列からなるポリヌクレオチド
(b2’)前記(b1’)の22番目の塩基(C)以外の塩基配列において、1もしくは数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなるポリヌクレオチド
(b3’)前記(b1’)の22番目の塩基(C)以外の塩基配列に対して、80%以上の同一性を有する塩基配列からなるポリヌクレオチド
(b1’’)配列番号2の塩基配列からなるポリヌクレオチド
(b2’’)前記(b1’’)の22番目の塩基(C)および40番目の塩基(T)以外の塩基配列において、1もしくは数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなるポリヌクレオチド
(b3’’)前記(b1’’)の22番目の塩基(C)および40番目の塩基(T)以外の塩基配列に対して、80%以上の同一性を有する塩基配列からなるポリヌクレオチド
前記(a)のポリヌクレオチドおよび前記(b)のポリヌクレオチドの組合せ
前記第1の抵抗性遺伝子座は、前記(1-3)に示すように、例えば、前記2つのSNPマーカーの部位間の領域の塩基配列によって規定されてもよい。前記2つのSNPマーカーの部位間の領域の塩基配列は、特に制限されず、例えば、前記染色体における、solcap_snp_sl_60250、solcap_snp_sl_60160、solcap_snp_sl_8658、solcap_snp_sl_8658’、およびsolcap_snp_sl_60089からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列、solcap_snp_sl_60250、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_60089からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列等があげられる。
配列番号11
5’-GATGAACTATGGAGAAATACTGTAGTGAGTAAATATTTGAGAATAAGCA[G]ATAGATGTAAGATAAAATGAATACATGACAAGA-3’
配列番号12
5’-GATGAACTATGGAGAAATACTGTAGTGAGTAAATATTTGAGAATAAGCA[G]ATAGATGTAAGATAAAATGAATACATGACAAGA-3’
配列番号13
5’-TTTCGTGTTTCGTATTTAGAACTTCTTAATGAAAAT[C]CTGGCTCAGCAGCTGGTAGTTAGTAGAGTGATTTGTGGGTGTATAT-3’
配列番号14
5’-TTTCGTGTTTCGTATTTAGAACTTCTTAATGAAAAT[C]CTGGCTCCGCCGCTGGTAGTTAGTAGAGTGATTTGTGGGTGTATAT-3’
solcap_snp_sl_60250およびsolcap_snp_sl_60160の組合せ
solcap_snp_sl_60250およびsolcap_snp_sl_8658’の組合せ
solcap_snp_sl_60250およびsolcap_snp_sl_8658の組合せ
solcap_snp_sl_60250およびsolcap_snp_sl_60089の組合せ
solcap_snp_sl_60160およびsolcap_snp_sl_8658’の組合せ
solcap_snp_sl_60160およびsolcap_snp_sl_8658の組合せ
solcap_snp_sl_60160およびsolcap_snp_sl_60089の組合せ
solcap_snp_sl_8658’およびsolcap_snp_sl_60089の組合せ
solcap_snp_sl_8658’およびsolcap_snp_sl_8658の組合せ
solcap_snp_sl_8658およびsolcap_snp_sl_60089の組合せ
条件(i)
前記染色体における、solcap_snp_sl_60250およびsolcap_snp_sl_60089の部位間の領域の塩基配列を含み、且つ、
前記領域の塩基配列において、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも一つのSNPマーカーを含む
2個の組合せ
SNP(a)およびSNP(b)の組合せ
SNP(a)およびSNP(b’)の組合せ
SNP(b)およびSNP(b’)の組合せ
3個の組合せ
SNP(a)、SNP(b)、およびSNP(b’)の組合せ
本発明において、前記第2の抵抗性遺伝子座は、前述のように、(2-1)前記SNPマーカーによって規定される。前記第2の抵抗性遺伝子座は、さらに、(2-2)前記SNPマーカーを含む塩基配列によって規定されてもよいし、(2-3)2つの前記SNPマーカーの部位間の領域の塩基配列によって規定されてもよいし、これらの組合せにより規定されてもよい。前記組合せによって規定する場合、前記組合せは、特に制限されず、例えば、以下の組合せが例示できる。また、前記第2の抵抗性遺伝子座は、前記(2-1)により特定されているが、前記第2の抵抗性遺伝子座は、これに限定されず、例えば、前記(2-1)に代えて、前記(2-2)または(2-3)で特定されてもよいし、これらの組合せで特定されてもよい。
前記(2-1)および前記(2-2)の組合せ
前記(2-1)および前記(2-3)の組合せ
前記(2-1)、前記(2-2)および前記(2-3)の組合せ
前記第2の抵抗性遺伝子座は、前記(2-1)に示すように、例えば、前記SNPマーカーによって規定されてもよい。前記SNPマーカーは、特に制限されず、例えば、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、SG_70’等があげられる。なお、solcap_snp_sl_25252’を除く、「solcap_snp_sl_番号」で表されるSNPマーカーは、前述のwebサイトで閲覧できる。また、solcap_snp_sl_25252’、SG_70、およびSG_70’は、本発明者らが新たに同定したSNPマーカーであり、当該技術分野における当業者であれば、後述するこれらのSNPマーカーを含む塩基配列に基づき、前記SNPマーカーの座乗位置を特定できる。
配列番号4
5’-TCTAATACCATCTGCAAGTTTCTGAGCCTC[A]TCTGACTTCAGCGGACATCTACTTAAG-3’
配列番号9
5’-CTCGAAATTGATAAGTTTTCTCTAATACCATCTGCAAGTTTCTGAGCCTC[A]TCTGACTTCAGCGGACATCTACTTAAGGTCTGTAAAACAATCAACTTTAT-3’
2個の組合せ
SNP(c)およびSNP(c’)の組合せ
SNP(c)およびSNP(d)の組合せ
SNP(c)およびSNP(e)の組合せ
SNP(c)およびSNP(e’)の組合せ
SNP(c’)およびSNP(d)の組合せ
SNP(c’)およびSNP(e)の組合せ
SNP(c’)およびSNP(e’)の組合せ
SNP(d)およびSNP(e)の組合せ
SNP(d)およびSNP(e’)の組合せ
SNP(e)およびSNP(e’)の組合せ
3個の組合せ
SNP(c)、SNP(c’)、およびSNP(d)の組合せ
SNP(c)、SNP(c’)、およびSNP(e)の組合せ
SNP(c)、SNP(c’)、およびSNP(e’)の組合せ
SNP(c)、SNP(d)、およびSNP(e)の組合せ
SNP(c)、SNP(d)、およびSNP(e’)の組合せ
SNP(c)、SNP(e)、およびSNP(e’)の組合せ
SNP(c’)、SNP(d)、およびSNP(e)の組合せ
SNP(c’)、SNP(d)、およびSNP(e’)の組合せ
SNP(c’)、SNP(e)、およびSNP(e’)の組合せ
SNP(d)、SNP(e)、およびSNP(e’)の組合せ
4個の組合せ
SNP(c)、SNP(c’)、SNP(d)、およびSNP(e)の組合せ
SNP(c)、SNP(c’)、SNP(d)、およびSNP(e’)の組合せ
SNP(c)、SNP(c’)、SNP(e)、およびSNP(e’)の組合せ
SNP(c)、SNP(d)、SNP(e)、およびSNP(e’)の組合せ
SNP(c’)、SNP(d)、SNP(e)、およびSNP(e’)の組合せ
5個の組合せ
SNP(c)、SNP(c’)、SNP(d)、SNP(e)、およびSNP(e’)の組合せ
前記組合せのうち、葉かび病抵抗性との相関性がより高いことから、好ましくは、例えば、以下の組合せである。
SNP(c)およびSNP(c’)の組合せ
SNP(e)およびSNP(e’)の組合せ
SNP(c)、SNP(d)、およびSNP(e)の組合せ
SNP(c)、SNP(c’)、SNP(d)、SNP(e)、およびSNP(e’)の組合せ
前記第2の抵抗性遺伝子座は、前記(2-2)に示すように、例えば、前記SNPマーカーを含む塩基配列によって規定されてもよい。前記第2の抵抗性遺伝子座は、例えば、前記塩基配列からなるものでもよいし、前記塩基配列を含むものでもよい。
(c1)配列番号3の塩基配列からなるポリヌクレオチド
(c2)前記(c1)の34番目の塩基(G)以外の塩基配列において、1もしくは数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなるポリヌクレオチド
(c3)前記(c1)の34番目の塩基(G)以外の塩基配列に対して、80%以上の同一性を有する塩基配列からなるポリヌクレオチド
(c1’)配列番号3の塩基配列からなるポリヌクレオチド
(c2’)前記(c1’)の46番目の塩基(T)以外の塩基配列において、1もしくは数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなるポリヌクレオチド
(c3’)前記(c1’)の46番目の塩基(T)以外の塩基配列に対して、80%以上の同一性を有する塩基配列からなるポリヌクレオチド
(c1’’)配列番号3の塩基配列からなるポリヌクレオチド
(c2’’)前記(c1’’)の34番目の塩基(G)および46番目の塩基(T)以外の塩基配列において、1もしくは数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなるポリヌクレオチド
(c3’’)前記(c1’’)の34番目の塩基(G)および46番目の塩基(T)以外の塩基配列に対して、80%以上の同一性を有する塩基配列からなるポリヌクレオチド
(d1)配列番号4の塩基配列からなるポリヌクレオチド
(d2)前記(d1)の31番目の塩基(A)以外の塩基配列において、1もしくは数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなるポリヌクレオチド
(d3)前記(d1)の31番目の塩基(A)以外の塩基配列に対して、80%以上の同一性を有する塩基配列からなるポリヌクレオチド
(e1)配列番号5の塩基配列からなるポリヌクレオチド
(e2)前記(e1)の40番目の塩基(A)以外の塩基配列において、1もしくは数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなるポリヌクレオチド
(e3)前記(e1)の40番目の塩基(A)以外の塩基配列に対して、80%以上の同一性を有する塩基配列からなるポリヌクレオチド
(e1’)配列番号5の塩基配列からなるポリヌクレオチド
(e2’)前記(e1’)の75番目の塩基(G)以外の塩基配列において、1もしくは数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなるポリヌクレオチド
(e3’)前記(e1’)の75番目の塩基(G)以外の塩基配列に対して、80%以上の同一性を有する塩基配列からなるポリヌクレオチド
(e1’’)配列番号5の塩基配列からなるポリヌクレオチド
(e2’’)前記(e1’’)の40番目の塩基(A)および75番目の塩基(G)以外の塩基配列において、1もしくは数個の塩基が欠失、置換、挿入および/または付加された塩基配列からなるポリヌクレオチド
(e3’’)前記(e1’’)の40番目の塩基(A)および75番目の塩基(G)以外の塩基配列に対して、80%以上の同一性を有する塩基配列からなるポリヌクレオチド
2個の組合せ
(c)のポリヌクレオチドおよび(d)のポリヌクレオチドの組合せ
(c)のポリヌクレオチドおよび(e)のポリヌクレオチドの組合せ
(d)のポリヌクレオチドおよび(e)のポリヌクレオチドの組合せ
3個の組合せ
(c)のポリヌクレオチド、(d)のポリヌクレオチド、および(e)のポリヌクレオチドの組合せ
前記第2の抵抗性遺伝子座は、前記(2-3)に示すように、例えば、前記2つのSNPマーカーの部位間の領域の塩基配列によって規定されてもよい。前記2つのSNPマーカーの部位間の領域の塩基配列は、特に制限されず、例えば、前記染色体における、solcap_snp_sl_25325、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、SG_70’、およびSL10401_823からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列、solcap_snp_sl_25325、solcap_snp_sl_25252、solcap_snp_sl_35221、SG_70、およびSL10401_823からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列等があげられる。
配列番号15
5’-TCACGAGCAGCAAGAAAGCTATTATCGCTAGTATCCAATA[G]TACGAAAGAGAACACTCCATCCAACATGT-3’
配列番号16
5’-TCCAATGGCATCACGAGCAGCAAGAAAGCTATTATCGCGAGTATCCAATA[G]TACGAAAGAGAACACTCCATCCAACATGTCAACAAAATTTTCTCCATACT-3’
配列番号17
5’-TACATTCTGTTTTATTTGTTCCTTACTTGTCT[T]AGTTGGGCTCATATCTCATTTTGAATGTAAATTACAGGTGCAAGACAA-3’
配列番号18
5’-TACATTCTGTTTTATTTGTTCCTTACTTGTCT[T]AGTTGGGCTCATATCTCATTTTGAATGTAAATTACAGGTGCAAGACAA-3’
solcap_snp_sl_25325およびsolcap_snp_sl_25252の組合せ
solcap_snp_sl_25325およびsolcap_snp_sl_25252’の組合せ
solcap_snp_sl_25325およびsolcap_snp_sl_35221の組合せ
solcap_snp_sl_25325およびSG_70の組合せ
solcap_snp_sl_25325およびSG_70’の組合せ
solcap_snp_sl_25325およびSL10401_823の組合せ
solcap_snp_sl_25252およびsolcap_snp_sl_25252’の組合せ
solcap_snp_sl_25252およびsolcap_snp_sl_35221の組合せ
solcap_snp_sl_25252およびSG_70の組合せ
solcap_snp_sl_25252およびSG_70’の組合せ
solcap_snp_sl_25252およびSL10401_823の組合せ
solcap_snp_sl_25252’およびsolcap_snp_sl_35221の組合せ
solcap_snp_sl_25252’およびSG_70の組合せ
solcap_snp_sl_25252’およびSG_70’の組合せ
solcap_snp_sl_25252’およびSL10401_823の組合せ
solcap_snp_sl_35221およびSG_70の組合せ
solcap_snp_sl_35221およびSG_70’の組合せ
solcap_snp_sl_35221およびSL10401_823の組合せ
SG_70およびSG_70’の組合せ
SG_70およびSL10401_823の組合せ
SG_70’およびSL10401_823の組合せ
条件(ii)
前記染色体における、solcap_snp_sl_25325およびSL10401_823の部位間の領域の塩基配列を含み、且つ、
前記領域の塩基配列において、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも1つのSNPマーカーを含む
2個の組合せ
SNP(c)およびSNP(c’)の組合せ
SNP(c)およびSNP(d)の組合せ
SNP(c)およびSNP(e)の組合せ
SNP(c)およびSNP(e’)の組合せ
SNP(c’)およびSNP(d)の組合せ
SNP(c’)およびSNP(e)の組合せ
SNP(c’)およびSNP(e’)の組合せ
SNP(d)およびSNP(e)の組合せ
SNP(d)およびSNP(e’)の組合せ
SNP(e)およびSNP(e’)の組合せ
3個の組合せ
SNP(c)、SNP(c’)、およびSNP(d)の組合せ
SNP(c)、SNP(c’)、およびSNP(e)の組合せ
SNP(c)、SNP(c’)、およびSNP(e’)の組合せ
SNP(c)、SNP(d)、およびSNP(e)の組合せ
SNP(c)、SNP(d)、およびSNP(e’)の組合せ
SNP(c)、SNP(e)、およびSNP(e’)の組合せ
SNP(c’)、SNP(d)、およびSNP(e)の組合せ
SNP(c’)、SNP(d)、およびSNP(e’)の組合せ
SNP(c’)、SNP(e)、およびSNP(e’)の組合せ
SNP(d)、SNP(e)、およびSNP(e’)の組合せ
4個の組合せ
SNP(c)、SNP(c’)、SNP(d)、およびSNP(e)の組合せ
SNP(c)、SNP(c’)、SNP(d)、およびSNP(e’)の組合せ
SNP(c)、SNP(c’)、SNP(e)、およびSNP(e’)の組合せ
SNP(c)、SNP(d)、SNP(e)、およびSNP(e’)の組合せ
SNP(c’)、SNP(d)、SNP(e)、およびSNP(e’)の組合せ
5個の組合せ
SNP(c)、SNP(c’)、SNP(d)、SNP(e)、およびSNP(e’)の組合せ
参考情報4:南信試病害虫土壌肥料部、 “トマトのトマト葉かび病防除にエコショットが有効である” 、 [online]、長野県、[平成27年3月3日検索]、インターネット〈URL: http://www.pref.nagano.lg.jp/nogi/sangyo/nogyo/gijutsu/fukyugijutsu/200802/200802fukyu.html〉
本発明の葉かび病抵抗性トマト植物は、前述のように、第1染色体上の葉かび病抵抗性遺伝子座および第6染色体上の葉かび病抵抗性遺伝子座の少なくとも一方の葉かび病抵抗性遺伝子座を含み、前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定されることを特徴とする。本発明の葉かび病抵抗性トマト植物は、第1染色体上の葉かび病抵抗性遺伝子座および第6染色体上の葉かび病抵抗性遺伝子座の少なくとも一方の葉かび病抵抗性遺伝子座を含み、前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定されることを特徴とし、その他の構成および条件は、特に制限されない。本発明の葉かび病抵抗性トマト植物は、前記第1の抵抗性遺伝子座および前記第2の抵抗性遺伝子座の少なくとも一方を含む前記本発明の抵抗性マーカーを含むことから、例えば、前記本発明の抵抗性マーカーの説明を援用できる。本発明において、前記第1染色体上の葉かび病抵抗性遺伝子座および前記第6染色体上の葉かび病抵抗性遺伝子座は、例えば、それぞれ、本発明の抵抗性マーカーにおける第1の抵抗性遺伝子座および第2の抵抗性遺伝子座と読み替え可能である。本発明の葉かび病抵抗性トマト植物は、例えば、前記本発明の抵抗性マーカー等の説明を援用できる。
寄託の種類:国際寄託
寄託機関名:独立行政法人製品評価技術基盤機構 特許生物寄託センター
あて名:日本国 〒292-0818 千葉県木更津市かずさ鎌足2-5-8 120号室
受託番号:FERM BP-22282
識別のための表示:Takii7
受領日:2015年1月28日
つぎに、本発明の葉かび病抵抗性トマト植物の製造方法(以下、「製造方法」ともいう。)について説明する。なお、以下の方法は、例示であって、本発明は、これらの方法に制限されない。本発明において、製造方法は、例えば、育成方法ということもできる。また、本発明において、前記葉かび病抵抗性遺伝子座は、前記本発明の抵抗性マーカーと言い換えることができる。
(a)本発明の葉かび病抵抗性トマト植物と、他のトマト植物とを交雑する工程
(b)前記(a)工程より得られた(1以上の)トマト植物またはその後代系統から、葉かび病抵抗性を備えるトマト植物を選抜する工程
(x)(1以上の)被検トマト植物から、前記本発明の葉かび病抵抗性トマト植物を選抜する工程
(x1)前記(1以上の)被検トマト植物の染色体上における、前記葉かび病抵抗性遺伝子座の有無を検出する検出工程
(x2)前記葉かび病抵抗性遺伝子座の存在により、前記被検トマト植物を、葉かび病抵抗性トマト植物として選抜する選抜工程
前記(x)工程における選抜が、前記第1の抵抗性遺伝子座の選抜である場合、前記第1の抵抗性遺伝子座は、前述のように、(1-1)前記SNPマーカーによって選抜される。前記第1の抵抗性遺伝子座は、さらに、(1-2)前記SNPマーカーを含む塩基配列によって選抜されてもよいし、(1-3)2つの前記SNPマーカーの部位間の領域の塩基配列によって選抜されてもよいし、これらの組合せにより選抜されてもよい。前記組合せによって選抜する場合、前記組合せは、特に制限されず、例えば、以下の組合せが例示できる。また、前記第1の抵抗性遺伝子座は、前記(1-1)により選抜されているが、前記第1の抵抗性遺伝子座は、これに限定されず、例えば、前記(1-1)に代えて、前記(1-2)または(1-3)で選抜されてもよいし、前記(1-2)および前記(1-3)の組合せで選抜されてもよい。
前記(1-1)および前記(1-2)の組合せ
前記(1-1)および前記(1-3)の組合せ
前記(1-1)、前記(1-2)および前記(1-3)の組合せ
前記第1の抵抗性遺伝子座において、前記選択されるSNPマーカーは、特に制限されず、例えば、前記本発明の抵抗性マーカーにおける「(1-1)SNPマーカーによる特定」の説明を援用できる。
前記(x)工程における前記選抜は、例えば、前記第1染色体上の葉かび病抵抗性遺伝子座を含む葉かび病抵抗性トマト植物の選抜であり、前記第1染色体上の葉かび病抵抗性遺伝子が、前記(a)および(b)の少なくとも一方のポリヌクレオチドで特定される。前記(a)および(b)のポリヌクレオチドは、例えば、前記本発明の抵抗性マーカーにおける「(1-2)SNPマーカーを含む塩基配列による特定」の説明を援用でききる。
前記(x)工程における前記選抜は、例えば、前記第1染色体上の葉かび病抵抗性遺伝子座を含む葉かび病抵抗性トマト植物の選抜であり、前記第1染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_60250、solcap_snp_sl_60160、solcap_snp_sl_8658、solcap_snp_sl_8658’、およびsolcap_snp_sl_60089からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列を含む。前記2つのSNPマーカーの部位間の領域の塩基配列は、例えば、前記本発明の抵抗性マーカーにおける「(1-3)2つのSNPマーカーの部位間の領域の塩基配列による特定」の説明を援用できる。
前記選抜が、前記第2の抵抗性遺伝子座の選抜である場合、前記第2の抵抗性遺伝子座は、前述のように、(2-1)前記SNPマーカーによって選抜される。前記第2の抵抗性遺伝子座は、さらに、(2-2)前記SNPマーカーを含む塩基配列によって選抜されてもよいし、(2-3)2つの前記SNPマーカーの部位間の領域の塩基配列によって選抜されてもよいし、これらの組合せにより選抜されてもよい。前記組合せによって選抜する場合、前記組合せは、特に制限されず、例えば、以下の組合せが例示できる。また、前記第2の抵抗性遺伝子座は、前記(2-1)により選抜されているが、前記第2の抵抗性遺伝子座は、これに限定されず、例えば、前記(2-1)に代えて、前記(2-2)または(2-3)で選抜されてもよいし、これらの組合せで選抜されてもよい。
前記(2-1)および前記(2-2)の組合せ
前記(2-1)および前記(2-3)の組合せ
前記(2-1)、前記(2-2)および前記(2-3)の組合せ
前記第2の抵抗性遺伝子座において、前記選択されるSNPマーカーは、特に制限されず、例えば、前記本発明の抵抗性マーカーにおける「(2-1)SNPマーカーによる特定」の説明を援用できる。
前記(x)工程における前記選抜は、例えば、前記第6染色体上の葉かび病抵抗性遺伝子座を含む葉かび病抵抗性トマト植物の選抜であり、前記第6染色体上の葉かび病抵抗性遺伝子座が、前記(c)、(d)および(e)からなる群から選択された少なくとも1つで特定される。前記(c)、(d)および(e)のポリヌクレオチドは、例えば、前記本発明の抵抗性マーカーにおける「(2-2)SNPマーカーを含む塩基配列による特定」の説明を援用でききる。
前記(x)工程における前記選抜は、例えば、前記第6染色体上の葉かび病抵抗性遺伝子座を含む葉かび病抵抗性トマト植物の選抜であり、前記第6染色体上の葉かび病抵抗性遺伝子が、前記染色体における、solcap_snp_sl_25325、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、SG_70’、およびSL10401_823からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列を含む。前記2つのSNPマーカーの部位間の領域の塩基配列は、例えば、前記本発明の抵抗性マーカーにおける「(2-3)2つのSNPマーカーの部位間の領域の塩基配列による特定」の説明を援用できる。
(b1)前記(a)工程より得られた(1以上の)トマト植物またはその後代系統について、染色体上における、葉かび病抵抗性遺伝子座の有無を検出する検出工程
(b2)前記葉かび病抵抗性遺伝子座の存在により、前記(a)工程により得られた(1以上の)トマト植物またはその後代系統を、葉かび病抵抗性トマト植物として選抜する選抜工程
本発明の葉かび病抵抗性トマト植物のスクリーニング方法(以下、「スクリーニング方法」ともいう。)は、交雑により葉かび病抵抗性トマト植物を生産するための親として、(1以上の)被検トマト植物から、トマト植物の葉かび病抵抗性マーカーとして第1染色体上の葉かび病抵抗性遺伝子座および第6染色体上の葉かび病抵抗性遺伝子座の少なくとも一方を含む葉かび病抵抗性トマト植物を選抜する工程を含み、前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定されることを特徴とする。
新規な葉かび病抵抗性トマト植物について、葉かび病抵抗性遺伝子座の遺伝様式の解析、前記葉かび病抵抗性遺伝子座の特定、および前記葉かび病抵抗性遺伝子座と葉かび病抵抗性との相関を確認することで、前記葉かび病抵抗性遺伝子座が、トマト植物の葉かび病抵抗性マーカーとなること、前記抵抗性遺伝子座を含むトマト植物が、葉かび病抵抗性トマト植物であること、ならびに前記トマト植物の葉かび病抵抗性マーカーを使用し、葉かび病抵抗性トマト植物をスクリーニングできることを確認した。
葉かび病抵抗性を示す新規トマト植物を開発するために、タキイ種苗株式会社農場で継代育種により採取された大量のトマト系統の種子について、育種を行い、葉かび病抵抗性の試験を行った。その結果、葉かび病抵抗性を示す、新規の葉かび病抵抗性トマト系統(S.lycopersicum)を製造(以下、「生産」ともいう。)した。この新規葉かび病抵抗性トマト植物は、受託番号FERM BP-22282で寄託した。以下、この葉かび病抵抗性トマト植物を寄託系統という。
前記寄託系統のトマト植物(受託番号FERM BP-22282)と、葉かび病罹病性トマト植物「桃太郎(タキイ種苗株式会社、S.lycopersicum)」(以下、「罹病性トマト植物」ともいう。)とを交雑することによって、114個体のF2分離集団(以下、「114系統」ともいう。)を生産した。さらに、前記114系統をそれぞれ自殖させることによって、自殖後代F3を生産した。各系統のF3をそれぞれ9~15個体使用し、以下に示すように、葉かび病菌の接種試験を行った。なお、前記寄託系統は、後述するように、第1染色体上のsolcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’を抵抗性のホモ接合型で含み、且つ第6染色体上のsolcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’を抵抗性のホモ接合型で含む。また、前記罹病性トマト植物は、後述するように、第1染色体上のsolcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’を罹病性のホモ接合型で含み、且つ第6染色体上のsolcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’を罹病性のホモ接合型で含む。
参考文献5:黒柳悟ら、「トマト葉かび病抵抗性遺伝子(Cf-9)に連鎖したDNAマーカーの開発」、愛知農総試研報、2010年、42巻、15-22頁
発病指数0:胞子形成が認められない(高い抵抗性)
発病指数1:胞子形成が認められるが、病斑面積が葉面積の50%未満である(抵抗性)
発病指数2:胞子形成が認められ、病斑面積が葉面積の50%以上である(罹病性)
発病指数=[(0×n0)+(1×n1)+(2×n2)]/調査個体数
前記式において、「0、1、2」は、それぞれ発病指数を示し、「n0、n1、n2」は、それぞれ、発病指数0、発病指数1、発病指数2の個体数を示す。
発病指数=0 : 25個体
0<発病指数≦0.1 : 1個体
0.1<発病指数≦0.5 : 22個体
0.5<発病指数≦1.0 : 32個体
1.0<発病指数≦1.5 : 11個体
1.5<発病指数<2.0 : 9個体
発病指数=2 : 14個体
合計 114個体
つぎに、前記114系統から、常法により、それぞれの系統のDNAを抽出した。そして、前記DNAについて、SOLCAPのSNPアッセイ(前記参考文献1~3参照)でジェノタイピングを行い、ソフトウェア(Join Map)を用いた連鎖地図の作成、およびソフトウェア(Win QTL cartographer)を用いた連鎖解析をおこなった。
前記114系統から、前記第1染色体上のsolcap_snp_sl_60160を抵抗性のホモ接合型、ヘテロ接合型、または罹病性のホモ接合型で含み、且つ前記第6染色体上のSG_70を抵抗性のホモ接合型、ヘテロ接合型、または罹病性のホモ接合型で含む個体を、それぞれ5個体ずつ選抜した。そして、前記114系統に代えて前記選抜したトマト植物を用い、葉かび病菌系統1に代えて、葉かび病菌系統1~4を用いた以外は、前記(2)と同様にして、発病指数を求めた。また、前記選抜したトマト植物に代えて、前記寄託系統または前記罹病性トマト植物を用いた以外は、同様にして発病指数を求めた。なお、葉かび病菌系統2は、岩手県のトマト栽培圃場において、葉かび病を自然発症したトマト植物から採取した葉かび病菌である。また、前記葉かび病菌系統2は、Cf-2、4、5、9、および11を、それぞれ含むトマト植物に対して、前記(2)と同様の接種試験を行うことにより、Cf-2および9を含むトマト植物に対して、感染可能であることを確認している。なお、前記葉かび病菌系統2と同様の感染性を示す葉かび病菌は、例えば、熊本県病害虫防除所から入手できる。前記葉かび病菌系統3は、福島県のトマト栽培圃場において、葉かび病を自然発症したトマト植物から採取した葉かび病菌である。また、前記葉かび病菌系統3は、Cf-2、4、5、9、および11を、それぞれ含むトマト植物に対して、前記(2)と同様の接種試験を行うことにより、Cf-4および9を含むトマト植物に対して、感染可能であることを確認している。前記葉かび病菌系統3と同様の感染性を示す葉かび病菌は、例えば、野菜茶業研究所から入手できる。前記葉かび病菌系統4は、Wageningen大学から入手した葉かび病菌であり、また、Cf-2、4および5を含むトマト植物に対して、感染可能な葉かび病菌であることが知られている。
前記114系統から、前記第1染色体上のsolcap_snp_sl_60160をヘテロ接合型で含み、且つ前記第6染色体上のSG_70を罹病性のホモ接合型で含む個体を選抜した(第1のF2選抜系統)。つぎに、前記第1のF2選抜系統をそれぞれ自殖させることによって、第1の自殖後代F3を生産した。そして、前記第1の自殖後代F3について、前記(3)と同様の方法により、SNPアッセイを行い、solcap_snp_sl_60160、ならびにsolcap_snp_sl_60160近傍のSNPマーカーであるsolcap_snp_sl_60303、solcap_snp_sl_60250、solcap_snp_sl_8658、solcap_snp_sl_8658’、solcap_snp_sl_60089、およびsolcap_snp_sl_60043に対応する多型の塩基を特定した。これらの個体について、前記(2)と同様にして、葉かび病菌の接種試験を行った。また、前記寄託系統および前記罹病性トマト植物についても、同様にSNPアッセイおよび接種試験を行った。得られた結果の内、前記第1の自殖後代F3において、特定した前記SNPマーカーの遺伝子型が互いに異なる6個体(植物体X1-1~6)、前記寄託系統、および前記罹病性トマト植物の結果を表6に示す。表6において、AおよびHは、網掛けで示している。表6に示すように、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’が、抵抗性のホモ接合型(A)またはヘテロ接合型(H)である個体においては、いずれの個体も、発病指数が1以下であった。これらの結果から、前記第1の抵抗性遺伝子座において、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’が葉かび病抵抗性と高い相関性を示すSNPマーカーであることが確認できた。また、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’と高い相関性を示すことから、前記第1染色体上において、前記SNPマーカーを含む領域であるsolcap_snp_sl_60250およびsolcap_snp_sl_60089の部位間の領域が、葉かび病抵抗性と高い相関性を示すことがわかった。これらの結果から、SNPマーカーであるsolcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’で特定される抵抗性遺伝子座、ならびにsolcap_snp_sl_60250およびsolcap_snp_sl_60089の部位間の領域で特定される抵抗性遺伝子座が、トマト植物の葉かび病抵抗性マーカーとなること、ならびに前記トマト植物の葉かび病抵抗性マーカーを使用し、葉かび病抵抗性トマト植物をスクリーニングできることがわかった。
前記114系統から、前記第1染色体上のsolcap_snp_sl_60160を罹病性のホモ接合型で含み、且つ前記第6染色体上のSG_70をヘテロ接合型で含む個体を選抜した(第2のF2選抜系統)。つぎに、前記第2のF2選抜系統をそれぞれ自殖させることによって、第2の自殖後代F3を生産した。そして、前記第2の自殖後代F3について、前記(3)と同様の方法により、SNPアッセイを行い、SG_70、ならびにSG_70近傍のSNPマーカーであるsolcap_snp_sl_25325、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70’およびSL10401_823に対応する多型の塩基を特定した。これらの個体について、前記(2)と同様にして、葉かび病菌の接種試験を行った。また、前記寄託系統および前記罹病性トマト植物についても、同様にSNPアッセイおよび接種試験を行った。得られた結果の内、前記第2の自殖後代F3において、特定した前記SNPマーカーの遺伝子型が互いに異なる7個体(植物体X6-1~7)、前記寄託系統、および前記罹病性トマト植物の結果を表7に示す。また、前記表7において、AおよびHは、網掛けで示している。前記表7に示すように、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’が、抵抗性のホモ接合型(A)またはヘテロ接合型(H)である個体においては、いずれの個体も、発病指数が1以下であった。これらの結果から、前記第2の抵抗性遺伝子座において、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’が葉かび病抵抗性と高い相関性を示すSNPマーカーであることが確認できた。また、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’と高い相関性を示すことから、前記第6染色体上において、前記SNPマーカーを含む領域であるsolcap_snp_sl_25325およびSL10401_823の部位間の領域が、葉かび病抵抗性と高い相関性を示すことがわかった。これらの結果から、SNPマーカーであるsolcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’で特定される抵抗性遺伝子座、ならびにsolcap_snp_sl_25325およびSL10401_823の部位間の領域で特定される抵抗性遺伝子座が、トマト植物の葉かび病抵抗性マーカーとなること、ならびに前記トマト植物の葉かび病抵抗性マーカーを使用し、葉かび病抵抗性トマト植物をスクリーニングできることがわかった。
新規な葉かび病抵抗性遺伝子座が、既知の葉かび病抵抗性遺伝子であるCf-2、4、5、9、および11と異なることを確認した。
前記第1の抵抗性遺伝子座および前記第2の抵抗性遺伝子座を含む本発明のトマト植物と他の抵抗性を有するトマト植物とを交雑することにより、高い葉かび病抵抗性と他の抵抗性とを有するトマト植物を容易に生産できることを確認した。
参考文献6:Alireza Seifi et.al., “Linked, if Not the Same, Mi-1 Homologues Confer Resistance to Tomato Powdery Mildew and Root-Knot Nematodes”, MPMI, 2011, Vol.24, No.4, pp.441-450
参考文献7:線虫学実験法(日本線虫学会)2004年3月25日発行
根こぶ程度0:根こぶなし
根こぶ程度1:根こぶがわずかに認められる
根こぶ程度2:一見して根こぶが認められる
参考文献8:G.B.Cap et al., “Inheritance of heat-stable resistance to Meloidogyne incognita in Lycopersicon peruvianum and its relationship to the Mi gene”, Theoretical and Applied Genetics, Springer, 1993, volume 85, pp. 777-783
発病度0:症状なし
発病度1:接種した葉にわずかにうどんこ病菌の胞子形成が見られる
発病度2:接種した葉に多くのうどんこ病菌の胞子形成が見られる
発病度=[(0×n0)+(1×n1)+(2×n2)]/調査個体数
ネコブセンチュウの接種試験における発病度の場合、前記式において、「0、1、2」は、それぞれ根こぶ程度を示し、「n0、n1、n2」は、それぞれ、根こぶ程度0、根こぶ程度1、根こぶ程度2の個体数を示す。また、うどんこ病の発病度の場合、前記式において、「0、1、2」は、それぞれうどんこ病の発病度を示し、「n0、n1、n2」は、それぞれ、発病度0、発病度1、発病度2の個体数を示す。
Claims (48)
- 第1染色体上の葉かび病抵抗性遺伝子座および第6染色体上の葉かび病抵抗性遺伝子座の少なくとも一方の葉かび病抵抗性遺伝子座を含み、
前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、
前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定されることを特徴とする、トマト植物の葉かび病抵抗性マーカー。 - 前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160およびsolcap_snp_sl_8658で特定される、請求項1記載のトマト植物の葉かび病抵抗性マーカー。
- 前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’で特定される、請求項1または2記載のトマト植物の葉かび病抵抗性マーカー。
- 前記第1染色体上の葉かび病抵抗性遺伝子座が、下記(a)および(b)の少なくとも一方のポリヌクレオチドで特定される、請求項1から3のいずれか一項に記載のトマト植物の葉かび病抵抗性マーカー。
(a) 下記(a1)のポリヌクレオチド
(a1)配列番号1の塩基配列からなるポリヌクレオチド
(b) 下記(b1)のポリヌクレオチド
(b1)配列番号2の塩基配列からなるポリヌクレオチド - 前記第1染色体上の葉かび病抵抗性遺伝子座が、前記(a)および(b)のポリヌクレオチドで特定される、請求項4記載のトマト植物の葉かび病抵抗性マーカー。
- 前記第1染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_60250、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_60089からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列を含む、請求項1から3のいずれか一項に記載のトマト植物の葉かび病抵抗性マーカー。
- 前記第1染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_60250およびsolcap_snp_sl_60089の部位間の領域の塩基配列を含む、請求項6記載のトマト植物の葉かび病抵抗性マーカー。
- 前記トマト植物の葉かび病抵抗性マーカーが、前記第1染色体上の葉かび病抵抗性遺伝子座および前記第6染色体上の葉かび病抵抗性遺伝子座を含む、請求項1から7のいずれか一項に記載のトマト植物の葉かび病抵抗性マーカー。
- 前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_35221、およびSG_70で特定される、請求項1から8のいずれか一項に記載のトマト植物の葉かび病抵抗性マーカー。
- 前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’で特定される、請求項1から9のいずれか一項に記載のトマト植物の葉かび病抵抗性マーカー。
- 前記第6染色体上の葉かび病抵抗性遺伝子座が、(c)、(d)および(e)からなる群から選択された少なくとも一つのポリヌクレオチドで特定される、請求項1から10のいずれか一項に記載のトマト植物の葉かび病抵抗性マーカー。
(c) 下記(c1)のポリヌクレオチド
(c1)配列番号3の塩基配列からなるポリヌクレオチド
(d) 下記(d1)のポリヌクレオチド
(d1)配列番号4の塩基配列からなるポリヌクレオチド
(e) 下記(e1)のポリヌクレオチド
(e1)配列番号5の塩基配列からなるポリヌクレオチド - 前記第6染色体上の葉かび病抵抗性遺伝子座が、(c)、(d)および(e)のポリヌクレオチドで特定される、請求項11記載のトマト植物の葉かび病抵抗性マーカー。
- 前記第6染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_25325、solcap_snp_sl_25252、solcap_snp_sl_35221、SG_70、およびSL10401_823からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列を含む、請求項1から10のいずれか一項に記載のトマト植物の葉かび病抵抗性マーカー。
- 前記第6染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_25325およびSL10401_823の部位間の領域の塩基配列を含む、請求項13記載のトマト植物の葉かび病抵抗性マーカー。
- 第1染色体上の葉かび病抵抗性遺伝子座および第6染色体上の葉かび病抵抗性遺伝子座の少なくとも一方の葉かび病抵抗性遺伝子座を含み、
前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、
前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定されることを特徴とする、葉かび病抵抗性トマト植物。 - 前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160およびsolcap_snp_sl_8658で特定される、請求項15記載の葉かび病抵抗性トマト植物。
- 前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’で特定される、請求項15または16記載の葉かび病抵抗性トマト植物。
- 前記第1染色体上の葉かび病抵抗性遺伝子座が、下記(a)および(b)の少なくとも一方のポリヌクレオチドで特定される、請求項15から17のいずれか一項に記載の葉かび病抵抗性トマト植物。
(a) 下記(a1)のポリヌクレオチド
(a1)配列番号1の塩基配列からなるポリヌクレオチド
(b) 下記(b1)のポリヌクレオチド
(b1)配列番号2の塩基配列からなるポリヌクレオチド - 前記第1染色体上の葉かび病抵抗性遺伝子座が、前記(a)および(b)のポリヌクレオチドで特定される、請求項18記載の葉かび病抵抗性トマト植物。
- 前記第1染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_60250、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_60089からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列を含む、請求項15から17のいずれか一項に記載の葉かび病抵抗性トマト植物。
- 前記第1染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_60250およびsolcap_snp_sl_60089の部位間の領域の塩基配列を含む、請求項20記載の葉かび病抵抗性トマト植物。
- 前記葉かび病抵抗性トマト植物が、前記第1染色体上の葉かび病抵抗性遺伝子座および前記第6染色体上の葉かび病抵抗性遺伝子座を含む、請求項15から21のいずれか一項に記載の葉かび病抵抗性トマト植物。
- 前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_35221、およびSG_70で特定される、請求項15から22のいずれか一項に記載の葉かび病抵抗性トマト植物。
- 前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’で特定される、請求項15から23のいずれか一項に記載の葉かび病抵抗性トマト植物。
- 前記第6染色体上の葉かび病抵抗性遺伝子座が、(c)、(d)および(e)からなる群から選択された少なくとも一つのポリヌクレオチドで特定される、請求項15から24のいずれか一項に記載の葉かび病抵抗性トマト植物。
(c) 下記(c1)のポリヌクレオチド
(c1)配列番号3の塩基配列からなるポリヌクレオチド
(d) 下記(d1)のポリヌクレオチド
(d1)配列番号4の塩基配列からなるポリヌクレオチド
(e) 下記(e1)のポリヌクレオチド
(e1)配列番号5の塩基配列からなるポリヌクレオチド - 前記第6染色体上の葉かび病抵抗性遺伝子座が、(c)、(d)および(e)のポリヌクレオチドで特定される、請求項25記載の葉かび病抵抗性トマト植物。
- 前記第6染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_25325、solcap_snp_sl_25252、solcap_snp_sl_35221、SG_70、およびSL10401_823からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列を含む、請求項15から24のいずれか一項に記載の葉かび病抵抗性トマト植物。
- 前記第6染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_25325およびSL10401_823の部位間の領域の塩基配列を含む、請求項27記載の葉かび病抵抗性トマト植物。
- 前記葉かび病抵抗性トマト植物が、受託番号FERM BP-22282で特定されるトマト植物またはその後代系統である、請求項15から28のいずれか一項に記載の葉かび病抵抗性トマト植物。
- 前記葉かび病抵抗性トマト植物が、植物体またはその部分である、請求項15から29のいずれか一項に記載の葉かび病抵抗性トマト植物。
- 前記葉かび病抵抗性トマト植物が、種子である、請求項15から30のいずれか一項に記載の葉かび病抵抗性トマト植物。
- 下記(a)および(b)工程を含むことを特徴とする、葉かび病抵抗性トマト植物の製造方法。
(a)請求項15から31のいずれか一項に記載の葉かび病抵抗性トマト植物と、他のトマト植物とを交雑する工程
(b)前記(a)工程より得られたトマト植物またはその後代系統から、葉かび病抵抗性トマト植物を選抜する工程 - 前記(a)工程に先立って、下記(x)工程を含む、請求項32記載の葉かび病抵抗性トマト植物の製造方法。
(x)被検トマト植物から、請求項15から31のいずれか一項に記載の葉かび病抵抗性トマト植物を選抜する工程 - 前記(x)工程における前記選抜が、前記第1染色体上の葉かび病抵抗性遺伝子座および前記第6染色体上の葉かび病抵抗性遺伝子座の少なくとも一方を含む葉かび病抵抗性トマト植物の選抜であり、
前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、
前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定される、請求項33記載の葉かび病抵抗性トマト植物の製造方法。 - 前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160およびsolcap_snp_sl_8658で特定される、請求項34記載の葉かび病抵抗性トマト植物の製造方法。
- 前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’で特定される、請求項34または35記載の葉かび病抵抗性トマト植物の製造方法。
- 前記(x)工程における前記選抜が、前記第1染色体上の葉かび病抵抗性遺伝子座を含む葉かび病抵抗性トマト植物の選抜であり、
前記第1染色体上の葉かび病抵抗性遺伝子座が、下記(a)および(b)の少なくとも一方のポリヌクレオチドで特定される、請求項34から36のいずれか一項に記載の葉かび病抵抗性トマト植物の製造方法。
(a) 下記(a1)のポリヌクレオチド
(a1)配列番号1の塩基配列からなるポリヌクレオチド
(b) 下記(b1)のポリヌクレオチド
(b1)配列番号2の塩基配列からなるポリヌクレオチド - 前記第1染色体上の葉かび病抵抗性遺伝子座が、前記(a)および(b)のポリヌクレオチドで特定される、請求項37記載の葉かび病抵抗性トマト植物の製造方法。
- 前記(x)工程における前記選抜が、前記第1染色体上の葉かび病抵抗性遺伝子座を含む葉かび病抵抗性トマト植物の選抜であり、
前記第1染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_60250、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_60089からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列を含む、請求項34から36のいずれか一項に記載の葉かび病抵抗性トマト植物の製造方法。 - 前記第1染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_60250およびsolcap_snp_sl_60089の部位間の領域の塩基配列を含む、請求項39記載の葉かび病抵抗性トマト植物の製造方法。
- 前記(x)工程における前記選抜が、前記第1染色体上の葉かび病抵抗性遺伝子座および前記第6染色体上の葉かび病抵抗性遺伝子座を含む前記葉かび病抵抗性トマト植物の選抜である、請求項34から40のいずれか一項に記載の葉かび病抵抗性トマト植物の製造方法。
- 前記(x)工程における前記選抜が、前記第6染色体上の葉かび病抵抗性遺伝子座を含む葉かび病抵抗性トマト植物の選抜であり、
前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_35221、およびSG_70で特定される、請求項34から41のいずれか一項に記載の葉かび病抵抗性トマト植物の製造方法。 - 前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’で特定される、請求項34から42のいずれか一項に記載の葉かび病抵抗性トマト植物の製造方法。
- 前記(x)工程における前記選抜が、前記第6染色体上の葉かび病抵抗性遺伝子座を含む葉かび病抵抗性トマト植物の選抜であり、
前記第6染色体上の葉かび病抵抗性遺伝子座が、(c)、(d)および(e)からなる群から選択された少なくとも一つのポリヌクレオチドで特定される、請求項34から43のいずれか一項に記載の葉かび病抵抗性トマト植物の製造方法。
(c) 下記(c1)のポリヌクレオチド
(c1)配列番号3の塩基配列からなるポリヌクレオチド
(d) 下記(d1)のポリヌクレオチド
(d1)配列番号4の塩基配列からなるポリヌクレオチド
(e) 下記(e1)のポリヌクレオチド
(e1)配列番号5の塩基配列からなるポリヌクレオチド - 前記第6染色体上の葉かび病抵抗性遺伝子座が、(c)、(d)および(e)のポリヌクレオチドで特定される、請求項44記載の葉かび病抵抗性トマト植物の製造方法。
- 前記(x)工程における前記選抜が、前記第6染色体上の葉かび病抵抗性遺伝子座を含む葉かび病抵抗性トマト植物の選抜であり、
前記第6染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_25325、solcap_snp_sl_25252、solcap_snp_sl_35221、SG_70、およびSL10401_823からなる群から選択された2つのSNPマーカーの部位間の領域の塩基配列を含む、請求項34から43のいずれか一項に記載の葉かび病抵抗性トマト植物の製造方法。 - 前記第6染色体上の葉かび病抵抗性遺伝子座が、前記染色体における、solcap_snp_sl_25325およびSL10401_823の部位間の領域の塩基配列を含む、請求項46記載の葉かび病抵抗性トマト植物の製造方法。
- 交雑により葉かび病抵抗性トマト植物を生産するための親として、被検トマト植物から、トマト植物の葉かび病抵抗性マーカーとして第1染色体上の葉かび病抵抗性遺伝子座および第6染色体上の葉かび病抵抗性遺伝子座の少なくとも一方を含む葉かび病抵抗性トマト植物を選抜する工程を含み、
前記第1染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_60160、solcap_snp_sl_8658、およびsolcap_snp_sl_8658’からなる群から選択された少なくとも1つのSNPマーカーで特定され、
前記第6染色体上の葉かび病抵抗性遺伝子座が、solcap_snp_sl_25252、solcap_snp_sl_25252’、solcap_snp_sl_35221、SG_70、およびSG_70’からなる群から選択された少なくとも一つのSNPマーカーで特定されることを特徴とする、葉かび病抵抗性トマト植物のスクリーニング方法。
Priority Applications (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| KR1020177034434A KR102334176B1 (ko) | 2015-04-30 | 2015-12-11 | 토마토 식물의 잎곰팡이병 저항성 마커, 잎곰팡이병 저항성 토마토 식물, 잎곰팡이병 저항성 토마토 식물의 제조 방법 및 잎곰팡이병 저항성 토마토 식물의 스크리닝 방법 |
| EP15890778.2A EP3290526B1 (en) | 2015-04-30 | 2015-12-11 | Marker for leaf mold resistance in tomato plant, leaf mold-resistant tomato plant, method for producing leaf mold-resistant tomato plant, and method for screening for leaf mold-resistant tomato plant |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2015093528A JP6527746B2 (ja) | 2015-04-30 | 2015-04-30 | トマト植物の葉かび病抵抗性マーカー、葉かび病抵抗性トマト植物、葉かび病抵抗性トマト植物の製造方法、および葉かび病抵抗性トマト植物のスクリーニング方法 |
| JP2015-093528 | 2015-04-30 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2016174795A1 true WO2016174795A1 (ja) | 2016-11-03 |
Family
ID=57199074
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/JP2015/084750 Ceased WO2016174795A1 (ja) | 2015-04-30 | 2015-12-11 | トマト植物の葉かび病抵抗性マーカー、葉かび病抵抗性トマト植物、葉かび病抵抗性トマト植物の製造方法、および葉かび病抵抗性トマト植物のスクリーニング方法 |
Country Status (4)
| Country | Link |
|---|---|
| EP (1) | EP3290526B1 (ja) |
| JP (1) | JP6527746B2 (ja) |
| KR (1) | KR102334176B1 (ja) |
| WO (1) | WO2016174795A1 (ja) |
Cited By (2)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN112695031A (zh) * | 2021-01-26 | 2021-04-23 | 北京市农林科学院 | 一种鉴定番茄品种dna指纹的方法及其使用的snp引物组合 |
| CN113736908A (zh) * | 2021-09-30 | 2021-12-03 | 中国农业科学院农业基因组研究所 | 用于检测番茄叶霉病抗性的snp位点组合及其应用 |
Families Citing this family (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN110129478B (zh) * | 2019-05-21 | 2022-06-21 | 上海市农业科学院 | 一种快速鉴定或辅助鉴定番茄可溶性固形物的dCAPS分子标记及应用 |
Family Cites Families (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US7906712B2 (en) * | 2008-08-14 | 2011-03-15 | Enza Zaden Beheer B.V. | Tomato hybrid annamay |
-
2015
- 2015-04-30 JP JP2015093528A patent/JP6527746B2/ja active Active
- 2015-12-11 WO PCT/JP2015/084750 patent/WO2016174795A1/ja not_active Ceased
- 2015-12-11 EP EP15890778.2A patent/EP3290526B1/en active Active
- 2015-12-11 KR KR1020177034434A patent/KR102334176B1/ko active Active
Non-Patent Citations (4)
| Title |
|---|
| DATABASE GenBank 20 November 1998 (1998-11-20), DIXON,M.S. ET AL.: "DEFINITION: Lycopersicon pimpinellifolium Hcr2- 2A (Hcr2-2A) gene , complete cds.", XP055326503, Database accession no. AF053996 * |
| HAANSTRA, J.P .W. ET AL.: "Mapping strategy for resistance genes against Cladosporium fulvum on the short arm of Chromosome 1 of tomato: Cf- ECP5 near the Hcr9 Milky Way cluster", THEOR APPL GENET., vol. 1, no. 4, 10 September 2000 (2000-09-10), pages 661 - 668, XP055326498 * |
| See also references of EP3290526A4 * |
| SOUMPOUROU,E. ET AL.: "The Solanum pimpinellifolium Cf-ECP1 and Cf-ECP4 genes for resistance to Cladosporium fulvum are located at the Milky Way locus on the short arm of chromosome 1", THEOR APPL GENET., vol. 115, no. 8, 11 November 2007 (2007-11-11), pages 1127 - 1136, XP019558609 * |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN112695031A (zh) * | 2021-01-26 | 2021-04-23 | 北京市农林科学院 | 一种鉴定番茄品种dna指纹的方法及其使用的snp引物组合 |
| CN112695031B (zh) * | 2021-01-26 | 2023-03-14 | 北京市农林科学院 | 一种鉴定番茄品种dna指纹的方法及其使用的snp引物组合 |
| CN113736908A (zh) * | 2021-09-30 | 2021-12-03 | 中国农业科学院农业基因组研究所 | 用于检测番茄叶霉病抗性的snp位点组合及其应用 |
| CN113736908B (zh) * | 2021-09-30 | 2024-05-28 | 中国农业科学院农业基因组研究所 | 用于检测番茄叶霉病抗性的snp位点组合及其应用 |
Also Published As
| Publication number | Publication date |
|---|---|
| KR20170141235A (ko) | 2017-12-22 |
| KR102334176B1 (ko) | 2021-12-03 |
| EP3290526A1 (en) | 2018-03-07 |
| JP2016208880A (ja) | 2016-12-15 |
| JP6527746B2 (ja) | 2019-06-05 |
| EP3290526A4 (en) | 2018-12-05 |
| EP3290526B1 (en) | 2025-11-19 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| Blanc et al. | A reference genetic map of Muscadinia rotundifolia and identification of Ren5, a new major locus for resistance to grapevine powdery mildew | |
| Zhang et al. | Mapping of clubroot (Plasmodiophora brassicae) resistance in canola (Brassica napus) | |
| KR102156692B1 (ko) | 서양 호박 식물의 흰가루병 저항성 마커, 흰가루병 저항성 서양 호박 식물, 그것을 이용한 흰가루병 저항성 서양 호박 식물의 제조 방법 및 서양 호박 식물에 대한 흰가루병 저항성의 부여 방법 | |
| JP6412497B2 (ja) | トマト植物のネコブセンチュウ抵抗性マーカー、ネコブセンチュウ抵抗性トマト植物、ネコブセンチュウ抵抗性トマト植物の製造方法、およびネコブセンチュウ抵抗性トマト植物のスクリーニング方法 | |
| JP6811526B2 (ja) | メロン植物のうどんこ病抵抗性マーカー、うどんこ病抵抗性メロン植物、およびそれを用いたうどんこ病抵抗性メロン植物の製造方法 | |
| Esmenjaud et al. | Selection and application of resistant germplasm for grapevine nematodes management | |
| JP6527746B2 (ja) | トマト植物の葉かび病抵抗性マーカー、葉かび病抵抗性トマト植物、葉かび病抵抗性トマト植物の製造方法、および葉かび病抵抗性トマト植物のスクリーニング方法 | |
| JP6936626B2 (ja) | トマト植物の半身萎凋病抵抗性マーカー、半身萎凋病抵抗性トマト植物、およびそれを用いた半身萎凋病抵抗性トマト植物の製造方法 | |
| Singh et al. | Screening of cauliflower (Brassica oleracea L. var. botrytis L.) germplasm for resistance to downy mildew [Hyaloperonospora parasitica Constant (Pers.: Fr) Fr.] and designing appropriate multiple resistance breeding strategies | |
| Bordenave et al. | Wild vitis as a germplasm resource for rootstocks | |
| EP2966167B1 (en) | Prsv-resistant cucumber, and method for producing cucumber exhibiting resistance to prsv and zymv | |
| JP7239790B1 (ja) | ユーストマ立枯病抵抗性ユーストマ植物 | |
| JP7357173B1 (ja) | すすかび病抵抗性トマト植物 | |
| JP6153245B2 (ja) | 褐斑病抵抗性キュウリ植物、ならびに褐斑病抵抗性およびうどんこ病抵抗性を示すキュウリ植物の製造方法 | |
| US20250280784A1 (en) | Resistance to leveillula taurica in pepper | |
| JP2023049045A (ja) | ホモプシス根腐病抵抗性カボチャ植物 | |
| JP2019024483A (ja) | セイヨウカボチャ植物のうどんこ病抵抗性マーカー、うどんこ病抵抗性セイヨウカボチャ植物、それを用いたうどんこ病抵抗性セイヨウカボチャ植物の製造方法、およびセイヨウカボチャ植物へのうどんこ病抵抗性の付与方法 | |
| Ndeve | Genetics of resistance to root-knot nematode and Fusarium wilt In cowpea germplasm from Mozambique | |
| WO2023068371A1 (ja) | ユーストマ立枯病抵抗性ユーストマ植物 | |
| Moreau | Big-Bracted Dogwood Genetic Diversity and Powdery Mildew Disease Resistance Investigation Using Genomic Tools | |
| Rashid | Strategies for improving Brassica napus resistance to Leptosphaeria maculans, a causal agent of blackleg disease in canola | |
| 王素卿 | Genetic Resources for Disease Resistance Breeding in Wheat |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 15890778 Country of ref document: EP Kind code of ref document: A1 |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| ENP | Entry into the national phase |
Ref document number: 20177034434 Country of ref document: KR Kind code of ref document: A |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2015890778 Country of ref document: EP |
|
| WWG | Wipo information: grant in national office |
Ref document number: 2015890778 Country of ref document: EP |