WO2020142676A1 - Systems and methods for modulating rna - Google Patents
Systems and methods for modulating rna Download PDFInfo
- Publication number
- WO2020142676A1 WO2020142676A1 PCT/US2020/012169 US2020012169W WO2020142676A1 WO 2020142676 A1 WO2020142676 A1 WO 2020142676A1 US 2020012169 W US2020012169 W US 2020012169W WO 2020142676 A1 WO2020142676 A1 WO 2020142676A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- rna
- cirts
- domain
- protein
- hairpin
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
- XDTMQSROBMDMFD-UHFFFAOYSA-N C1CCCCC1 Chemical compound C1CCCCC1 XDTMQSROBMDMFD-UHFFFAOYSA-N 0.000 description 1
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/62—DNA sequences coding for fusion proteins
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/46—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- C07K14/47—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
- C07K14/4701—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals not used
- C07K14/4702—Regulators; Modulating activity
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/79—Vectors or expression systems specially adapted for eukaryotic hosts
- C12N15/85—Vectors or expression systems specially adapted for eukaryotic hosts for animal cells
- C12N15/86—Viral vectors
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/14—Hydrolases (3)
- C12N9/16—Hydrolases (3) acting on ester bonds (3.1)
- C12N9/22—Ribonucleases [RNase]; Deoxyribonucleases [DNase]
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/14—Hydrolases (3)
- C12N9/78—Hydrolases (3) acting on carbon to nitrogen bonds other than peptide bonds (3.5)
-
- A—HUMAN NECESSITIES
- A61—MEDICAL OR VETERINARY SCIENCE; HYGIENE
- A61K—PREPARATIONS FOR MEDICAL, DENTAL OR TOILETRY PURPOSES
- A61K38/00—Medicinal preparations containing peptides
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/01—Fusion polypeptide containing a localisation/targetting motif
- C07K2319/09—Fusion polypeptide containing a localisation/targetting motif containing a nuclear localisation signal
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/35—Fusion polypeptide containing a fusion for enhanced stability/folding during expression, e.g. fusions with chaperones or thioredoxin
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/85—Fusion polypeptide containing an RNA binding domain
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/20—Type of nucleic acid involving clustered regularly interspaced short palindromic repeats [CRISPR]
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/30—Chemical structure
- C12N2310/35—Nature of the modification
- C12N2310/351—Conjugate
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/30—Chemical structure
- C12N2310/35—Nature of the modification
- C12N2310/351—Conjugate
- C12N2310/3519—Fusion with another nucleic acid
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/50—Physical structure
- C12N2310/53—Physical structure partially self-complementary or closed
- C12N2310/531—Stem-loop; Hairpin
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2750/00—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA ssDNA viruses
- C12N2750/00011—Details
- C12N2750/14011—Parvoviridae
- C12N2750/14111—Dependovirus, e.g. adenoassociated viruses
- C12N2750/14141—Use of virus, viral particle or viral elements as a vector
- C12N2750/14143—Use of virus, viral particle or viral elements as a vector viral genome or elements thereof as genetic vector
Definitions
- the present invention relates generally to the field of chemistry and medicine. More particularly, it concerns the use of a system for modulating RNA.
- CRISPR/Cas9 system which evolved as a bacterial immune defense mechanism, has transformed the ability to study and manipulate cellular DNA site-specifically (Cong et al., 2013; Jiang et al., 2013; O’Connell et al., 2014; Wiedenheft et al., 2012).
- a key advantage of CRISPR/Cas systems compared to previous methods is that they are easily programmable to target virtually any locus of interest.
- the CRISPR/Cas system is a ribonucleoprotein complex that uses base pair interactions of a displayed guide RNA (gRNA) to interact with a target nucleic acid sequence.
- gRNA displayed guide RNA
- the simple nature of base pair-guided targeting opens up the possibility to program systems to interact with a defined nucleic acid sequence by simply changing the nucleic acid sequence on the guiding strand.
- RNA level presents several opportunities for therapeutic intervention, including but not limited to the ability to halt treatment if side effects emerge, the ability to target genes that would be too risky to alter at the DNA level, and the ability to manipulate gene expression without permanent alterations to the host genome.
- RNA epitranscriptomic regulatory mechanisms offer a broad range of RNA regulatory processes to target, including editing, degradation, transport, and translation of RNA transcripts (Nishikura, 2010; Roundtree et al., 2017; Zhao et al., 2017). Although the mechanisms and consequences of this epitranscriptomic regulatory layer are just beginning to be uncovered, it is apparent that the information flow through RNA is tightly regulated, offering many new opportunities for both basic research discoveries as well as therapeutic development.
- RNA-targeting tools analogous to the dCas9 DNA-targeting systems hold great promise for studying the mechanisms of epitranscriptomic regulation and for therapeutic applications.
- the current tools for RNA targeting involve the delivery of large complexes and pose immunogenicity issues. From a basic science perspective, the large size of the delivery vehicle could lead to potential perturbations to the RNA under interrogation, convoluting the study of RNA regulatory mechanisms. From a translational perspective, the large size presents challenges for viral packaging or direct protein delivery. Additionally, while DNA-editing therapies will likely consist of a one-time, irreversible treatment, RNA-targeting therapies will need to be continually administered, making delivery concerns especially important.
- CRISPR/Cas-inspired RNA targeting system a general method for engineering programmable RNA effector proteins.
- CIRTS is a ribonucleoprotein complex that uses Watson-Crick-Franklin base pair interactions to deliver protein cargo site-selectively in the transcriptome.
- the inventors show they can easily engineer CIRTS that deliver a range or regulatory proteins to transcripts, including nucleases for degradation, deadenylation regulatory machinery for degradation, or translational activation machinery for enhanced protein production.
- CIRTS are up to 5-fold smaller than the smallest current CRISPR/Cas systems and can be engineered entirely from human parts.
- RNA regulatory system or method comprising at least one of each: i) a RNA hairpin binding domain; ii) a RNA targeting molecule comprising a RNA targeting region and at least one hairpin structure, wherein the hairpin structure of the RNA targeting molecule specifically binds to i; and iii) a RNA regulatory domain.
- the following are included: i) and ii), i) and iii), ii) and iii), or i), ii), and iii). Any embodiment disclosed herein can contain any of these combinations.
- RNA hairpin binding domain a nucleotide encoding: i) a RNA hairpin binding domain; ii) a RNA targeting molecule comprising a RNA targeting region and at least one hairpin structure, wherein the hairpin structure of the RNA targeting molecule specifically binds to i), and iii) a RNA regulatory domain.
- fusion protein comprising a RNA hairpin binding protein and a RNA regulatory domain and nucleic acids encoding such fusion proteins.
- RNA regulatory domain operably linked to a RNA targeting molecule
- the RNA targeting molecule comprises a RNA targeting region and at least one hairpin structure.
- the RNA regulatory domain and the RNA targeting molecule are operably linked through a peptide bond.
- the polypeptide further comprises one or more linkers.
- the RNA regulatory domain and the RNA targeting molecule are operably linked through non- covalent interactions.
- the RNA regulatory domain is covalently linked to a first dimerization domain and the RNA targeting molecule is covalently linked to a second dimerization domain and wherein the first and second dimerization domain are capable of dimerizing to form a non-covalent or covalent linkage.
- the conjugate comprises one or more nuclear localization signals (NLS)s.
- NLS nuclear localization signals
- Yet further aspects relate to a delivery vehicle comprising a system of the disclosure.
- the delivery vehicle comprises liposome(s), particle(s), exosome(s), microvesicle(s), a gene-gun or one or more nucleic acid vector(s).
- compositions or a cell comprising a system, delivery vehicle, or fusion protein of the disclosure.
- modulating at least one target RNA comprises cleaving, demethylating, methylating, activating translation, repressing translation, promoting degradation, or binding to the RNA.
- at least two target RNAs are modulating.
- at least 2, 3, 4, 5, 6, 7, 8, 9, 10, 15, 20, or 25 (or any derivable range therein) target RNAs are modulated.
- the multiple RNAs are modulated by the same RNA regulatory domain or by a regulatory domain with the same activity.
- the different target RNAs are modulated with a different activity, such as by cleaving, demethylating, methylating, activating translation, repressing translation, promoting degradation, or binding to the RNA.
- the RNA regulatory domain does not bind to RNA.
- the RNA regulatory domain comprises a polypeptide that does not have RNA binding activity.
- the RNA regulatory domain does not bind to modified RNA.
- the RNA regulatory domain does not bind to m6A modified RNA.
- Further aspects relate to a cell or progeny thereof comprising modulated target RNA, wherein the target RNA has been modulated according to a method of the disclosure. Further aspects relate to a multicellular organism comprising one or more cells of the disclosure. Further aspects relate to a plant or animal comprising one or more cells of the disclosure. Further aspects relate to a kit comprising a system, vector, delivery vehicle, or fusion protein of the disclosure.
- Further aspects relate to a method for modulating a target RNA in a subject, the method comprising administering a system or composition of the disclosure to the subject.
- RNA hairpin refers to a RNA molecule with stem-loop intramolecular base pairing.
- a hairpin can occur when two regions of the same strand, usually complementary in nucleotide sequence when read in opposite directions, base-pair to form a double helix that ends in an unpaired loop.
- the disclosure relates to engineered RNA targeting molecules comprising a RNA targeting region and one or more hairpins. Accordingly, the engineered RNA molecules of the disclosure are chimeric molecules that are non-naturally occurring.
- RNA targeting region refers to a region of the RNA that is capable of hybridizing to a target RNA.
- the target RNA may be a disease associated RNA or one that is a modulation target according to the current systems and methods.
- RNA regulatory domain refers to a peptide or polypeptide that has activity directed to RNA.
- activity include methylation activity, RNA-binding activity, nuclease activity, and translational activation or repression activity. Further examples of activities and proteins comprising RNA regulatory domains are described throughout the disclosure.
- the RNA hairpin binding domain and the RNA regulatory domain are operably linked.
- the term“operably linked” refers to two proteins that are linked through either covalent or non-covalent interactions.
- the two proteins may be covalently linked through a peptide bond.
- the proteins are non- covalently linked.
- One or more proteins of the disclosure may be operably linked to another protein through linkage to a pair of accessory proteins that have a strong affinity for each other.
- accessory proteins are known in the art.
- the SunTag is one such system that includes an antibody with a strong affinity for a peptide.
- One protein, polypeptide, or domain of the disclosure may be linked to a SunTag peptide and another protein, polypeptide, or domain of the disclosure may be linked to an antibody to allow operable linkage of the two proteins, polypeptides, or domains through the interaction of the SunTag peptide and antibody.
- Further examples include biotin and avidin/streptavidin and spytag and spycatcher.
- the system is inducible by providing the RNA regulatory domain and the hairpin binding domain as two unlinked polypeptides that become linked upon the presence of a stimulant.
- the induction may be, for example, by light induction or by chemical induction. Such inducibility allows for activation of the RNA regulation at a desired moment in time.
- the RNA regulatory domain is covalently linked to a first dimerization domain and the RNA hairpin binding domain is covalently linked to a second dimerization domain and wherein the first and second dimerization domain are capable of dimerizing to form a non-covalent or covalent linkage.
- the dimerization is inducible.
- the dimerization is induced through binding of the dimerization domains to a ligand.
- the term inducible refers to dimerization that is formed in response to a stimulus, such as a ligand, a chemical, a temperature change, or light, for example.
- Light inducibility is for instance achieved by designing a fusion complex wherein the first and second dimerization domains comprise CRY2PHR and CIBN. This system is particularly useful for light induction of protein interactions in living cells and is further described in Konermann S, et al. Nature. 2013;500:472-476, which is herein incorporated by reference.
- Suitable dimerization domains and corresponding ligands are known in the art. For example, Liang, F.S., Ho, W.Q., and Crabtree, G.R. (2011). Engineering the ABA plant stress pathway for regulation of induced proximity. Sci. Signal. 4, rs2, which is incorporated by reference, describes suitable dimerization/ligand systems that are useful in embodiments of the disclosure.
- one of the first or second dimerization domain comprises PYR/PYRl-like (PYL1)
- the other of the first or second domain comprises ABA insensitive 1 (ABI1)
- the ligand comprises abscisic acid (ABA) or derivatives or fragments thereof.
- the dimerization domain may be a fragment or portion of the whole protein and may be a substituted or modified.
- the first and/or second dimerization domain comprises FKBP12 and the ligand comprises FK1012 or derivatives or fragments thereof.
- one of the first or second dimerization domain comprises FK506 binding protein (FKBP)
- the other of the first or second domain comprises FKBP-Rap binding domain of mammalian target of Rap mTOR (Frb)
- the ligand comprises rapamycin (Rap) or derivatives or fragments thereof.
- Derivatives refer to modifiied ligands and domains that retain binding or have enhanced binding to their dimerization domain or ligand, respectively. Fragments refer to contiguous portions of the dimerization domains that retain binding to the ligand. In some embodiments, the dimerization domain may be a modified fragment.
- i, ii, and/or iii are human or are human-derived.
- the system, conjugate, and/or fusion protein is non-immunogenic.
- a human protein, polypeptide, domain, or nucleic acid refers to a protein, polypeptide, domain, or nucleic acid that is from the human genome, although it may be produced recombinantly in non-human systems.
- the term“human-derived” refers to a protein, polypeptide, domain, or nucleic acid that is a variant or fragment of a protein, polypeptide, domain, or nucleic acid from the human genome, although it may be produced recombinantly in non-human systems.
- the fusion protein, conjugate, system, or parts thereof, such as parts i, ii, and/or iii are non-immunogenic and/or non-toxic when expressed in or administered to humans.
- the nucleic acids or polypeptides of the disclosure are synthetic, are non-natural, and/or do not occur naturally in nature.
- the system further comprises a stabilizer polypeptide; wherein the stabilizer polypeptide comprises a cationic polypeptide that binds non-specifically to nucleic acids.
- the stabilizer polypeptide is human-derived.
- the stabilizer polypeptide is operably linked to the RNA regulatory domain and/or RNA hairpin binding domain.
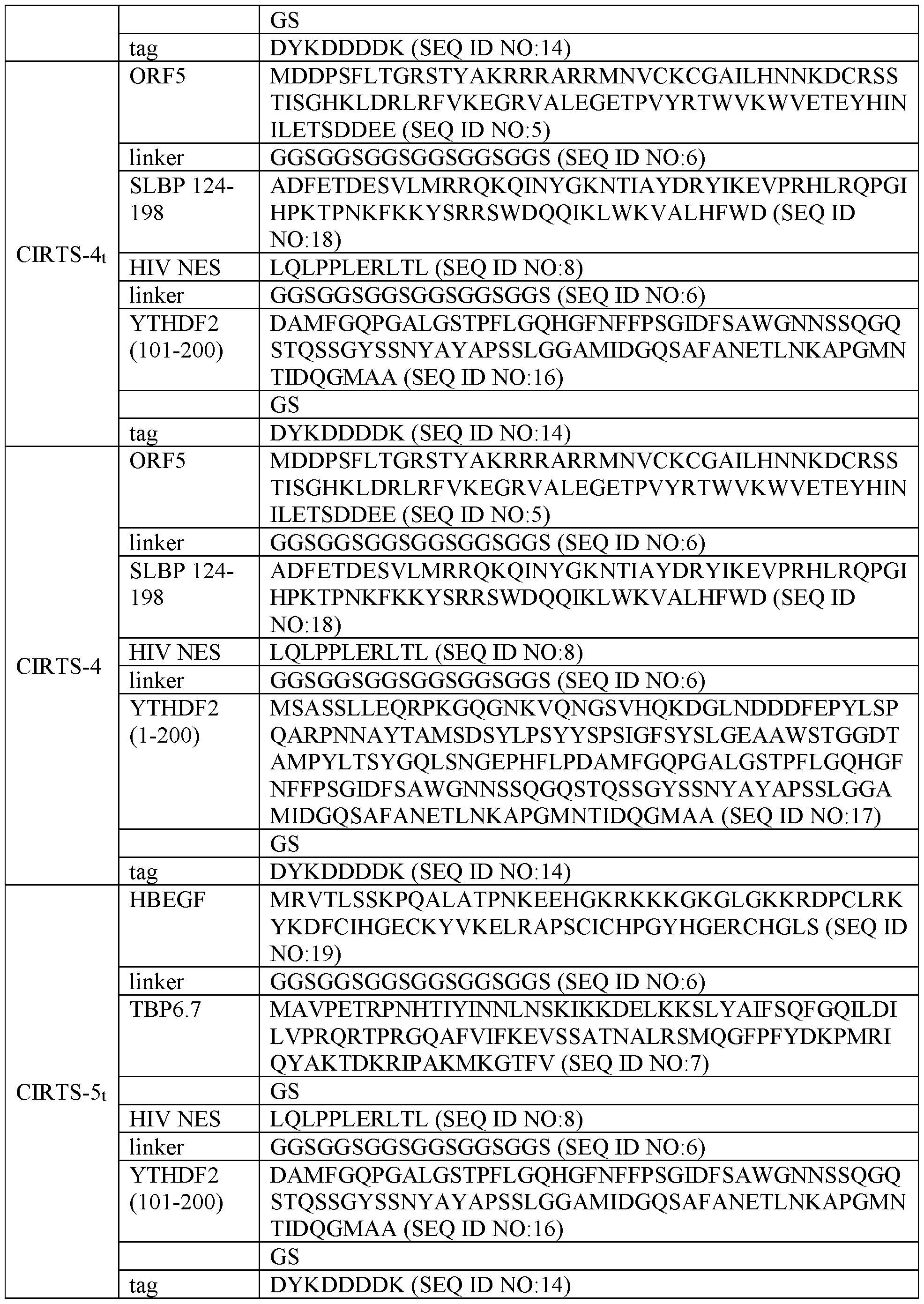
- the stabilizer polypeptide comprises ORF5 or a fragment thereof.
- the stabilizer polypeptide comprises SEQ ID NO:5, a variant thereof, or a polypeptide with at least 60, 61, 62, 63, 64, 65,
- the stabilizer polypeptide comprises HEBGF or a fragment thereof.
- the stabilizer polypeptide comprises SEQ ID NO: 19, a variant thereof, or a polypeptide with at least 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 76, 77, 78, 79, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% (or any derivable range therein) identity or homology to SEQ ID NO: 19.
- the stabilizer polypeptide comprises b-defensin 3 or a fragment thereof.
- the stabilizer polypeptide comprises SEQ ID NO:20, a variant thereof, or a polypeptide with at least 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 76, 77, 78, 79, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% (or any derivable range therein) identity or homology to SEQ ID NO:20.
- the stabilizer polypeptide, conjugate, fusion protein, conjugate, RNA regulatory domain, and/or RNA hairpin binding domain are less than, more than, or are at most or at least 175, 170, 165, 160, 155, 150, 145, 140, 135, 130, 125, 120, 115, 110, 105, 100, 95, 90, 85, 80, 75, 70, 65, 60, 55, 50, 45, 40, 35, 30, 25, 20, 15, 10, or 5 kDa (or any derivable range therein).
- the total complex comprising the RNA regulatory domain and hairpin binding domain is less than, more than, or is at most or at least 175, 170, 165, 160, 155, 150, 145, 140, 135, 130, 125, 120, 115, 110, 105, 100, 95, 90, 85, 80,
- the total complex comprising the stabilizer polypeptide, RNA regulatory domain and hairpin binding domain is less than, more than, or is at most or at least
- the RNA hairpin binding domain comprises a RNA hairpin binding domain from U1A (TBP6.7), SLBP, or variants thereof. In some embodiments, the RNA hairpin binding domain comprises a RNA hairpin binding domain from U1A (TBP6.7), SLBP, Ku70, nucleolin, or variants thereof. In some embodiments, the RNA hairpin binding domain comprises SEQ ID NO: 7 or 18, a variant thereof, or a polypeptide with at least 60, 61,
- the RNA targeting molecule comprises a TAR hairpin scaffold.
- the RNA targeting molecule comprises the TAR hairpin scaffold of SEQ ID NO: 1 or a nucleotide with at least 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 76, 77, 78, 79, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, or 99% (or any derivable range therein) identity to SEQ ID NO: 1.
- the RNA targeting molecule comprises a SLBP hairpin scaffold.
- the RNA targeting molecule comprises the SLBP hairpin scaffold of SEQ ID NO:2 or a nucleotide with at least
- the RNA targeting molecule comprises exactly one hairpin. In some embodiments, the RNA targeting molecule comprises at least one hairpin. In some embodiments, the RNA targeting molecule comprises exactly two hairpins. In some embodiments, the RNA targeting molecule comprises at least two hairpins. In some embodiments, the RNA targeting molecule comprises exactly three hairpins. In some embodiments, the RNA targeting molecule comprises at least three hairpins. In some embodiments, the RNA targeting molecule comprises exactly four hairpins. In some embodiments, the RNA targeting molecule comprises at least four hairpins. In some embodiments, the RNA targeting molecule comprises exactly five hairpins. In some embodiments, the RNA targeting molecule comprises at least five hairpins.
- the RNA targeting molecule comprises 1-4 hairpins. In some embodiments, the RNA targeting molecule comprises 1-3 hairpins. In some embodiments, the RNA targeting molecule comprises 1-2 hairpins. In some embodiments, the RNA targeting molecule comprises 2-4 hairpins. In some embodiments, the RNA targeting molecule comprises 2-3 hairpins. In some embodiments, the RNA targeting molecule comprises at least, at most, or exactly 1, 2, 3, 4, 5, or 6 hairpins (or any range derivable therein). In some embodiments, the RNA targeting molecule comprises at least one hairpin that does not bind to the RNA hairpin binding protein and at least one hairpin that binds to the RNA hairpin binding protein.
- the RNA targeting molecule binds to more than one RNA binding protein. In some embodiments, the RNA targeting molecule comprises two, three, or four hairpin structures and binds to at least two RNA binding proteins. In some embodiments, the RNA regulatory system comprises at least two regulatory domains, wherein each regulatory domain binds to a different RNA binding molecule.
- the RNA targeting molecule comprises one or more modified nucleotides.
- the modified nucleotides comprise a modification such as a phosphorothioate, locked nucleotides, ethylene bridged nucleotides, peptide nucleic acids, 5’E-VP, or is modified to a morpholino.
- the modification includes one described herein.
- the RNA hairpin binding domain comprises the RNA hairpin binding domain of U1 A, a variant thereof, or a polypeptide with at least 60, 61, 62, 63, 64, 65,
- RNA targeting molecule comprises a TAR hairpin scaffold or a nucleotide with at least 60, 61, 62,
- the RNA hairpin binding domain comprises the RNA hairpin binding domain of SLBP, a variant thereof, or a polypeptide with at least 60, 61, 62, 63, 64,
- RNA targeting molecule comprises a SLBP hairpin scaffold or a nucleotide with at least 60,
- the RNA hairpin binding domain comprises the RNA hairpin binding domain of ku70 or a variant thereof
- the RNA targeting molecule comprises a hairpin scaffold or a nucleotide with at least 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 76, 77, 78, 79, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, or 100% (or any derivable range therein) identity to SEQ ID NO: 83.
- the RNA hairpin binding domain comprises the RNA hairpin binding domain of nucleolin or a variant thereof
- the RNA targeting molecule comprises a hairpin scaffold or a nucleotide with at least 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 76, 77, 78, 79, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, or 100% (or any derivable range therein) identity to one of SEQ ID NO:84-86.
- the RNA hairpin binding domain, stabilizer polypeptide, or RNA hairpin binding domain comprises a linker.
- the linker comprises a polypeptide comprising SEQ ID NO:6, 21, 22, 23, or 25 or a polypeptide with at least 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 76, 77, 78, 79, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94
- the linker is a rigid linker. In some embodiments, the linker is a flexible linker. In some embodiments, the linker comprises glycine and serine residues. In some embodiments, the linker is at least 4 amino acids. In some embodiments, the linker is at least or at most or exactly 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18,
- the stabilizer polypeptide comprises a polypeptide, such as a RNA-binding polypeptide, from cJun, HBEGF, HRX, NDEK, NHGF, beta-defensin3, or scGFP.
- the RNA regulatory domain is operably linked to the stabilizer polypeptide at the carboxy terminus of the RNA regulatory domain.
- the RNA regulatory domain is operably linked to the stabilizer polypeptide at the amino terminus of the RNA regulatory domain.
- the RNA regulatory domain is operably linked to the RNA hairpin binding domain polypeptide at the carboxy terminus of the RNA regulatory domain.
- the RNA regulatory domain is operably linked to the RNA hairpin binding domain polypeptide at the amino terminus of the RNA regulatory domain. In some embodiments, the RNA hairpin binding domain polypeptide is operably linked to the stabilizer polypeptide at the carboxy terminus of the RNA hairpin binding domain polypeptide. In some embodiments, the RNA hairpin binding domain polypeptide is operably linked to the stabilizer polypeptide at the amino terminus of the RNA hairpin binding domain polypeptide.
- the RNA targeting region comprises at least 12 nucleotides. In some embodiments, the RNA targeting region comprises at least, at most, or exactly 8, 9,
- nucleotides 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, or 79 nucleotides (or any derivable range therein).
- the RNA regulatory domain comprises a nuclease, methylase, demethylase, translational activator, translational repressor, single-stranded RNA cleavage activity, double-stranded RNA cleavage activity, or RNA binding activity. In some embodiments, the RNA regulatory domain comprises an activity described herein.
- the RNA regulatory domain comprises a Pin nuclease domain or a m6A reader protein or portion thereof. In some embodiments, the RNA regulatory domain comprises a domain or polypeptide from SMG6, YTHDF1, or YTHDF2. In some embodiments, the RNA regulatory domain comprises a domain or polypeptide from an ADAR protein. In some embodiments, the RNA regulatory domain comprises a domain or polypeptide from a human ADAR protein.
- the RNA regulatory domain comprises a polypeptide that has at least, at most, or exactly 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 76, 77, 78, 79, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, or 100% (or any derivable range therein) identity or homology to SEQ ID NO:9, 11, 15, 16, 17, or 123-125.
- the RNA regulatory domain further comprises a helical region.
- the helical region comprises a polypeptide that has at least, at most, or exactly 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 76, 77, 78, 79, 80, 85, 86, 87, 88, 89, 90, 91, 92,
- the RNA regulatory domain increases translation of a target RNA. In some embodiments, the RNA regulatory domain increases degradation of a target RNA. In some embodiments, the RNA regulatory domain modifies the localization of a target RNA. In some embodiments, the RNA regulatory domain modifies the processing of the target RNA.
- the RNA regulatory domain comprises a polypeptide, such as a polypeptide having RNA regulatory activity from IFIT2, eIF4a, eIF4e, PABP, PAIP, SLBP, BOLL, ICP27, YTHDF1, YTHDF2, or YTHDF3.
- the RNA regulatory domain comprises a polypeptide, such as a polypeptide having RNA regulatory activity from YTHDF2, TOB2, ZFP36, CNOT7, RNaseA, RNaseL, RNaseP, RNase4, RNasel, RNaseU2, or HRSP12.
- the RNA regulatory domain increases the expression of a polypeptide encoded by the target RNA and wherein the RNA regulatory domain comprises IFIT2, eIF4a, eIF4e, PABP, PAIP, SLBP, BOLL, ICP27, YTHDF1, or YTHDF3.
- the RNA regulatory domain comprises a polypeptide, such as a polypeptide having RNA regulatory activity from YTHDF2, TOB2, ZFP36, CNOT7, RNaseA, RNaseL, RNaseP, RNase4, RNasel, RNaseU2, or HRSP12.
- the RNA regulatory domain decreases the expression of a polypeptide encoded by the target RNA and wherein the RNA regulatory domain comprises YTHDF2, TOB2, ZFP36, CNOT7, RNaseA, RNaseL, RNaseP, RNase4, RNasel, RNaseU2, or HRSP12.
- one or more nuclear export signals are fused to the RNA regulatory domain, the RNA hairpin binding domain, and/or the stabilizing polypeptide.
- the NES is at the carboxy terminus of the RNA regulatory domain, the RNA hairpin binding domain, and/or the stabilizing polypeptide.
- the NES is at the amino terminus of the RNA regulatory domain, the RNA hairpin binding domain, and/or the stabilizing polypeptide.
- the NES comprises a polypeptide that has at least, at most, or exactly 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 76, 77, 78, 79, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, or 100% (or any derivable range therein) identity or homology to SEQ ID NO: 8.
- one or more nuclear localization signals are fused to the RNA regulatory domain, the RNA hairpin binding domain, and/or the stabilizing polypeptide.
- the NLS is at the carboxy terminus of the RNA regulatory domain, the RNA hairpin binding domain, and/or the stabilizing polypeptide.
- the NLS is at the amino terminus of the RNA regulatory domain, the RNA hairpin binding domain, and/or the stabilizing polypeptide.
- the NES comprises a polypeptide that has at least, at most, or exactly 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 75, 76, 77, 78, 79, 80, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, or 100% (or any derivable range therein) identity or homology to SEQ ID NO: 13.
- the RNA targeting region of ii hybridizes to a target RNA in a prokaryotic or eukaryotic cell.
- the target RNA is in a human cell.
- the target RNA is in vitro or in vivo.
- the system comprises at least two of each i, ii, and iii. In some embodiments, the at least two of i, ii, and iii are expressed in the same cell. In some embodiments, the method comprises modulating at least two target RNAs. In some embodiments, the system comprises at least, at most, or exactly 2, 3, 4, 5, 6, 7, 8, 9, 10, 20, 30, 40, or 50 or more (or any derivable range therein) of i, ii, and iii. In some embodiments, at least, at most, or exactly 2, 3, 4, 5, 6, 7, 8, 9, 10, 20, 30, 40, or 50 or more (or any derivable range therein) target RNAs are modulated in a cell.
- the RNA regulatory domain cleaves RNA, promotes RNA translation, inhibits RNA translation, or modifies the base sequence of RNA.
- the vectors of the disclosure further comprise a regulatory element operably linked to the nucleotide encoding i, ii, and/or iii.
- Regulatory elements in addition to a NLS and NES, as previously described, also include promoters, polyadenylation signals, enhancers, etc.
- Other regulatory elements are known in the art and described herein and may be used in the embodiments of the disclosure.
- the one or more nucleic acid vectors are optimized for expression in an eukaryotic cell.
- the expression of the domains, RNA, or polypeptides in the cell or from a vector is constitutive.
- the expression of the domains, RNA, or polypeptides in the cell or from a vector is conditional.
- i, ii, and iii are on a single vector.
- i, ii, iii, and the stabilizer polypeptide are encoded on a single vector.
- i, iii, and the stabilizer polypeptide are encoded on a single vector.
- one or more of the vectors are viral vectors.
- the one or more vectors comprise one or more retroviral, lentiviral, adenoviral, adeno-associated or herpes simplex viral vectors.
- one or more of the vectors are non-viral vectors.
- the system or composition is non-viral, which denotes that it does not contain any viral components.
- a system or kit comprising one or more of the following components: a polypeptide comprising a RNA regulatory domain, a polypeptide comprising a RNA binding domain, a polypeptide comprising a stabilizer, a nucleic acid encoding for a RNA regulatory domain, a nucleic acid encoding for a RNA binding domain, a nucleic acid encoding a stabilizer, a nucleic acid encoding a RNA targeting molecule comprising a RNA targeting region and at least one hairpin structure; a conjugate of the disclosure; a vector of the disclosure, a fusion protein of the disclosure, a recombinant host cell, an expression construct, an engineered viral vector, or an engineered attenuated virus.
- a polypeptide of the disclosure is under the control of a heterologous promoter. It is specifically contemplated that any protein or polypeptide function that are used in embodiments, may be used a nucleic acid encoding that protein or polypeptide function. Also, any and all polypeptides, proteins, nucleic acid molecules may be contained within a cell or other living organism, such as a virus (for instance, a phage).
- a kit may include one or more components that are separate or together in a suitable container means, such as a sterile, non-reactive container.
- a suitable container means such as a sterile, non-reactive container.
- cells or viruses are provided that contain one or more nucleic acid constructs that encode the polypeptides of the disclosure.
- the term“promoter” is used according to its ordinary meaning to those in the field of molecular biology; it generally refers to a site on a nucleic acid in which a polymerase can bind to initiate transcription. In specific embodiments, the promoter is recognized by a T7 RNA polymerase.
- compositions, vectors, systems, methods, and proteins of the disclosure are useful for a variety of clinical and research-related applications.
- the embodiments of the disclosure may be useful for the treatment of a disease or condition, such as cancer or autoimmunity.
- the methods and compositions are for the acute treatment of a disease or condition.
- the methods and compositions are useful for the temporary modulation of RNA.
- the methods and compositions are safer due to the acute modulation of RNA and/or due to the ability to control the expression of the system in vivo.
- “a” or“an” may mean one or more.
- the words“a” or“an” when used in conjunction with the word“comprising”, may mean one or more than one.
- the terms“or” and“and/or” are utilized to describe multiple components in combination or exclusive of one another.
- “x, y, and/or z” can refer to“x” alone,“y” alone,“z” alone,“x, y, and z,”“(x and y) or z,”“x or (y and z),” or“x or y or z.” It is specifically contemplated that x, y, or z may be specifically excluded from an embodiment.
- any limitation discussed with respect to one embodiment of the invention may apply to any other embodiment of the invention.
- any composition of the invention may be used in any method of the invention, and any method of the invention may be used to produce or to utilize any composition of the invention.
- Aspects of an embodiment set forth in the Examples are also embodiments that may be implemented in the context of embodiments discussed elsewhere in a different Example or elsewhere in the application, such as in the Summary of Invention, Detailed Description of the Embodiments, Claims, and Description of Figure Legends.
- FIG. 1A-D Design of CRISPR/Cas-inspired RNA targeting system (CIRTS)
- CIRTS is composed of a ssRNA binding protein an RNA hairpin binding protein, an effector protein and a guiding RNA.
- B List of modular CIRTS constructs used in this work.
- C Design of the guiding RNA for TBP6.7. The HIV TAR hairpin was fused to a nucleotide linker (L)and a guide sequence.
- D Design of the guiding RNA for the RRM of SLBP. The human histone mRNA hairpin was fused to a flexible five nucleotide linker and a guide sequence.
- FIG. 2A-C CIRTS-1 in vitro binding and RNA cleavage assays
- Electrophoretic mobility assay evaluating the binding affinity of MBP-CIRTS- Lon- target gRNA (R3) complex to a labeled RNA substrate (Rl). EDTA was supplemented to the reaction buffer to avoid any cleavage.
- B Calculation of the binding affinity by fitting the fraction of TBP6.7:gRNA bound to substrate to a quadratic binding equation.
- C Cleavage assay run on a denaturing gel after 2h of incubation. A labeled RNA substrate (R2) is cleavage in a gRNA-dependent manner.
- FIG. 3A-G CIRTS mammalian cell reporter assays
- A General overview of the dual luciferase assay. A reporter construct that contains both firefly luciferase and Renilla luciferase is used in all assays. The inventors targeted their CIRTS to the firefly luciferase transcript, while keeping Renilla luciferase constant to use as a transfection control. For all subsequent assays, HEK293T cells were transfected with the reporter vector, the CIRTS vector, and a gRNA vector.
- B Catalytically-inactive CIRTS-0 was used as a control.
- C Comparison of CIRTS- 1 with Casl3b nuclease. Cells transfected with either CIRTS-1 or Casl3 and the corresponding gRNA targeting show reduced protein levels after incubation.
- D HEK293T cells transfected with CIRTS-2 show an increase in protein level after 48h
- E cells transfected with CIRTS-3, however, show the anticipated decrease in protein level.
- F Switching the hairpin-binding protein to SLBP still results in decrease protein levels after 48h.
- G Cells transfected with a fully humanized CIRTS (CIRTS-5 and CIRTS-6) system and an on-target gRNA for firefly luciferase result again in decreased protein levels.
- FIG. 4A-C Targeting endogenous transcripts with CIRTS
- A Nuclease- mediated knockdown of five endogenous transcripts upon transfection of cells with CIRTS- 1 as assayed using qPCR.
- CIRTS- 1 can be used to target endogenous transcripts of interest by co-transfecting a gRNA the corresponding on-target guiding sequence.
- B qPCR analysis of non-nuclease-mediated knockdown of endogenous transcripts with CIRTS-3. Cells transfected with CIRTS-3 show gRNA-dependent decreases in RNA level for all five transcripts tested.
- C Analysis of protein levels after transfection with CIRTS-2 or CIRTS-3 using Western blot. CIRTS-2 can induce an increase in protein levels, whereas CIRTS-3 shows the expected decrease in protein levels as a control.
- FIG. 5A-B Multidimensional targeting with CIRTS.
- A Schematic of vectors used for multiplexed targeting.
- CIRTS-6 was co-transfected with its gRNA construct targeting PPIB
- CIRTS-7 was co-transfected with the corresponding gRNA construct targeting SMARCA4.
- B Heat map showing knockdown of multiplexed targeting.
- co-transfection of either on-target or off-target control gRNA can guide the CIRTS to decrease endogenous transcripts.
- both CIRTS have an on-target gRNA for PPIB or SMARCA4 present, both transcripts can be knocked down in the same samples.
- FIG. 6 Comparing CIRTS to other DNA and RNA-targeting CRISPR/Cas systems. Schematic size comparison of currently used Cas9, Casl3, and fusion protein systems. Cas9 and Casl3-based delivery systems are substantially larger than engineered CIRTS systems.
- FIG. 7A-C Controls EMSA and Cleavage Assay.
- A EMSA evaluating binding shifts dependent on MBP-CIRTS-1 in the absence of any gRNA.
- B EMSA assaying binding shifts in the presence of only labeled substrate (Rl) and on-target gRNA (R3).
- C Full cleavage gel shown in main text FIG. 2C.
- FIG. 8A-D CIRTS linker and gRNA optimization.
- A Luciferase assay with the CIRTS nuclease system using different linkers between the hairpin-binding protein and the effector protein.
- B Luciferase assay with CIRTS-YTHDF2-mediated decay using different linkers between the hairpin-binding protein and the effector protein.
- C Different engineered gRNA for TBP6.7 based on the design shown in FIG. 1C. Two different targeting lengths of 20 and 40 nucleotides were used in combination with different numbers of linking nucleotides (L) between the hairpin and the guiding sequence. The dual luciferase assay was used to assess nuclease-mediated decay.
- FIG. 9A-H Control luciferase assays and RT-qPCRs.
- A RT-qPCR analysis of RNA levels with the‘dead’ Pin nuclease domain CIRTS (CIRTS-0).
- B Luciferase assay comparing the nuclease-mediated decay of TBP6.7-Pin nuclease domain without (CIRTS-8) and with (CIRTS-9) the additional ssRNA binding protein ORF5.
- FIG. 10A-C Immunoprecipitation, Control qPCR Western Blot, Y2 truncations.
- B RT-qPCR analysis of RNA level when cells were transfected with CIRTS-2. As anticipated, no significant changes in RNA level were observed when a YTHDF1 -containing protein was used.
- FIG. 11A-C Endogenous targeting with CIRTS.
- A Changes in RNA levels assessed after transfection of CIRTS5-7 alone for PPIB.
- B Similar to FIG. 11 A, SMARCA4 levels were assayed when cells were transfected with CIRTS5-7.
- FIG. 12A-D Design of CRISPR/Cas-inspired RNA targeting system (CIRTS)
- CIRTS is composed of a ssRNA binding protein, an RNA hairpin binding protein, an effector protein, and a guiding RNA.
- B List of key CIRTS used in this work.
- D Design of the guiding RNA for the RNA recognition motif (RRM) of SLBP. The human histone mRNA hairpin was fused to a flexible five nucleotide linker and a guide sequence.
- RRM RNA recognition motif
- FIG. 13A-B CIRTS-1 in vitro binding and RNA cleavage assays.
- A Filter binding assay evaluating the binding affinity of MBP-CIRTS-1 with on-target gRNA and non targeting RNA complex to a labeled RNA substrate. Fitting the data to a quadratic binding equation revealed an apparent KD of 22 ⁇ 7 nM for the on-target:protein complex and an apparent KD around 500 nM for the non-targeting: protein interaction.
- B Cleavage assay run on a 10% denaturing Urea PAGE gel in presence of 0.5 mM MnCb. An IR800-labeled RNA substrate is cleaved in a gRNA-dependent manner.
- FIG. 14A-G CIRTS mammalian cell reporter assays.
- A General overview of the dual luciferase assay. A reporter construct that contains both firefly luciferase and Renilla luciferase is used in all assays. The inventors targeted CIRTS to the firefly luciferase transcript, while using Renilla luciferase as an internal control. For all subsequent assays, HEK293T cells were transfected with the reporter vector, a CIRTS vector, and a gRNA vector.
- B Catalytically-inactive CIRTS-0 was used as a control. After 48 h of incubation, the inventors observed no decrease in protein readout.
- FIG. 15A-B CIRTS for RNA editing.
- B Delivery of CIRTS-7 (hADAR2 wt) and CIRTS-8 (hADAR E488Q) with an on-target gRNA shows significant RNA editing that results in measurable firefly luciferase signal.
- FIG. 16A-C Targeting endogenous transcripts with CIRTS.
- A Nuclease- mediated knockdown of five endogenous transcripts upon transfection of cells with CIRTS- 1 as assayed using qPCR.
- B qPCR analysis of YTHDF2-mediated knockdown of endogenous transcripts with CIRTS- 3.
- CIRTS-2 Analysis of protein levels after transfection with CIRTS-2 or CIRTS-3 by Western blot. CIRTS-2 induces an increase in protein levels, whereas CIRTS-3 shows the expected decrease in protein levels, both in a gRNA-dependent manner.
- FIG. 17 Targeting accessibility determines knockdown efficiency of CIRTS.
- gRNA screen along SMARCA4 using CIRTS-3 to induce gRNA-dependent RNA decay. The inventors observe significant changes in the amount of induced decay dependent on where the transcript is targeted (n 2 or 3).
- FIG. 18A-D Multidimensional targeting with CIRTS.
- A Schematic of delivery of CIRTS-6 and three gRNAs.
- C Schematic of simultaneous CIRTS delivery with different effector proteins.
- D Changes in luciferase protein levels and PPIB transcript levels when cells were transfected with both CIRTS-9 (YTHDF1) and CIRTS-10 (YTHDF2) and gRNAs for Flue and PPIB respectively.
- FIG. 19A-C AAV Delivery of CIRTS.
- A Transfer plasmid for AAV delivery containing both the CIRTS-6 (YTHDF2) as well as the gRNA component of the system. The total insert size between the two inverted terminal repeats (ITR) was 2.7 kb.
- B AAV- packaged CIRTS-6 and a gRNA targeting luciferase was delivered to HEK293T cells to knockdown firefly luciferase in the dual luciferase reporter assay.
- FIG. 20 Comparing CIRTS to other DNA and RNA-targeting CRISPR/Cas systems. Schematic size comparison of commonly used Cas9, Casl2, Casl3, and fusion protein systems.
- FIG. 21 CIRTS List continued from FIG. 12B. Reference list of all remaining CIRTS used in this work.
- FIG. 22A-J Control luciferase assays and RT-qPCRs.
- A Luciferase assay comparing the nuclease-mediated decay of TBP6.7-Pin nuclease domain without (CIRTS-11) and with (CIRTS-12) the additional ssRNA binding protein ORF5.
- C RT-qPCR analysis of RNA levels with the‘dead’ Pin nuclease domain CIRTS (CIRTS-0).
- RNA levels when cells were transfected with CIRTS-1 and active Casl3b nuclease were transfected with CIRTS-1 and active Casl3b nuclease.
- CIRTS-l-Pin mediated RNA cleavage showed substantially less RNA degradation compared to the Casl3b system.
- E-H All engineered CIRTS system tested in the dual luciferase assay were also subjected to RT-qPCR analysis to assess changes in RNA levels.
- CIRTS-2 which contain the YTHDF1 effector domain inducing translation activation showed no significant changes in RNA level while all YTHDF2-containing CIRTS show the expected decrease in RNA levels.
- FIG. 23A-D CIRTS linker and gRNA optimization.
- A Luciferase assay with the CIRTS nuclease system using different linkers between the hairpin-binding protein and the effector protein.
- C Different engineered gRNA for TBP6.7 based on the design shown in Figure 1C. Two different targeting lengths of 20 and 40 nucleotides were used in combination with different numbers of linking nucleotides (L) between the hairpin and the guiding sequence. The dual luciferase assay was used to assess nuclease-mediated decay.
- FIG. 24A-D Control qPCR, Western Blot, YTHDF2 truncations.
- CIRTS- 1 can be delivered to RNA species other than mRNA.
- CIRTS-1 Pieris nuclease
- two different gRNAs for the IncRNA MALAT1 assessed RNA levels by RT-qPCR.
- B RT-qPCR analysis of RNA level when cells were transfected with CIRTS-2. As anticipated, no significant changes in RNA level were observed when a YTHDF1 -containing protein was used.
- FIG. 25A-H Targeting Specificity of CIRTS.
- A Schematic of the KRAS4b- luciferase mismatch reporter assay. The inventors chose four KRAS4b variants that have an increasing number of mismatches to the designed 20 nt length gRNA and fused it N-terminal to the dual luciferase reporter.
- B CIRTS-mediated knockdown of KRAS4b-Fluc with different numbers of mismatches between the gRNA and target RNA as described in Figure S5A. CIRTS was found to be most sensitive to mismatches in the middle of its guiding sequence.
- C Casl3b-mediated knockdown in the same KRAS4b-Fluc reporter assay as described above. Casl3b shows a higher knockdown efficiency but is also less sensitive to mismatches introduced. Similar to CIRTS, Casl3b is knockdown is most affected by mismatches at the center of the guiding target duplex region.
- D Knockdown efficiency of CIRTS on the KRAS4b-luciferase mismatch reporter when using a 40 nt gRNA length. A longer guiding sequence in the gRNA can rescue some of the loss in knockdown efficiency.
- G Knockdown levels of SMARCA4 as determined by RNA sequencing.
- FIG. 26A-C Endogenous targeting with CIRTS.
- A Changes in RNA levels as assessed by RT-qPCR after transfection of CIRTS5-7 alone for PPIB.
- B Similar to Figure S6A, SMARCA4 levels were assayed when cells were transfected with CIRTS5-7.
- FIG. 27A-C Multiplexed targeting with CIRTS.
- A Schematic of vectors used for multiplexed targeting. Cells were transfected with an expression vector for CIRTS-6, and an expression vector for CIRTS-10, an expression vector for a CIRTS-6 gRNA construct targeting PPIB or a non-targeting control, and an expression vector for a CIRTS-9 gRNA targeting SMARCA4 or a non-targeting control.
- B Heat map showing knockdown of multiplexed targeting described in (A). When both CIRTS have an on-target gRNA for PPIB or SMARCA4 present, both transcripts can be knocked down in the same samples.
- FIG. 28 gRNA screen with ADAR. Testing whether guide RNA designs that feature multiple hairpins increase the potency of CIRTS.
- Left panel shows gRNA designs feature either the origins design, or guides with one TAR hairpin on either end of the guide, or two hairpins on either end. The additional hairpin guides increase the potency of CIRTS in an ADAR activity assay in cells.
- Right panel is testing whether the second hairpin needs to be a TAR hairpin, or if just a "stabilizing hairpin” can function - meaning a hairpin that does not directly interact with the CIRTS protein but slows degradation. As seen in the data, the second hairpin increase potency compared to one hairpin gRNA design.
- FIG. 29A-B shows gRNA screen with ADAR. Testing whether guide RNA designs that feature multiple hairpins increase the potency of CIRTS.
- A C-to-U Editor: The data in A demonstrates that a CIRTS based on a C-to-U base editor is also functional using a mammalian cell reporter assay.
- B ssRNA binding proteins. The data in B demonstrates that other ssRNA binding proteins can function as a RNA binding proteins in the systems and methods of the disclosure.
- FIG. 30A-C RNA regulatory domain-containing proteins that may activate translation, arraying gRNAs at various locales on a reporter RNA and measuring translational activation of each.
- B-C RNA regulatory domain-containing proteins that potentially degrade or destabilize an RNA, arraying gRNAs at various locales on a reporter RNA and measuring RNA degradation of each.
- FIG. 31A-B (A) Embodiments demonstrating the different orientations of the elements of the systems of the disclosure. (B) Data using CNOT7 in different orientations (as shown in A) and on two RNA targets: a luciferase reporter (left) and an endogenous RNA (right). Several different orientations of the proteins still function, indicating the proteins can be engineered in differed orders depending on the needs of the effector.
- FIG. 32A-D A CIRTS biosensor for inducible RNA targeting.
- A Schematic overview of the abscisic acid (ABA) CIRTS biosensor design. The gRNA-mediated targeting component of CIRTS is fused to one of the ABA heterodimerization domains (ABI) while the effector component of CIRTS is fused to its binding partner (PYL). Upon addition of the small molecule ABA, the two CIRTS components dimerize and bring the effector in proximity of the targeted transcript.
- ABA-inducible RNA degradation of an red-fluorescent protein (RFP) reporter transcript 48h after transfection can be mediated by the Pin nuclease domain or YTHDF2.
- RFP red-fluorescent protein
- C Translation activation of RFP by ABA-induced CIRTS-YTHDFl 48h after transfection.
- D Delivery of CIRTS-hADAR with on-target gRNA in the presence of ABA to cells transfected with a mutation-deactivated luciferase reporter (FlucW417X for A-to-I editing or the GlucC82R for C-to-U editing) induces AB A-dependent RNA editing.
- a mutation-deactivated luciferase reporter FlucW417X for A-to-I editing or the GlucC82R for C-to-U editing
- CRISPR/Cas-inspired RNA targeting system CRISPR/Cas-inspired RNA targeting system
- the inventors show that CIRTS is a simple and generalizable approach to deliver a range of effector proteins, including nucleases, degradation machinery, and translational activators, to target transcripts.
- CIRTS are not only smaller than naturally-occurring CRISPR/Cas programmable RNA binding systems, but can be built entirely from human protein parts.
- the small size and human-derived nature of CIRTS provides a less perturbative method for fundamental RNA regulatory studies as well as a potential strategy to avoid immune issues when applied to epitranscriptome-modulating therapies.
- RNA regulatory domain may be used in the methods and systems of the current disclosure.
- a RNA regulatory domain with one or more of the following activities may be used: methylation, 5'-3' guanylylation, phosphoribosylation, deamination, carbamoylation, isopentenylation, agmatinylation, acetylation, lysylation, O/S exchange, galactosylation, glutamyl ati on, mannosylation, hydrogenation, pseudouridine formation, carboxymethylaminomethylation, aminomethylation, decarboxymethylation, dehydrogenation, carboxymethylation, hydroxylation, methylthiolation, 3 -amino-3 - carboxypropylation, dem ethylation, 5 '-5' guanylylation, and dephosphorylation.
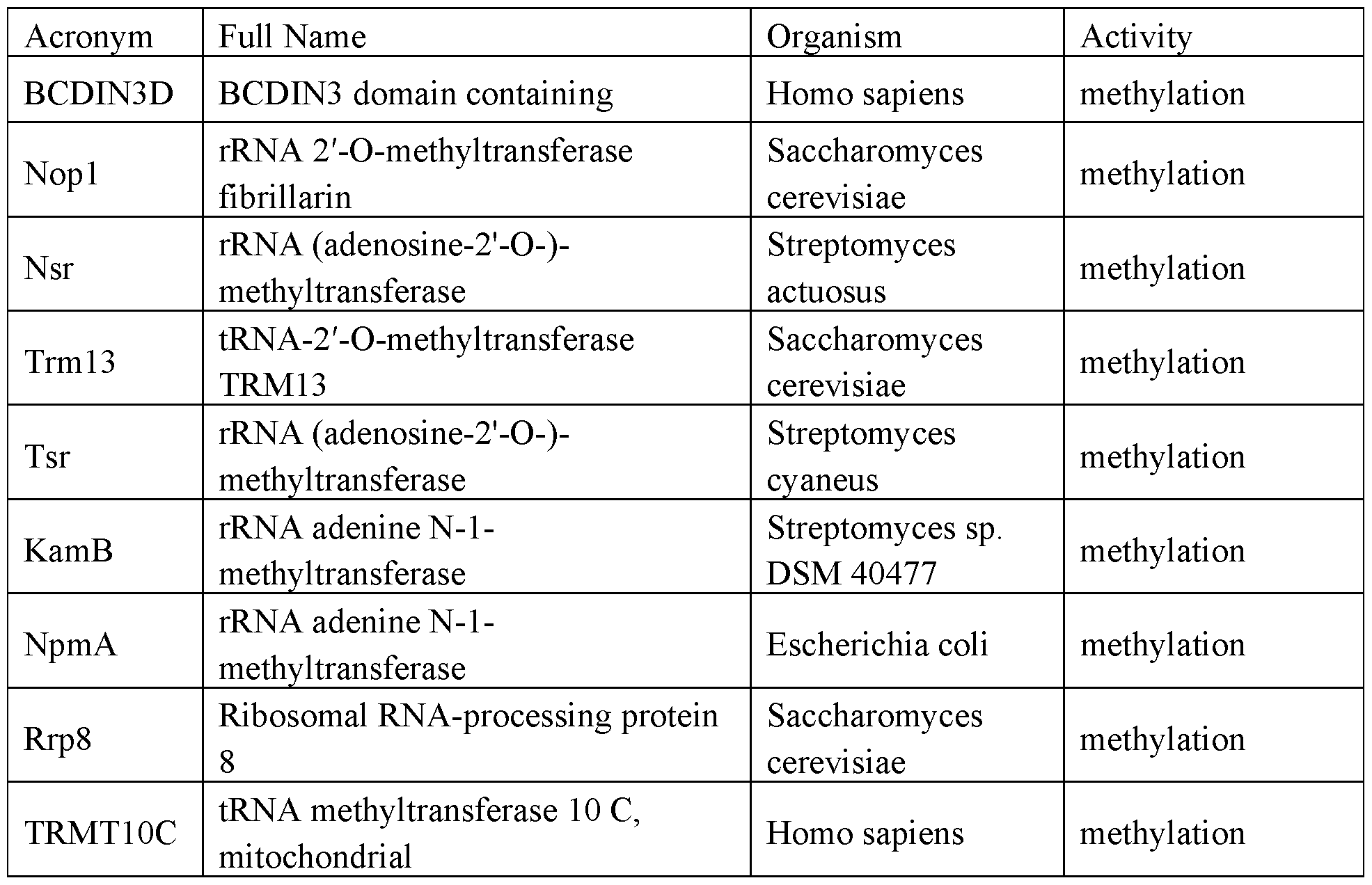
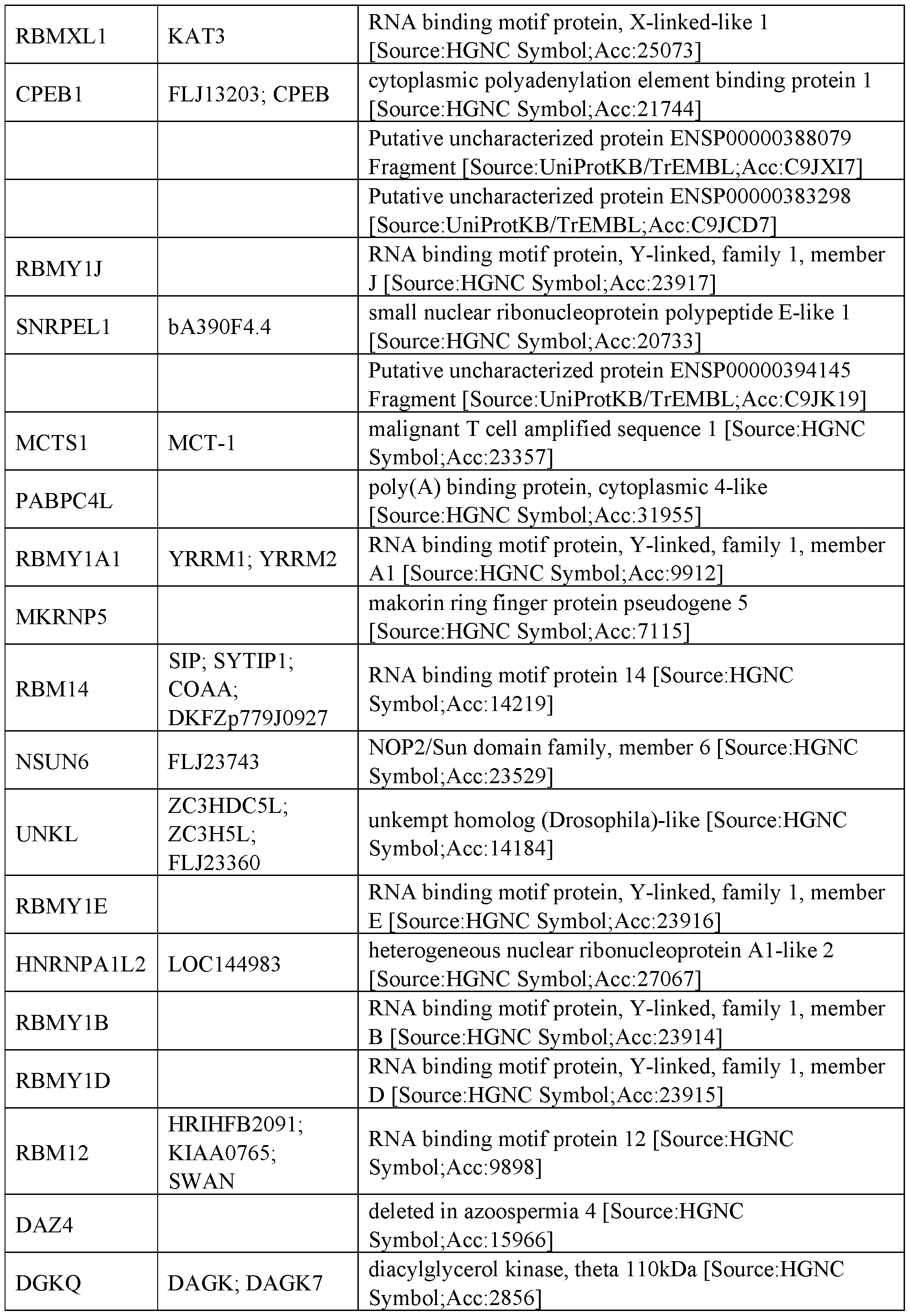
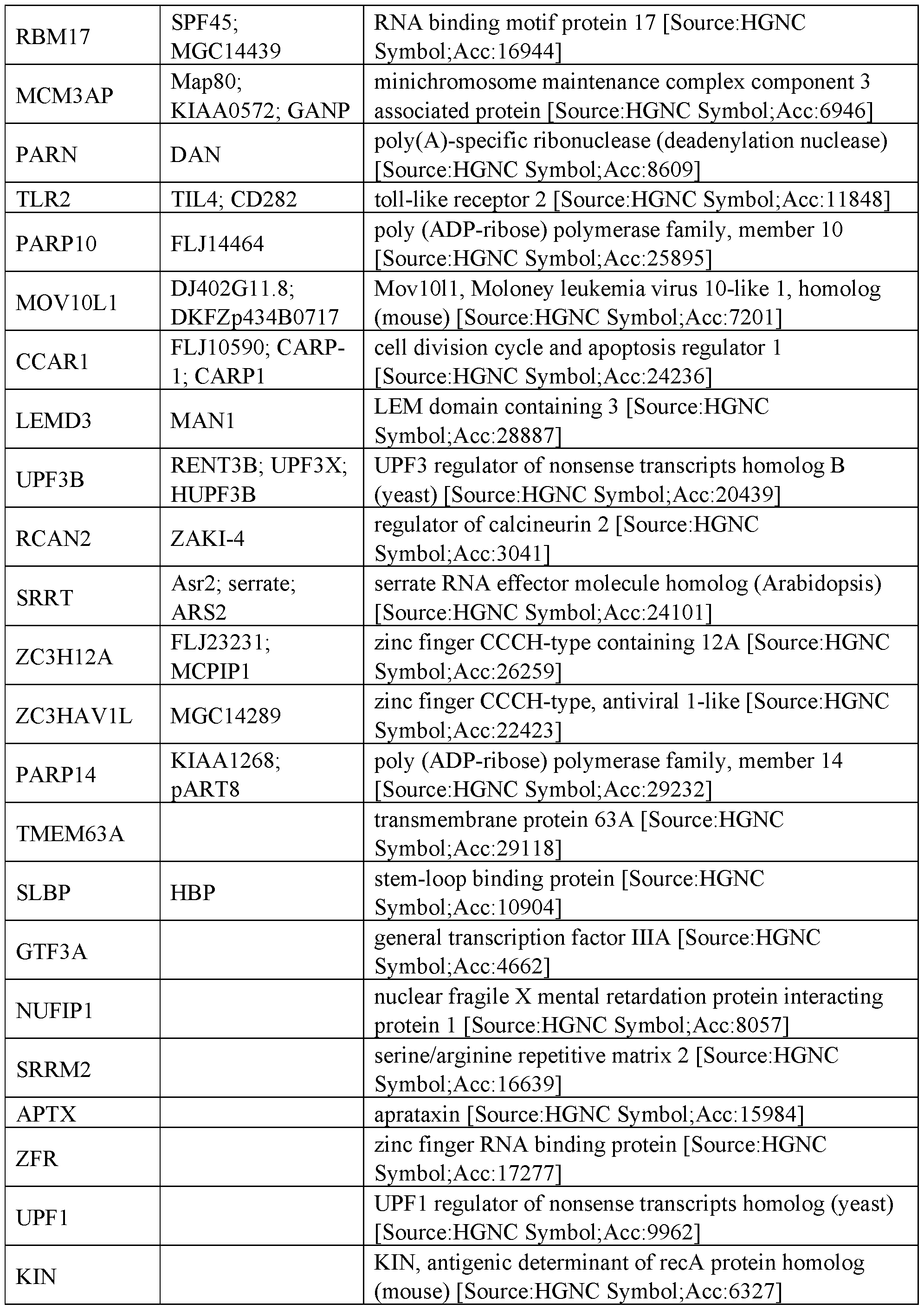
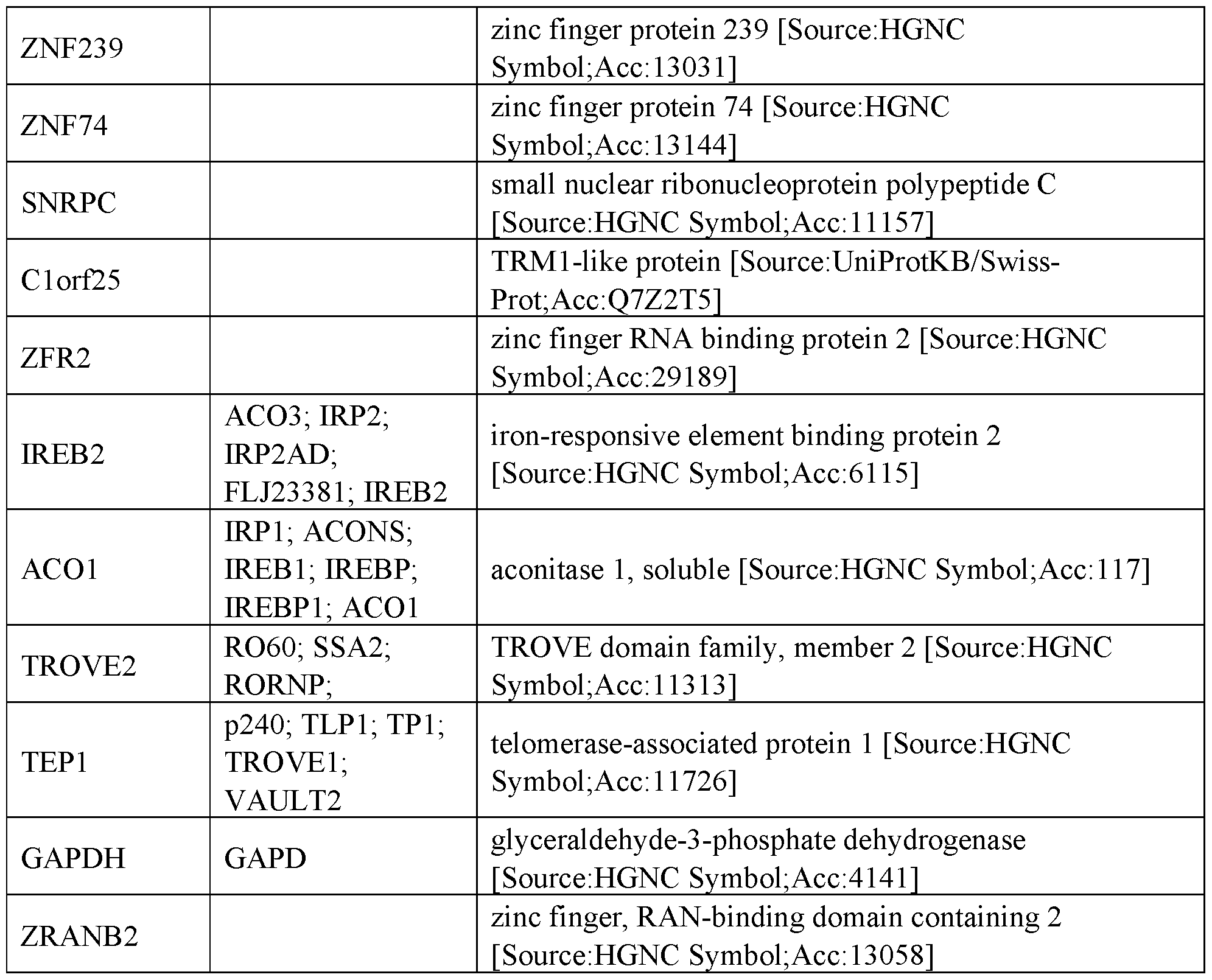
- RNA regulatory domains include domains from the following proteins of Table 1 (or functional fragments thereof):
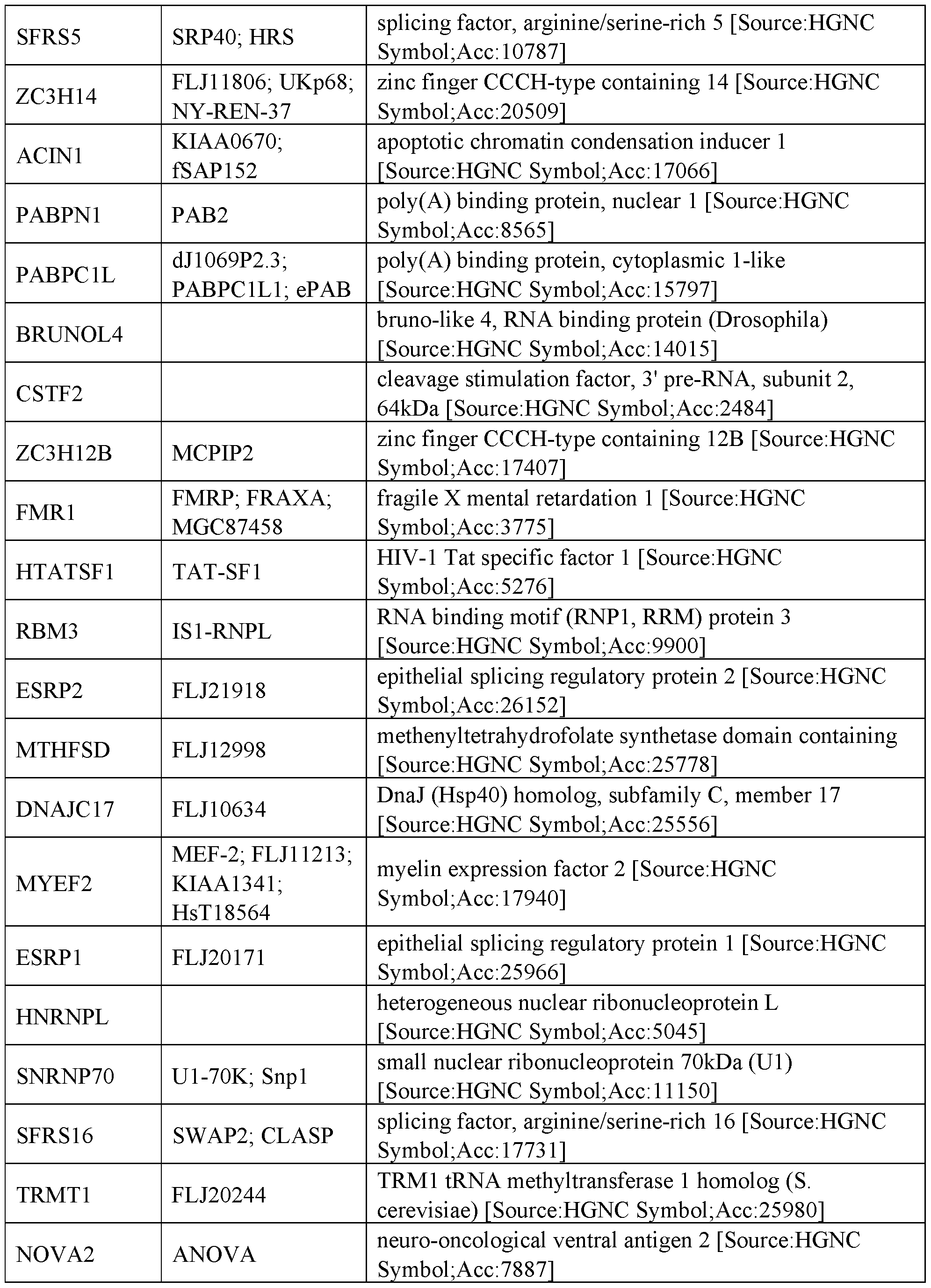
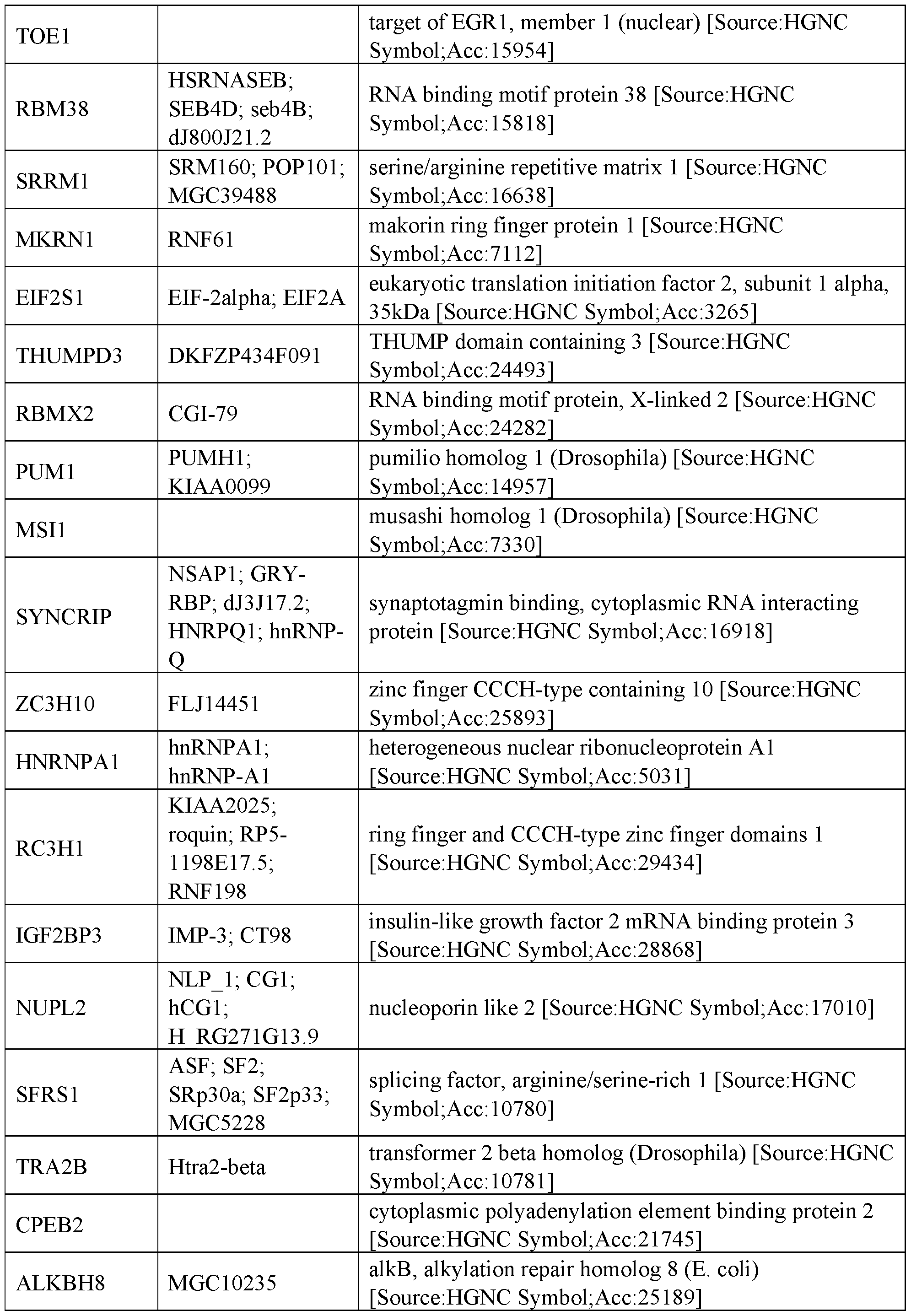
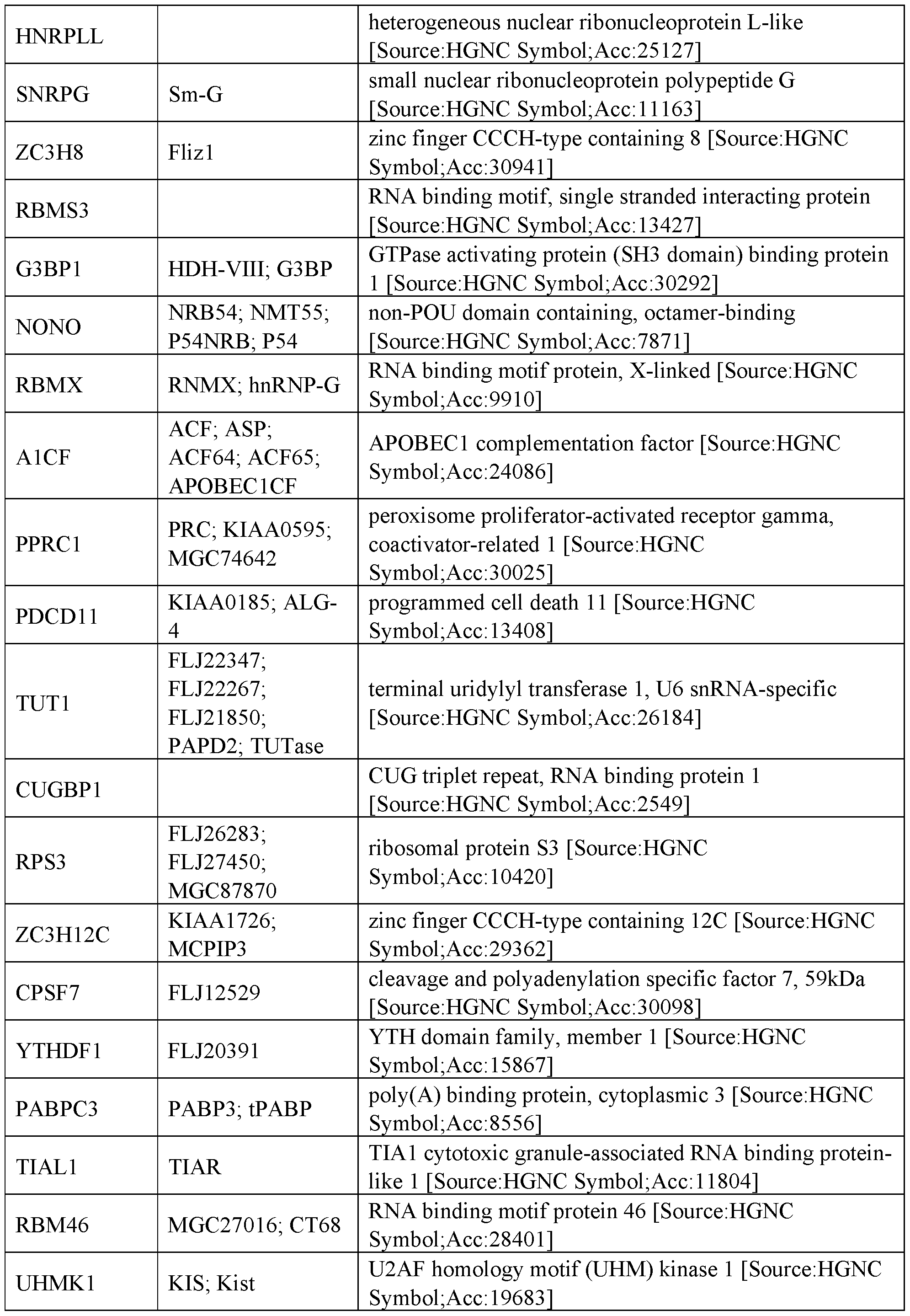
- RNA regulatory domains include functional domains from the following human proteins of Table 2:
- the RNA regulatory domain may be a protein selected from Table 1 or Table 2 or a functional domain from a protein selected from the list of proteins in Table 1 or 2.
- the RNA regulatory domain comprises a fragment from a protein selected from the list of proteins in Table 1 or 2.
- the RNA regulatory domain comprises a protein having at least, at most, or exactly 100, 99, 98, 97, 96, 95, 94, 93, 92, 91,
- the RNA regulatory domain comprises at least, at most, or exactly 100, 150, 200, 250, 300, 350, 400, 450, 500, 550, 600, 650, 700, 750, 800, 850, 900, 950, 1000, 1050, 1100, 1150, 1200, 1250, 1300, 1350, 1400, 1450, 1500, 1550, 1600, 1650,
- the RNA regulatory domain comprises a fragment of a protein from Table 1 or 2, wherein the fragment has one or more of the following activities: methylation, 5'-3' guanylylation, phosphoribosylation, deamination, carbamoylation, isopentenylation, agmatinylation, acetylation, lysylation, O/S exchange, galactosylation, glutamyl ati on, mannosylation, hydrogenation, pseudouridine formation, carb oxy methyl aminomethy 1 ati on, aminomethy 1 ati on, decarb oxy methyl ati on, dehydrogenation, carboxymethylation, hydroxylation, methylthiolation, 3 -amino-3 - carboxypropylation, dem ethylation, 5 '-5'
- the RNA regulatory domain is at or near the carboxy- terminus of the RNA hairpin binding protein. In some embodiments, the RNA regulatory domain is at or near the amino-terminus of the RNA hairpin binding protein. In some embodiments, the RNA regulatory domain is fused by way of a peptide bond to the RNA hairpin binding protein. In some embodiments, the RNA regulatory domain is linked to the RNA hairpin binding protein by a linker moiety.
- RNA hairpin binding domains and hairpin structures that they bind are known in the art and can be used in the systems, compositions, fusion proteins, kits, vectors, and methods of the disclosure.
- embodiments include a RNA hairpin binding domain and hairpin structure according to the following table (Table 3), which lists proteins comprising RNA hairpin binding domains and the hairpin structure that they specifically bind to:
- RNA hairpin binding domains and/or the RNA regulatory domain may be used in a multiplexed fashion by using RNA hairpin binding domains that bind to different hairpin structures to target multiple different RNAs in the same cell.
- the different RNAs may be modulated in the same or in different ways. For example, one RNA may be modulation with translational activation, while a second RNA may be modulated with translational repression in the same cell. Therefore, the systems of the disclosure can be used in a multiplexed fashion for the modulation of at least 2, 3, 4, 5, 6, 7, 8, 9, 10 or more RNAs in one cell, tissue, or organisms.
- nucleic acids encoding the proteins, polypeptides, regulatory domains, or RNA targeting molecules described herein.
- polynucleotide refers to a nucleic acid molecule that either is recombinant or has been isolated free of total genomic nucleic acid. Included within the term“polynucleotide” are oligonucleotides (nucleic acids 100 residues or fewer in length), recombinant vectors, including, for example, plasmids, cosmids, phage, viruses, and the like. Polynucleotides include, in certain aspects, regulatory sequences, isolated substantially away from their naturally occurring genes or protein encoding sequences.
- Polynucleotides may be single-stranded (coding or antisense) or double-stranded, and may be RNA, DNA (genomic, cDNA or synthetic), analogs thereof, or a combination thereof. Additional coding or non-coding sequences may, but need not, be present within a polynucleotide.
- the term“gene,”“polynucleotide,” or“nucleic acid” is used to refer to a nucleic acid that encodes a protein, polypeptide, or peptide (including any sequences required for proper transcription, post-translational modification, or localization).
- this term encompasses genomic sequences, expression cassettes, cDNA sequences, and smaller engineered nucleic acid segments that express, or may be adapted to express, proteins, polypeptides, domains, peptides, fusion proteins, and mutants.
- a nucleic acid encoding all or part of a polypeptide may contain a contiguous nucleic acid sequence encoding all or a portion of such a polypeptide. It also is contemplated that a particular polypeptide may be encoded by nucleic acids containing variations having slightly different nucleic acid sequences but, nonetheless, encode the same or substantially similar protein (see above).
- nucleic acid segments and recombinant vectors incorporating nucleic acid sequences that encode a polypeptides e.g., a polymerase, RNA polymerase, one or more truncated polymerase domains or interaction components that are polypeptides
- a polypeptides e.g., a polymerase, RNA polymerase, one or more truncated polymerase domains or interaction components that are polypeptides
- the term“recombinant” may be used in conjunction with a polypeptide or the name of a specific polypeptide, and this generally refers to a polypeptide produced from a nucleic acid molecule that has been manipulated in vitro or that is a replication product of such a molecule.
- nucleic acid segments regardless of the length of the coding sequence itself, may be combined with other nucleic acid sequences, such as promoters, polyadenylation signals, additional restriction enzyme sites, multiple cloning sites, other coding segments, and the like, such that their overall length may vary considerably. It is therefore contemplated that a nucleic acid fragment of almost any length may be employed, with the total length preferably being limited by the ease of preparation and use in the intended recombinant nucleic acid protocol.
- a nucleic acid sequence may encode a polypeptide sequence with additional heterologous coding sequences, for example to allow for purification of the polypeptide, transport, secretion, post-translational modification, or for therapeutic benefits such as targeting or efficacy.
- a tag or other heterologous polypeptide may be added to the modified polypeptide-encoding sequence, wherein“heterologous” refers to a polypeptide that is not the same as the modified polypeptide.
- polynucleotide variants having substantial identity to the sequences disclosed herein; those comprising at least 70%, 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, or 99% or higher sequence identity, including all values and ranges there between, compared to a polynucleotide sequence provided herein using the methods described herein (e.g ., BLAST analysis using standard parameters).
- the isolated polynucleotide will comprise a nucleotide sequence encoding a polypeptide that has at least 90%, preferably 95% and above, identity to an amino acid sequence described herein, over the entire length of the sequence; or a nucleotide sequence complementary to said isolated polynucleotide.
- Polypeptides may be encoded by a nucleic acid molecule.
- the nucleic acid molecule can be in the form of a nucleic acid vector.
- vector is used to refer to a carrier nucleic acid molecule into which a heterologous nucleic acid sequence can be inserted for introduction into a cell where it can be replicated and expressed.
- a nucleic acid sequence can be“heterologous,” which means that it is in a context foreign to the cell in which the vector is being introduced or to the nucleic acid in which is incorporated, which includes a sequence homologous to a sequence in the cell or nucleic acid but in a position within the host cell or nucleic acid where it is ordinarily not found.
- Vectors include DNAs, RNAs, plasmids, cosmids, viruses (bacteriophage, animal viruses, and plant viruses), and artificial chromosomes (e.g., YACs).
- viruses bacteriophage, animal viruses, and plant viruses
- artificial chromosomes e.g., YACs.
- Vectors may be used in a host cell to produce a polymerase, RNA polymerase, one or more truncated polymerase domains or interaction components that are fused, attached or linked to the one or more truncated RNA polymerase domains.
- expression vector refers to a vector containing a nucleic acid sequence coding for at least part of a gene product capable of being transcribed. In some cases, RNA molecules are then translated into a protein, polypeptide, or peptide.
- Expression vectors can contain a variety of“control sequences,” which refer to nucleic acid sequences necessary for the transcription and possibly translation of an operably linked coding sequence in a particular host organism. In addition to control sequences that govern transcription and translation, vectors and expression vectors may contain nucleic acid sequences that serve other functions as well and are described herein.
- the disclosure provides methods for modifying a target RNA of interest, in particular in prokaryotic cells, eukaryotic cells, tissues, organs, or organisms, more in particular in mammalian cells, tissues, organs, or organisms.
- the target RNA may be comprised in a nucleic acid molecule within a cell.
- the target RNA is in a eukaryotic cell, such as a mammalian cell or a plant cell.
- the mammalian cell many be a human, non human primate, bovine, porcine, rodent or mouse cell.
- the cell may be a non-mammalian eukaryotic cell such as poultry, fish or shrimp.
- the plant cell may be of a crop plant such as cassava, com, sorghum, wheat, or rice.
- the plant cell may also be of an algae, tree or vegetable.
- the modulation of the RNA induced in the cell by the methods, systems, and compositions of the disclosure may be such that the cell and progeny of the cell are altered for improved production of biologic products such as an antibody, starch, alcohol or other desired cellular output.
- the modulation of the RNA induced in the cell may be such that the cell and progeny of the cell include an alteration that changes the biologic product produced.
- the mammalian cell may be a human or non-human mammal, e.g., primate, bovine, ovine, porcine, canine, rodent, Leporidae such as monkey, cow, sheep, pig, dog, rabbit, rat or mouse cell.
- the cell may be a non-mammalian eukaryotic cell such as poultry bird (e.g., chicken), vertebrate fish (e.g., salmon) or shellfish (e.g., oyster, clam, lobster, shrimp) cell.
- the cell may also be a plant cell.
- the plant cell may be of a monocot or dicot or of a crop or grain plant such as cassava, com, sorghum, soybean, wheat, oat or rice.
- the plant cell may also be of an algae, tree or production plant, fruit or vegetable (e.g., trees such as citrus trees, e.g., orange, grapefruit or lemon trees; peach or nectarine trees; apple or pear trees; nut trees such as almond or walnut or pistachio trees; nightshade plants; plants of the genus Brassica; plants of the genus Lactuca; plants of the genus Spinacia; plants of the genus Capsicum; cotton, tobacco, asparagus, carrot, cabbage, broccoli, cauliflower, tomato, eggplant, pepper, lettuce, spinach, strawberry, blueberry, raspberry, blackberry, grape, coffee, cocoa, etc.).
- “cell,” “cell line,” and“cell culture” may be used interchangeably. All of these terms also include their progeny, which is any and all subsequent generations. It is understood that all progeny may not be identical due to deliberate or inadvertent mutations.
- “host cell” refers to a prokaryotic or eukaryotic cell, and it includes any transformable organism that is capable of replicating a vector or expressing a heterologous gene encoded by a vector.
- a host cell can, and has been, used as a recipient for vectors or viruses.
- a host cell may be “transfected” or“transformed,” which refers to a process by which exogenous nucleic acid, such as a recombinant protein-encoding sequence, is transferred or introduced into the host cell.
- a transformed cell includes the primary subject cell and its progeny.
- Some vectors may employ control sequences that allow it to be replicated and/or expressed in both prokaryotic and eukaryotic cells.
- control sequences that allow it to be replicated and/or expressed in both prokaryotic and eukaryotic cells.
- One of skill in the art would further understand the conditions under which to incubate all of the above described host cells to maintain them and to permit replication of a vector. Also understood and known are techniques and conditions that would allow large-scale production of vectors, as well as production of the nucleic acids encoded by vectors and their cognate polypeptides, proteins, or peptides.
- Prokaryote- and/or eukaryote-based systems can be employed for use with an embodiment to produce nucleic acid sequences, or their cognate polypeptides, proteins and peptides.
- the vectors, fusion proteins, RNA hairpin binding proteins, RNA targeting molecules, RNA regulatory domain, and accessory proteins of the disclosure may utilize an expression system, such as an inducible or constitutive expression system. Many such systems are commercially and widely available.
- the insect cell/baculovirus system can produce a high level of protein expression of a heterologous nucleic acid segment, such as described in U.S. Patents 5,871,986, 4,879,236, both herein incorporated by reference, and which can be bought, for example, under the name MAXBAC® 2.0 from INVITROGEN® and BACPACKTM BACULOVIRUS EXPRESSION SYSTEM FROM CLONTECH®.
- a heterologous nucleic acid segment such as described in U.S. Patents 5,871,986, 4,879,236, both herein incorporated by reference, and which can be bought, for example, under the name MAXBAC® 2.0 from INVITROGEN® and BACPACKTM BACULOVIRUS EXPRESSION SYSTEM FROM CLONTECH®.
- STRATAGENE® COMPLETE CONTROL Inducible Mammalian Expression System, which involves a synthetic ecdysone-inducible receptor, or its pET Expression System, an E. coli expression system.
- INVITROGEN ® which carries the T-REXTM (tetracycline-regulated expression) System, an inducible mammalian expression system that uses the full-length CMV promoter.
- INVITROGEN ® also provides a yeast expression system called the Pichia methanolica Expression System, which is designed for high-level production of recombinant proteins in the methyl otrophic yeast Pichia methanolica.
- a vector such as an expression construct, to produce a nucleic acid sequence or its cognate polypeptide, protein, or peptide.
- Embodiments of the disclosure relate to the conjugation of nucleic acids to polypeptides. Methods of conjugation of nucleic acids to polypeptides are known in the art and include those described below. Embodiments of the disclosure relate to methods of making nucleic acid-polypeptide molecules and the molecules themselves wherein the nucleic acid has been conjugated to the polypeptide by way of a method described herein.
- One such example includes click chemistry.
- the "click reaction”, also known as "click chemistry” is a name often used to describe a stepwise variant of the Huisgen 1,3-dipolar cycloaddition of azides and alkynes to yield 1,2,3-triazole.
- This reaction is carried out under ambient conditions, or under mild microwave irradiation, typically in the presence of a Cu(I) catalyst, and with exclusive regioselectivity for the 1,4-di substituted triazole product when mediated by catalytic amounts of Cu(I) salts [V. Rostovtsev, L. G. Green, V. V. Fokin, K. B. Sharpless, Angew. Chem. Int. Ed. 2002, 41, 2596; H. C. Kolb, M. Finn, K. B. Sharpless, Angew Chem., Int. Ed. 2001, 40, 2004]
- a mutant form of the human DNA repair protein 06- alkylguanine-DNA alkyltransferase reacts rapidly and specifically with 06-benzylguanin (BG) and also with derivatives that carry a large moiety linked to the benzyl group.
- BG 06-benzylguanin
- the benzyl moiety With guanine as the leaving group, the benzyl moiety becomes covalently attached to a cysteine in the active site of the enzyme.
- the enzyme has also been mutagenized to become specific for 06- benzylcytosine (BC) in a similar manner.
- BC 06- benzylcytosine
- a further conjugation method utilizes the Halo tag.
- the Halo tag makes use of a chemical reaction orthogonal to eukaryotes, i.e. the dehalogenation of haloalkane ligands, thus, leading to highly specific covalent labelling of the tag, and therefore protein, in both live and fixed cells.
- RNA targeting molecules such as the RNA targeting molecules and other nucleic acids described herein may have modifications that increase the stability of the nucleic acid.
- the RNA targeting molecule is an oligonucleotide analogs.
- oligonucleotide analog refers to compounds which function like oligonucleotides but which have non-naturally occurring portions. Oligonucleotide analogs can have altered sugar moieties, altered base moieties or altered inter-sugar linkages.
- oligomers is intended to encompass oligonucleotides, oligonucleotide analogs or oligonucleosides.
- oligomers reference is made to a series of nucleosides or nucleoside analogs that are joined via either natural phosphodiester bonds or other linkages, including the four atom linkers.
- linkage generally is from the 3’ carbon of one nucleoside to the 5’ carbon of a second nucleoside
- the term“oligomer” can also include other linkages such as T - 5’ linkages.
- Oligonucleotide analogs also can include other modifications, particularly modifications that increase nuclease resistance, improve binding affinity, and/or improve binding specificity. For example, when the sugar portion of a nucleoside or nucleotide is replaced by a carbocyclic moiety, it is no longer a sugar. Moreover, when other substitutions, such a substitution for the inter-sugar phosphodiester linkage are made, the resulting material is no longer a true nucleic acid species. All such compounds are considered to be analogs. Throughout this specification, reference to the sugar portion of a nucleic acid species shall be understood to refer to either a true sugar or to a species taking the structural place of the sugar of wild type nucleic acids. Moreover, reference to inter-sugar linkages shall be taken to include moieties serving to join the sugar or sugar analog portions in the fashion of wild type nucleic acids.
- modified oligonucleotides i.e., oligonucleotide analogs or oligonucleosides
- modified oligonucleotides and oligonucleotide analogs may exhibit increased chemical and/or enzymatic stability relative to their naturally occurring counterparts.
- Extracellular and intracellular nucleases generally do not recognize and therefore do not bind to the backbone-modified compounds. When present as the protonated acid form, the lack of a negatively charged backbone may facilitate cellular penetration.
- the modified internucleoside linkages are intended to replace naturally-occurring phosphodiester-5’ -methylene linkages with four atom linking groups to confer nuclease resistance and enhanced cellular uptake to the resulting compound.
- Preferred linkages have structure CH2 -RA— N 1 CH2, CH2— N 1 -RA -CH2, RA— NR1 -CH2 -CH2, CH2 - CH2— NR1 -RA, CH2 -CH2 -RA— NR1, or NR1 -RA -CH2 -CH2 where RA is O or NR2.
- Modifications may be achieved using solid supports which may be manually manipulated or used in conjunction with a DNA synthesizer using methodology commonly known to those skilled in DNA synthesizer art. Generally, the procedure involves functionalizing the sugar moieties of two nucleosides which will be adjacent to one another in the selected sequence. In a 5’ to 3’ sense, an“upstream” synthon is modified at its terminal 3’ site, while a“downstream” synthon is modified at its terminal 5’ site.
- Oligonucleosides linked by hydrazines, hydroxylarnines, and other linking groups can be protected by a dimethoxytrityl group at the 5’ -hydroxyl and activated for coupling at the 3’ -hydroxyl with cyanoethyldiisopropyl-phosphite moieties. These compounds can be inserted into any desired sequence by standard, solid phase, automated DNA synthesis techniques. One of the most popular processes is the phosphoramidite technique. Oligonucleotides containing a uniform backbone linkage can be synthesized by use of CPG- solid support and standard nucleic acid synthesizing machines such as Applied Biosystems Inc.
- the initial nucleotide (number 1 at the 3’ -terminus) is attached to a solid support such as controlled pore glass. In sequence specific order, each new nucleotide is attached either by manual manipulation or by the automated synthesizer system.
- Free amino groups can be alkylated with, for example, acetone and sodium cyanoboro hydride in acetic acid.
- the alkylation step can be used to introduce other, useful, functional molecules on the macromolecule.
- useful functional molecules include but are not limited to reporter molecules, RNA cleaving groups, groups for improving the pharmacokinetic properties of an oligonucleotide, and groups for improving the pharmacodynamic properties of an oligonucleotide.
- Such molecules can be attached to or conjugated to the macromolecule via attachment to the nitrogen atom in the backbone linkage. Alternatively, such molecules can be attached to pendent groups extending from a hydroxyl group of the sugar moiety of one or more of the nucleotides. Examples of such other useful functional groups are provided by WO1993007883, which is herein incorporated by reference, and in other of the above-referenced patent applications.
- Solid supports may include any of those known in the art for polynucleotide synthesis, including controlled pore glass (CPG), oxalyl controlled pore glass [53], TentaGel Support— an aminopolyethyleneglycol derivatized support [54] or Poros— a copolymer of polystyrene/divinylbenzene. Attachment and cleavage of nucleotides and oligonucleotides can be effected via standard procedures [55] As used herein, the term solid support further includes any linkers (e.g., long chain alkyl amines and succinyl residues) used to bind a growing oligonucleoside to a stationary phase such as CPG.
- CPG controlled pore glass
- TentaGel Support an aminopolyethyleneglycol derivatized support
- Poros a copolymer of polystyrene/divinylbenzene. Attachment and cleavage of nucleo
- the nucleic acid of the disclosure such as the RNA targeting molecule comprises a locked nucleic acid.
- a locked nucleic acid (LNA or Ln), also referred to as inaccessible RNA, is a modified RNA nucleotide.
- the ribose moiety of an LNA nucleotide is modified with an extra bridge connecting the T oxygen and 4’ carbon. The bridge “locks” the ribose in the 3’-endo (North) conformation, which is often found in the A-form duplexes.
- LNA nucleotides can be mixed with DNA or RNA residues in the oligonucleotide whenever desired and hybridize with DNA or RNA according to Watson-Crick base-pairing rules. Such oligomers are synthesized chemically and are commercially available.
- the locked ribose conformation enhances base stacking and backbone pre-organization. This significantly increases the hybridization properties (melting temperature) of oligonucleotides.
- the nucleic acid of the disclosure such as the RNA targeting molecule comprises one or more ethylene bridged nucleotides.
- Ethylene-bridged nucleic acids are modified nucleotides with a 2’-0, 4’C ethylene linkage. Like locked nucleotides, these nucleotides also restrict the sugar puckering to the N-conformation of RNA.
- the nucleic acid of the disclosure such as the RNA targeting molecule comprises one or more peptide nucleic acids.
- Peptide nucleic acids (PNA or Pn) mimic the behavior of DNA and binds complementary nucleic acid strands.
- the term, “peptide,” when used herein may also refer to a peptide nucleic acid.
- PNA is an artificially synthesized polymer similar to DNA or RNA. DNA and RNA have a deoxyribose and ribose sugar backbone, respectively, whereas PNA’s backbone is composed of repeating N-(2- aminoethyl)-glycine units linked by peptide bonds.
- PNAs are depicted like peptides, with the N-terminus at the first (left) position and the C- terminus at the last (right) position.
- PNAs Since the backbone of PNAs contains no charged phosphate groups, the binding between PNA/DNA strands is stronger than between DNA/DNA strands due to the lack of electrostatic repulsion. PNAs are not easily recognized by either nucleases or proteases, making them resistant to degradation by enzymes. PNAs are also stable over a wide pH range. In some aspects, the PNAs described herein have improved cytosolic delivery over other PNAs.
- the nucleic acid of the disclosure such as the RNA targeting molecule comprises one or more 5’(E)-vinyl-phosphonate (VP) modifications.
- 5’-Vinyl - phosphonate modifications have been reported to enhance the metabolic stability and the potency of oligonucleotides. 5. Morpholinos
- the nucleic acid of the disclosure such as the RNA targeting molecule comprises a morpholino.
- Morpholinos are synthetic molecules that are the product of a redesign of natural nucleic acid structure. Usually 25 bases in length, they bind to complementary sequences of RNA or single-stranded DNA by standard nucleic acid base pairing.
- the difference between Morpholinos and DNA is that, while Morpholinos have standard nucleic acid bases, those bases are bound to methylenemorpholine rings linked through phosphorodiamidate groups instead of phosphates. The figure compares the structures of the two strands depicted there, one of RNA and the other of a Morpholino.
- the current disclosure contemplates several delivery systems compatible with nucleic acids that provide for roughly uniform distribution and have controllable rates of release.
- a variety of different media are described below that are useful in creating nucleic acid delivery systems. It is not intended that any one medium or carrier is limiting to the present invention. Note that any medium or carrier may be combined with another medium or carrier; for example, in one embodiment a polymer microparticle carrier attached to a compound may be combined with a gel medium.
- Carriers or mediums contemplated by this disclosure comprise a material selected from the group comprising gelatin, collagen, cellulose esters, dextran sulfate, pentosan polysulfate, chitin, saccharides, albumin, fibrin sealants, synthetic polyvinyl pyrrolidone, polyethylene oxide, polypropylene oxide, block polymers of polyethylene oxide and polypropylene oxide, polyethylene glycol, acrylates, acrylamides, methacrylates including, but not limited to, 2-hydroxyethyl methacrylate, poly(ortho esters), cyanoacrylates, gelatin- resorcinol-aldehyde type bioadhesives, polyacrylic acid and copolymers and block copolymers thereof.
- microparticles comprise liposomes, nanoparticles, microspheres, nanospheres, microcapsules, and nanocapsules.
- some microparticles contemplated by the present invention comprise poly(lactide-co-glycolide), aliphatic polyesters including, but not limited to, poly-glycolic acid and poly-lactic acid, hyaluronic acid, modified polysaccharides, chitosan, cellulose, dextran, polyurethanes, polyacrylic acids, pseudo-poly(amino acids), polyhydroxybutyrate-related copolymers, polyanhydrides, polymethylmethacrylate, polyethylene oxide), lecithin and phospholipids.
- Liposomes capable of attaching and releasing nucleic acids conjugates, polypeptides, and fusion proteins as described herein.
- Liposomes are microscopic spherical lipid bilayers surrounding an aqueous core that are made from amphiphilic molecules such as phospholipids.
- a liposome may trap a nucleic acid between the hydrophobic tails of the phospholipid micelle.
- Water soluble agents can be entrapped in the core and lipid-soluble agents can be dissolved in the shell-like bilayer. Liposomes have a special characteristic in that they enable water soluble and water insoluble chemicals to be used together in a medium without the use of surfactants or other emulsifiers.
- Liposomes can form spontaneously by forcefully mixing phospholipids in aqueous media. Water soluble compounds are dissolved in an aqueous solution capable of hydrating phospholipids. Upon formation of the liposomes, therefore, these compounds are trapped within the aqueous liposomal center. The liposome wall, being a phospholipid membrane, holds fat soluble materials such as oils. Liposomes provide controlled release of incorporated compounds. In addition, liposomes can be coated with water soluble polymers, such as polyethylene glycol to increase the pharmacokinetic half-life.
- One embodiment of the present invention contemplates an ultra high-shear technology to refine liposome production, resulting in stable, unilamellar (single layer) liposomes having specifically designed structural characteristics. These unique properties of liposomes allow the simultaneous storage of normally immiscible compounds and the capability of their controlled release.
- the disclosure contemplates cationic and anionic liposomes, as well as liposomes having neutral lipids.
- cationic liposomes comprise negatively- charged materials by mixing the materials and fatty acid liposomal components and allowing them to charge-associate.
- the choice of a cationic or anionic liposome depends upon the desired pH of the final liposome mixture. Examples of cationic liposomes include lipofectin, lipofectamine, and lipofectace.
- liposomes that provide controlled release of at least one molecule described herein.
- liposomes that are capable of controlled release i) are biodegradable and non-toxic; ii) carry both water and oil soluble compounds; iii) solubilize recalcitrant compounds; iv) prevent compound oxidation; v) promote protein stabilization; vi) control hydration; vii) control compound release by variations in bilayer composition such as, but not limited to, fatty acid chain length, fatty acid lipid composition, relative amounts of saturated and unsaturated fatty acids, and physical configuration; viii) have solvent dependency; iv) have pH-dependency and v) have temperature dependency.
- compositions of liposomes are broadly categorized into two classifications.
- Conventional liposomes are generally mixtures of stabilized natural lecithin (PC) that may comprise synthetic identical-chain phospholipids that may or may not contain glycolipids.
- Special liposomes may comprise: i) bipolar fatty acids; ii) the ability to attach antibodies for tissue-targeted therapies; iii) coated with materials such as, but not limited to lipoprotein and carbohydrate; iv) multiple encapsulation and v) emulsion compatibility.
- Liposomes may be easily made in the laboratory by methods such as, but not limited to, sonication and vibration.
- compound-delivery liposomes are commercially available.
- Collaborative Laboratories, Inc. are known to manufacture custom designed liposomes for specific delivery requirements.
- Microspheres and microcapsules are useful due to their ability to maintain a generally uniform distribution, provide stable controlled compound release and are economical to produce and dispense.
- an associated delivery gel or the compound-impregnated gel is clear or, alternatively, said gel is colored for easy visualization by medical personnel.
- Microspheres are obtainable commercially (ProleaseTM, Alkerme's: Cambridge, Mass.). For example, a freeze dried medium comprising at least one therapeutic agent is homogenized in a suitable solvent and sprayed to manufacture microspheres in the range of 20 to 90 pm Techniques are then followed that maintain sustained release integrity during phases of purification, encapsulation and storage. Scott et ak, Improving Protein Therapeutics With Sustained Release Formulations, Nature Biotechnology, Volume 16: 153-157 (1998). Modification of the microsphere composition by the use of biodegradable polymers can provide an ability to control the rate of nucleic acid release.
- a sustained or controlled release microsphere preparation is prepared using an in-water drying method, where an organic solvent solution of a biodegradable polymer metal salt is first prepared. Subsequently, a dissolved or dispersed medium of a nucleic acid is added to the biodegradable polymer metal salt solution.
- the weight ratio of a nucleic acid to the biodegradable polymer metal salt may for example be about 1 : 100000 to about 1 : 1, preferably about 1 :20000 to about 1 :500 and more preferably about 1 : 10000 to about 1 :500.
- the organic solvent solution containing the biodegradable polymer metal salt and nucleic acid is poured into an aqueous phase to prepare an oil/water emulsion.
- the solvent in the oil phase is then evaporated off to provide microspheres.
- these microspheres are then recovered, washed and lyophilized. Thereafter, the microspheres may be heated under reduced pressure to remove the residual water and organic solvent.
- Other methods useful in producing microspheres that are compatible with a biodegradable polymer metal salt and nucleic acid mixture are: i) phase separation during a gradual addition of a coacervating agent; ii) an in-water drying method or phase separation method, where an antiflocculant is added to prevent particle agglomeration and iii) by a spray drying method.
- the present invention contemplates a medium comprising a microsphere or microcapsule capable of delivering a controlled release of a nucleic acid for a duration of approximately between 1 day and 6 months.
- the microsphere or microparticle may be colored to allow the medical practitioner the ability to see the medium clearly as it is dispensed.
- the microsphere or microcapsule may be clear.
- the microsphere or microparticle is impregnated with a radio opaque fluoroscopic dye.
- Controlled release microcapsules may be produced by using known encapsulation techniques such as centrifugal extrusion, pan coating and air suspension. Such microspheres and/or microcapsules can be engineered to achieve desired release rates.
- OliosphereTM Macromed
- OliosphereTM is a controlled release microsphere system. These particular microsphere's are available in uniform sizes ranging between 5-500 pm and composed of biocompatible and biodegradable polymers. Specific polymer compositions of a microsphere can control the nucleic acid release rate such that custom-designed microspheres are possible, including effective management of the burst effect.
- ProMaxxTM (Epic Therapeutics, Inc.) is a protein-matrix delivery system. The system is aqueous in nature and is adaptable to standard pharmaceutical delivery models. In particular, ProMaxxTM are bioerodible protein microspheres that deliver both small and macromolecular drugs, and may be customized regarding both microsphere size and desired release characteristics.
- a microsphere or microparticle comprises a pH sensitive encapsulation material that is stable at a pH less than the pH of the internal mesentery.
- the typical range in the internal mesentery is pH 7.6 to pH 7.2. Consequently, the microcapsules should be maintained at a pH of less than 7.
- the pH sensitive material can be selected based on the different pH criteria needed for the dissolution of the microcapsules. The encapsulated nucleic acid, therefore, will be selected for the pH environment in which dissolution is desired and stored in a pH preselected to maintain stability.
- lipids comprise the inner coating of the microcapsules.
- these lipids may be, but are not limited to, partial esters of fatty acids and hexitiol anhydrides, and edible fats such as triglycerides. Lew C. W., Controlled-Release pH Sensitive Capsule And Adhesive System And Method. U.S. Pat. No. 5,364,634 (herein incorporated by reference).
- the present invention contemplates a microparticle comprising a gelatin, or other polymeric cation having a similar charge density to gelatin (i.e., poly-L- lysine) and is used as a complex to form a primary microparticle.
- a gelatin or other polymeric cation having a similar charge density to gelatin (i.e., poly-L- lysine) and is used as a complex to form a primary microparticle.
- a primary microparticle is produced as a mixture of the following composition: i) Gelatin (60 bloom, type A from porcine skin), ii) chondroitin 4-sulfate (0.005%-0.1%), iii) glutaraldehyde (25%, grade 1), and iv) 1- ethyl-3-(3-dimethylaminopropyl)-carbodiimide hydrochloride (EDC hydrochloride), and ultra-pure sucrose (Sigma Chemical Co., St. Louis, Mo.).
- the source of gelatin is not thought to be critical; it can be from bovine, porcine, human, or other animal source.
- the polymeric cation is between 19,000-30,000 daltons. Chondroitin sulfate is then added to the complex with sodium sulfate, or ethanol as a coacervation agent.
- a nucleic acid is directly bound to the surface of the microparticle or is indirectly attached using a "bridge" or "spacer".
- the amino groups of the gelatin lysine groups are easily derivatized to provide sites for direct coupling of a compound.
- spacers i.e., linking molecules and derivatizing moieties on targeting ligands
- avidin-biotin are also useful to indirectly couple targeting ligands to the microparticles.
- Stability of the microparticle is controlled by the amount of glutaraldehyde- spacer crosslinking induced by the EDC hydrochloride.
- a controlled release medium is also empirically determined by the final density of glutaraldehyde-spacer crosslinks.
- the present invention contemplates microparticles formed by spray-drying a composition comprising fibrinogen or thrombin with a nucleic acid.
- these microparticles are soluble and the selected protein (i.e., fibrinogen or thrombin) creates the walls of the microparticles. Consequently, the nucleic acids are incorporated within, and between, the protein walls of the microparticle. Heath et ah, Microparticles And Their Use In Wound Therapy. U.S. Pat. No. 6,113,948 (herein incorporated by reference).
- the subsequent reaction between the fibrinogen and thrombin creates a tissue sealant thereby releasing the incorporated compound into the immediate surrounding area.
- microparticles need not be exactly spherical; only as very small particles capable of being sprayed or spread into or onto a surgical site (i.e., either open or closed).
- microparticles are comprised of a biocompatible and/or biodegradable material selected from the group consisting of polylactide, polyglycolide and copolymers of lactide/glycolide (PLGA), hyaluronic acid, modified polysaccharides and any other well known material.
- polypeptides or polynucleotides of the disclosure such as the CIRT fusion proteins, stabilizer polypeptide, linker, RNA hairpin binding domain, NES, RNA regulatory domain, tag, NLS, RNA targeting molecule, hairpin region of the RNA targeting molecule, helical region, or targeting region of the RNA targeting molecule may include 1, 2, 3, 4, 5, 6,
- polypeptides or polynucleotides of the disclosure such as the CIRT fusion proteins, stabilizer polypeptide, linker, RNA hairpin binding domain, NES, RNA regulatory domain, tag, NLS, RNA targeting molecule, hairpin region of the RNA targeting molecule, helical region, or targeting region of the RNA targeting molecule may include 3, 4, 5, 6, 7, 8,
- the fusion protein may comprise amino acids 1 to 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81,
- 504 505, 506, 507, 508, 509, 510, 511, 512, 513, 514, 515, 516, 517, 518, 519, 520, 521, 522,
- the fusion protein may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10,
- the fusion protein may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10,
- the stabilizer polypeptide may comprise amino acids 1 to 2,
- the stabilizer polypeptide may comprise 1, 2, 3, 4, 5, 6, 7, 8,
- the stabilizer polypeptide may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58,
- the RNA hairpin binding domain may comprise amino acids 1 to 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53,
- the RNA hairpin binding domain may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56,
- the RNA hairpin binding domain may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56,
- SEQ ID NOs:7 or 18 contiguous amino acids of SEQ ID NOs:7 or 18 that are at least, at most, or exactly 60%, 61%, 62%, 63%, 64%, 65%, 66%, 67%, 68%, 69%, 70%, 71%, 72%, 73%, 74%, 75%, 76%, 77%, 78%, 79%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% similar, identical, or homologous with one of SEQ ID NOS:7 or 18.
- the RNA regulatory domain may comprise amino acids 1 to 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54,
- the RNA regulatory domain may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58,
- nucleic acid molecule or polypeptide starting at position 1 there is a nucleic acid molecule or polypeptide starting at position
- the RNA regulatory domain may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 66, 67, 68, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88, 89, 90, 91, 92, 93, 94, 95, 96, 97, 98, 99, 100, 101, 102, 103, 104, 105, 106, 107, 108, 109, 110, 111, 112, 113, 114
- the hairpin structure, such as the stem, loop, or both stem and loop, of the RNA targeting molecule may comprise nucleic acids 1 to 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, or 43 (or any derivable range therein) of SEQ ID NOs: l, 2, 26, 28, or 83-86.
- the hairpin structure, such as the stem, loop, or both stem and loop, of the RNA targeting molecule may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14,
- the hairpin structure, such as the stem, loop, or both stem and loop, of the RNA targeting molecule may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14,
- the RNA targeting region or the RNA targeting molecule may comprise nucleic acids 1 to 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, or 40 (or any derivable , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , ,
- the RNA targeting region or the RNA targeting molecule may comprise 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24,
- the stabilizer polypeptide may have one or more substitutions that reduce or eliminate binding to endogenous proteins. In some aspects, the stabilizer polypeptide may have one or more substitutions that reduce or eliminate an activity directed to an endogenous protein.
- the polypeptides and nucleic acids of the disclosure such as the CIRT fusion proteins, stabilizer polypeptide, linker, RNA hairpin binding domain, NES, RNA regulatory domain, tag, NLS, RNA targeting molecule, hairpin region of the RNA targeting molecule, helical region, or targeting region of the RNA targeting molecule may include at least, at most, or exactly 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25,
- substitution may be at amino acid position or nucleic acid position 1, 2, 3, 4, 5,
- Substitutional variants typically contain the exchange of one amino acid for another at one or more sites within the protein, and may be designed to modulate one or more properties of the polypeptide, with or without the loss of other functions or properties. Substitutions may be conservative, that is, one amino acid is replaced with one of similar shape and charge.
- Conservative substitutions are well known in the art and include, for example, the changes of: alanine to serine; arginine to lysine; asparagine to glutamine or histidine; aspartate to glutamate; cysteine to serine; glutamine to asparagine; glutamate to aspartate; glycine to proline; histidine to asparagine or glutamine; isoleucine to leucine or valine; leucine to valine or isoleucine; lysine to arginine; methionine to leucine or isoleucine; phenylalanine to tyrosine, leucine or methionine; serine to threonine; threonine to serine; tryptophan to tyrosine; tyrosine to tryptophan or phenylalanine; and valine to isoleucine or leucine.
- substitutions may be non-conservative such that a function or activity of the polypeptide is affected.
- Non conservative changes typically involve substituting a residue with one that is chemically dissimilar, such as a polar or charged amino acid for a nonpolar or uncharged amino acid, and vice versa.
- Proteins may be recombinant or synthesized in vitro. Alternatively, a non recombinant or recombinant protein may be isolated from bacteria. It is also contemplated that bacteria containing such a variant may be implemented in compositions and methods. Consequently, a protein need not be isolated.
- the term“functionally equivalent codon” is used herein to refer to codons that encode the same amino acid, such as the six codons for arginine or serine, and also refers to codons that encode biologically equivalent amino acids.
- amino acid and nucleic acid sequences may include additional residues, such as additional N- or C-terminal amino acids, or 5' or 3' sequences, respectively, and yet still be essentially as set forth in one of the sequences disclosed herein, so long as the sequence meets the criteria set forth above, including the maintenance of biological protein activity where protein expression is concerned.
- the addition of terminal sequences particularly applies to nucleic acid sequences that may, for example, include various non coding sequences flanking either of the 5' or 3' portions of the coding region.
- amino acids of a protein may be substituted for other amino acids in a protein structure without appreciable loss of interactive binding capacity.
- Structures such as, for example, an enzymatic catalytic domain or interaction components may have amino acid substituted to maintain such function. Since it is the interactive capacity and nature of a protein that defines that protein’s biological functional activity, certain amino acid substitutions can be made in a protein sequence, and in its underlying DNA coding sequence, and nevertheless produce a protein with like properties. It is thus contemplated by the inventors that various changes may be made in the DNA sequences of genes without appreciable loss of their biological utility or activity.
- alteration of the function of a polypeptide is intended by introducing one or more substitutions.
- certain amino acids may be substituted for other amino acids in a protein structure with the intent to modify the interactive binding capacity of interaction components. Structures such as, for example, protein interaction domains, nucleic acid interaction domains, and catalytic sites may have amino acids substituted to alter such function. Since it is the interactive capacity and nature of a protein that defines that protein’s biological functional activity, certain amino acid substitutions can be made in a protein sequence, and in its underlying DNA coding sequence, and nevertheless produce a protein with different properties. It is thus contemplated by the inventors that various changes may be made in the DNA sequences of genes with appreciable alteration of their biological utility or activity.
- the hydropathic index of amino acids may be considered.
- the importance of the hydropathic amino acid index in conferring interactive biologic function on a protein is generally understood in the art (Kyte and Doolittle, 1982). It is accepted that the relative hydropathic character of the amino acid contributes to the secondary structure of the resultant protein, which in turn defines the interaction of the protein with other molecules, for example, enzymes, substrates, receptors, DNA, antibodies, antigens, and the like.
- amino acid substitutions generally are based on the relative similarity of the amino acid side-chain substituents, for example, their hydrophobicity, hydrophilicity, charge, size, and the like.
- Exemplary substitutions that take into consideration the various foregoing characteristics are well known and include: arginine and lysine; glutamate and aspartate; serine and threonine; glutamine and asparagine; and valine, leucine and isoleucine.
- all or part of proteins described herein can also be synthesized in solution or on a solid support in accordance with conventional techniques.
- Various automatic synthesizers are commercially available and can be used in accordance with known protocols. See, for example, Stewart and Young, (1984); Tam etal. , (1983); Merrifield, (1986); and Barany and Merrifield (1979), each incorporated herein by reference.
- recombinant DNA technology may be employed wherein a nucleotide sequence that encodes a peptide or polypeptide is inserted into an expression vector, transformed or transfected into an appropriate host cell and cultivated under conditions suitable for expression.
- One embodiment includes the use of gene transfer to cells, including microorganisms, for the production and/or presentation of proteins.
- the gene for the protein of interest may be transferred into appropriate host cells followed by culture of cells under the appropriate conditions.
- a nucleic acid encoding virtually any polypeptide may be employed.
- the generation of recombinant expression vectors, and the elements included therein, are discussed herein.
- the protein to be produced may be an endogenous protein normally synthesized by the cell used for protein production.
- TAR hairpin Scaffold ggccagaucugagccugggagcucucuggcc (SEQ ID NO: 1)
- Human SLBP hairpin scaffold ccaaaggcucuucucagagccaccca (SEQ ID NO:2)
- gRNA sequences used in this study (all gRNAs are expressed from a hU6 promoter)
- gRNA guiding sequencing all expressed by the same hU6 promoter as TBP-OT
- RNA hairpin structure [0191]
- Nucleolin recognition sequence UCCCGA (SEQ ID NO:84).
- RNA hairpin structure GGCCGAAAUCCCGAAUGAGGCC (SEQ ID NO:85) and
- GGAUGCCUCCCGAGUGCAUCC (SEQ ID NO: 86).
- Example 1 Programmable RNA-guided RNA effector proteins built from human parts
- CIRTS CRISPR/Cas-inspired RNA targeting system
- the inventors show that CIRTS permits mining the human proteome for functional parts to build programmable RNA regulatory proteins.
- CIRTS is a ribonucleoprotein complex that uses Watson-Crick-Franklin base pair interactions to deliver protein cargo site-selectively in the transcriptome.
- the inventors show they can easily engineer CIRTS that deliver a range or regulatory proteins to transcripts, including nucleases for degradation, deadenylation regulatory machinery for degradation, or translational activation machinery for enhanced protein production.
- CIRTS are up to 5-fold smaller than the smallest current CRISPR/Cas systems and can be engineered entirely from human parts.
- RNA hairpin-binding protein that serves as the core of the system and is a selective, high affinity binder to a specific RNA structure displayed on an engineered gRNA
- a gRNA that features both the structure that interacts with the engineered hairpin-binding protein and a sequence with complementarity to the target RNA of interest
- an effector protein such as a nuclease or epitranscriptomic regulator, that acts on the targeted RNA in a proximity-dependent manner.
- a charged protein that binds to the displayed gRNA sequence non-specifically is used to stabilize and protect the guiding RNA prior to target engagement. (FIG. 1A).
- CIRTS combines multiple protein domains performing these functions in an engineered system.
- the inventors present the design and validation of CIRTS.
- the inventors engineered a programmable CIRTS ribonuclease, which they used for both in vitro and mammalian cell reporter assay optimization and validation.
- the inventors demonstrate the versatility of CIRTS by showing that all four of the constituent parts, including the gRNA, the hairpin binding protein, the ssRNA binding protein, and the effector domain of CIRTS- 1 can be substituted for other parts and developed five additional CIRTS (FIG. IB).
- the inventors are able to target endogenous epitranscriptomic regulatory pathways, including degradation machinery and translational activation, using CIRTS engineered as programmable m 6 A reader proteins.
- the inventors show that they can target endogenous transcripts for proximity-dependent nuclease-mediated decay, non-nuclease-mediated decay, or translation activation, using CIRTS. Finally, the inventors show they can target multiple genes simultaneously using orthogonal CIRTS. Taken together, this work validates the CIRTS strategy as a viable new approach to engineer RNA effector proteins that are small and assembled from human parts.
- CIRTS-1 first-generation system
- TBP6.7 human hairpin-binding protein U1A protein
- L nucleotide linker sequence
- FIG. 1C guiding sequence
- the inventors first engineered a programmable nuclease by fusing TBP6.7 to the Pin nuclease domain of human SMG6, which has been previously used as a non-specific proximity-dependent RNA endonuclease (Batra et al., 2017; Choudhury et al., 2012). Although this simplest design already displayed gRNA-mediated transcript degradation in cell-based assays (FIG. 9B, left), the performance was quite poor, which the inventors attributed to the potential degradation of the displayed targeting sequence. The protein surface and hairpin channel of Casl3 systems tend to be highly charged, likely to non-specifically bind and stabilize the guiding RNA sequence (Liu et al., 2017).
- CIRTS-1 is composed of a protein fusion complex of ORF5-TBP6.7-Pin nuclease domain along with a corresponding gRNA (FIG. 1 A).
- the inventors first characterized the programmable RNA binding and RNA nuclease activity of CIRTS-1 in vitro on model RNA target substrates. Using purified MBP-CIRTS-1 protein and gRNA in electrophoretic mobility shift assays (EMSA) run in the presence of EDTA to inactive the nuclease, the inventors found MBP-CIRTS-1 binds a target RNA in a gRNA-dependent manner with an apparent KD of 105 nM (FIG. 2A and 2B). If either the gRNA (FIG. 7A) or CIRTS-1 protein (FIG.
- the inventors To test the target nuclease activity of CIRTS-1 in live mammalian cells, the inventors established a dual luciferase reporter assay that reports on gRNA-dependent transcriptional changes on a target RNA (FIG. 3A). Using this system, the inventors optimized the deployment of CIRTS-1 by assaying different protein linker types (FIG. 8A), gRNA structures (FIG. 8C), and CIRTS-1 cellular localization (FIG. 9C). After optimization, the inventors compared the ability of optimized CIRTS-1 system to degrade the target reporter RNA to the current state- of-the-art Casl3 system.
- the inventors designed gRNAs that target firefly luciferase mRNAs in the dual-luciferase reporter assay for both CIRTS-1 and Casl3b, as well as control, off-target gRNAs for each programmable nuclease.
- CIRTS-0 which contains a previously reported ‘dead’ mutation in the nuclease domain of CIRTS-1 (Eberle et al., 2009), serving as a negative control.
- CIRTS-0 has no significant effect on the target transcript (FIG. 3B and FIG.
- the inventors next tested different protein domains for each component of the CIRTS protein delivery system.
- CIRTS RNA epitranscriptomic regulatory“reader” proteins, which they previously delivered using the dCasl3b system (Rauch et al., 2018).
- the inventors swapped the Pin nuclease effector protein of CIRTS-1 for the N-terminal domain of YTHDF1, a cytoplasmic A ⁇ - ethyl adenosine (m 6 A) reader protein that recruits the translation machinery, to generate CIRTS-2.
- m 6 A cytoplasmic A ⁇ - ethyl adenosine
- RNA levels are relatively unchanged (FIG. 9E), but a significant increase in protein levels from the RNA are generated (FIG. 3D).
- the inventors then exchanged the YTHDF1 fragment for a fragment of YTHDF2, an m 6 A reader protein that recruits the RNA deadenylation machinery and induces RNA degradation, to generate CIRTS-3.
- Delivery of CIRTS-3 to the reporter mRNA induces degradation of the target transcript as measured by both RNA (FIG. 9F) and protein levels (FIG. 3E).
- CIRTS-1 through -3 demonstrate the versatility of the design strategy to deliver a range of effector protein cargoes to target RNA in live cells.
- the inventors set out to assess if other human parts could also be used for the RNA hairpin binding domain and non specific ssRNA binding protein.
- the inventors replaced TBP6.7 in CIRTS-3 with the RNA hairpin binding domain of the human histone stem loop binding protein (SLBP) to generate CIRTS-4.
- the inventors designed a gRNA based on the histone mRNA stem loop structure (FIG. ID).
- Assaying CIRTS-4 in the reporter assay (FIG. 3F) and by RT-qPCR to assess RNA levels (FIG. 9G) revealed similar performance as CIRTS-3.
- the inventors sought to engineer entirely humanized versions of the CIRTS system.
- the inventors designed the first proof-of-concept systems based on the viral non-specific, single- stranded RNA binding protein, ORF5. Although there are no annotated human single-stranded, non-specific RNA binding proteins, the inventors reasoned highly charged, cationic human proteins could fulfill the role of ORF5 in the CIRTS system (Cronican et al., 2011). The inventors therefore engineered HBEGF and b-defensin 3, two cationic human proteins, in the place of ORF5 in CIRTS-3 to generate CIRTS-5 and CIRTS-6, respectively. Again, deploying these programmable effectors in the luciferase reporter assay revealed gRNA-dependent degradation of the target gene (FIG.
- the inventors also assayed changes of RNA levels by qPCR and found a decrease for both CRITS-5 and CIRTS-6 (FIG. 9H).
- CIRTS-1 through CIRTS-6 in the reporter assay demonstrates the modularity of the CIRTS design in all dimensions, including the hairpin-binding domain and corresponding gRNA, the single-stranded RNA binding protein, and the effector protein.
- the inventors sought to test whether the CIRTS can also be used to target endogenous transcripts.
- the inventors verified that CIRTS-0 could bind a target endogenous transcript by analysis by RNA immunoprecipitation followed by RT-qPCR.
- the inventors designed gRNAs to target two endogenous transcripts that were previously targeted by Casl3 systems, PPIB, and SMARCA4 (Cox et al., 2017; Konermann et al., 2018).
- the inventors separately delivered each gRNA along with CIRTS-0 fused to a 3x FLAG-tag.
- the inventors then isolated total RNA, immunoprecipitated with an anti-Flag antibody, and quantified the relative amounts of each target RNA bound to the protein.
- both endogenous transcripts were enriched between 2.5- and 5-fold in a gRNA-dependent manner (FIG. 10A), validating CIRTS can function as a programmable RNA-guided RNA binding protein on endogenous transcripts.
- the inventors next sought to assess whether the CIRTS system could deliver an effector protein to a target endogenous transcript, using the CIRTS- 1 programmable nuclease and CIRTS-3 programmable YTHDF2-mediated decay systems as exemplars.
- the inventors selected five RNA transcripts that have been previously validated as Casl3 targets, reasoning that these are accessible for RNA targeting by programmable RNA-binding systems.
- the inventors then designed gRNAs for each target, using the same binding sites on the targets that were previously used in Casl3 experiments, postulating these sites would also be accessible to CIRTS targeting (Konermann et al., 2018).
- the inventors assayed the effects of the CIRT system on RNA levels of each target by RT-qPCR.
- RT-qPCR When cells were transfected with either CIRTS-1 or CIRTS-3, along with a specific gRNA expressing vector, the inventors saw a significant decrease in RNA level by RT-qPCR for each of the five endogenous transcripts: RRP3, NFKB1, NRAS, B4GALTN1, and SMARCA4 (FIG. 4A and 4B).
- CIRTS-2 could trigger protein production of endogenous transcripts through a YTHDF1 -mediated epitranscriptomic pathways.
- the inventors selected an abundant transcript (CypB, the protein product of PPIB) with a reported, reliable antibody for analysis of protein production by Western blotting. Indeed, cells transfected with CIRTS-2 and an on-target gRNA showed an increase in protein level (FIG. 4C) without a change in RNA levels (FIG. 10B), confirming the YTHDF1 effect on the transcript.
- CypB the protein product of PPIB
- the CIRTS platform is functional on endogenous transcripts in a gRNA-dependent manner, and can actively degrade a target transcript, trigger degradation machinery to act on the target transcript, or to active translation and increased protein production from the target transcript.
- the versatility of the CIRTS inspired the inventors to deploy them simultaneously to target more than one transcript.
- the inventors aimed to assess whether CIRTS engineered with different hairpin binding domains could functional orthogonally in live cells to selectively target different transcripts.
- the CIRTS built from the TBP hairpin binding domain and the CIRTS built from the SLBP hairpin binding domain, which each use separately engineered gRNAs (FIG. 1C and ID) should be orthogonal to one another.
- the inventors deployed two fully-humanized CIRTS that each deliver YTHDF2, CIRTS-6 and CIRTS-7, which each bind different hairpin structures and should have minimal crosstalk between one another.
- the inventors aimed to use CIRTS-6 to target PPIB and CIRTS-7 to target SMARCA4 (FIG. 5A).
- the inventors co-transfected cells with expression vectors for both CIRTS proteins, along with expression vectors for gRNAs for each CIRTS displaying either a control, non-targeted gRNA sequence, or a gRNA sequence targeting the respective endogenous gene.
- the inventors then assessed changes in relative RNA levels of the two target genes by RT-qPCR. Indeed, each CIRTS system degraded the target transcript in an on-target gRNA-dependent manner, without any crosstalk between the two systems (FIG. 5B).
- the inventors were able to selectively degrade one transcript or the other in the presence of both CIRTS proteins and were also able to simultaneously degrade both transcripts.
- RNA degradation of CIRTS-6 showed levels comparable to when only CIRTS-6 is delivered to cells (FIG. 11 A), targeting with CIRTS-7, however, resulted in slightly reduced RNA degradation as compared with individual transfection (FIG. 11B).
- the inventors transfected cells sequentially on two consecutive days with a CIRTS and a gRNA vector each day. The inventors attribute this decreased activity to decreased transfection efficiency on the second day of transfection with CIRTS-7.
- the SLBP-based constructs could theoretically still bind to cellular histone mRNA, which could lead to a decrease in overall activity.
- both CIRTS were delivered with on-target gRNA, they performed at their best in this assay.
- the inventors conclude that the two orthogonal CIRTS systems can each be orthogonal to one another and can each simultaneously target endogenous transcripts in a gRNA-dependent manner the inventors are currently working on engineering orthogonal CIRTS that have no more endogenous binding partners and can act without potentially perturbing cellular events.
- CIRTS a new strategy for engineering programmable RNA effector proteins.
- CIRTS are small, can be fully humanized, can target endogenous RNAs in live cells, and can work orthogonally and synergistically together for multidimensional control.
- CIRTS should provide advantages to previous methods because of their smaller size.
- CIRTS-2 and CIRTS-3 are 65 and 36 kDa respectively, while the comparable Casl3b-based programmable YTHDF1 and YTHDF2 systems are 155 and 126 kDa, respectively (FIG. 6).
- CIRTS-1 is even smaller than the smallest DNA-targeting Cas protein found to date, Casl4a (Harrington et ak, 2018).
- the smaller size of the delivery system should be less perturbative to the endogenous transcript under study presenting opportunities for uncovering the roles of RNA regulatory proteins in live cells.
- the CIRTS should offer several key advantages and opportunities.
- the humanized nature of the CIRTS will provide a pathway toward avoiding immune responses, opening up the potential for continuously-delivered therapies.
- accessory proteins such as human b-defensin 3
- this protein has been extensively studied and the structural bases for its function has been elucidated (Dhople et ak, 2006; Kluver et ak, 2005). This knowledge will enable one skilled in the art to engineer a b-defensin 3 peptide that retains its charged nature but abolishes its endogenous functions.
- the small size of the CIRTS will still allow for multiple regulatory proteins to be simultaneously delivered in a viral delivery system, for example to target one transcript for degradation and another for translational activation.
- the multiplexable capacity of the CIRTS couple with the small size and diversity of effector proteins that can be delivered opens up possibilities for cell reprogramming by targeting multiple genes at once in multiple dimensions.
- the CIRTS platform demonstrates the potential of combining parts contained within the human protein toolbox to engineer proteins with new properties.
- the CIRTS system provides a new approach for studying and exploiting RNA regulation and will open up many future opportunities to intervene in cell regulation for disease treatment.
- Example 2 Programmable RNA-guided RNA effector proteins built from human parts A. RESULTS
- RNA targeting is key requirements for RNA targeting.
- the inventors sought to engineer a Cas 13 -inspired system that uses a defined protein-RNA interaction to display a gRNA sequence to deliver protein cargoes to a target RNA, similar to previous RNA tethering assays with overexpressed reporter constructs (Coller and Wickens, 2007). Indeed, hairpin-binding proteins and covalent RNA fusions have been used to deliver RNA editing machinery to transcripts (Monti el -Gonzalez et al., 2016; Sinnamon et al., 2017; Vogel et al., 2018).
- RNA hairpin-binding protein that serves as the core of the system and is a selective, high affinity binder to a specific RNA structure displayed on an engineered gRNA
- a gRNA that features both the structure that interacts with the engineered hairpin-binding protein and a sequence with complementarity to the target RNA of interest
- a charged protein that could bind to the displayed gRNA sequence non-specifically to stabilize and protect the guiding RNA prior to target engagement
- an effector protein such as a ribonuclease or epitranscriptomic regulator, that acts on the targeted RNA in a proximity-
- CRISPR/Cas-inspired RNA targeting system CRISPR/Cas-inspired RNA targeting system
- CIRTS-1 For the first-generation system, CIRTS-1, the inventors used an evolved version of the human hairpin-binding protein U1 A protein (TBP6.7), which was previously engineered to bind the HIV trans-activation response (TAR) hairpin and has no endogenous human RNA hairpin targets (Blakeley and McNaughton, 2014; Crawford et al., 2016) (FIG. 12B). The inventors designed a gRNA that includes the TAR hairpin, a nucleotide linker sequence (L), and then a guiding sequence (FIG. 12C).
- TBP6.7 human hairpin-binding protein
- L nucleotide linker sequence
- FIG. 12C guiding sequence
- the inventors first engineered a programmable ribonuclease by fusing TBP6.7 to the Pin nuclease domain of human nonsense-mediated mRNA decay factor SMG6, which has been previously used as a non-specific proximity-dependent RNA endonuclease (Batra et al., 2017; Choudhury et al., 2012). Although this simplest design already displayed promising gRNA-mediated transcript degradation in cell-based luciferase assays (FIG. 22A, left), the performance was quite poor, which the inventors attributed to the degradation of the displayed guiding sequence.
- CIRTS-1 is composed of a protein fusion complex of ORF5-TBP6.7-Pin nuclease domain along with a corresponding gRNA.
- the inventors first characterized the programmable RNA binding and RNA nuclease activity of CIRTS-1 in vitro on model RNA target substrates.
- the Pin nuclease domain was previously shown to be active in the presence of Mn2+ and activity could be quenched by the addition of EDTA (Choudhury et al., 2012).
- Directly overexpressing CIRTS-1 led to insoluble protein in the cell pellet, which the inventors resolved by fusing an N-terminal MBP tag to CIRTS-1 (MBP-CIRTS-1).
- MBP-CIRTS-1 binds a target RNA in a gRNA-dependent manner with an apparent binding dissociation constant (KD) of 22 nM (FIG. 13 A).
- KD apparent binding dissociation constant
- the inventors see ⁇ 50-fold weaker binding.
- MBP-CIRTS-1 degrades an RNA substrate in a gRNA- and Mn2+-dependent manner, confirming the activity of the Pin ribonuclease domain in the fusion context (FIG. 13B).
- FIG. 14A To test the target nuclease activity of CIRTS-1 in live mammalian cells, the inventors established a dual luciferase reporter assay that reports on gRNA-dependent transcriptional changes on a target firefly luciferase (Flue) RNA (FIG. 14A). Using this system, the inventors optimized the deployment of the Pin nuclease CIRTS by assaying different protein linker types (FIG. 23 A), gRNA structures and lengths (FIG. 23C), and CIRTS nuclease cellular localization (FIG. 22B).
- Flue firefly luciferase
- the inventors compared the ability of optimized CIRTS-1 (Pin nuclease) system to degrade the target reporter RNA to the Casl3b system (Cox et ah, 2017).
- the inventors designed gRNAs that target the firefly luciferase mRNA in the dual-luciferase reporter assay for both CIRTS-1 and Casl3b, as well as control, non-targeting gRNAs (targeting a lambda phage sequence) for each programmable nuclease.
- CIRTS-0 which contains a previously reported‘dead’ mutation in the Pin nuclease domain of CIRTS-1 (Eberle et ah, 2009), serving as a negative control.
- CIRTS-1 containing an active nuclease, could mediate degradation of the target.
- CIRTS-1 Pin nuclease
- the inventors next sought to assess the versatility of the design by testing whether each component of the system, including the gRNA, hairpin binding domain, non-specific RNA binding domain, and effector protein, could be swapped for other parts to achieve CIRTS with diverse functions.
- the inventors next tested different protein domains for each component of the CIRT protein delivery system.
- the inventors assayed whether CIRTS could deliver RNA epitranscriptomic regulatory“reader” proteins, which the inventors previously delivered using the dCasl3b system (Rauch et ak, 2018).
- the inventors chose to focus on regulatory proteins of N6-methyladenosine, the most prevalent mRNA modification.
- each transcript contains three modifications sites with high m6A abundance detected in the 3’UTR, and m6A has been shown to have regulatory roles in splicing (Xiao et ak, 2016), translation (Meyer et ak, 2015; Wang et ak, 2015), and stability (Du et ak, 2016; Wang et ak, 2013).
- the inventors exchanged the Pin nuclease effector protein of CIRTS-1 for the N-terminal domain of the YT521-B homology domain family protein 1 (YTHDF1), a cytoplasmic m6A reader protein that recruits the translation machinery (Wang et al., 2015), to generate CIRTS-2.
- YTHDF1 YT521-B homology domain family protein 1
- the inventors then exchanged the YTHDF1 fragment for a fragment of YTHDF2, a different m6A reader protein that recruits the RNA deadenylation machinery and induces RNA degradation (Du et al., 2016; Wang et al., 2013), to generate CIRTS-3.
- Delivery of CIRTS-3 (YTHDF2) to the reporter mRNA induces degradation of the target transcript as measured by both RNA (FIG. 22F) and protein levels (FIG. 14E).
- CIRTS-1 through -3 demonstrate the versatility of the design strategy to deliver a range of effector protein cargoes to target RNA in live cells.
- the inventors set out to assess if other human parts could also be used for the RNA hairpin binding domain and non specific ssRNA binding protein.
- the inventors replaced TBP6.7 in CIRTS-3 (YTHDF2) with the RNA hairpin binding domain of the human histone stem loop binding protein (SLBP) to generate CIRTS-4 (YTHDF2).
- CIRTS-4 YTHDF2
- the inventors designed a gRNA based on the histone mRNA stem loop structure (FIG. 12D).
- Assaying CIRTS-4 (YTHDF2) in the reporter assay (FIG. 14F) and by RT-qPCR to assess RNA levels (FIG. 22G) revealed similar performance as CIRTS-3, confirming other hairpin binding domains can be used as the core of the CIRTS.
- the inventors sought to engineer entirely humanized versions of the CIRTS system.
- the inventors designed the initial proof-of-concept systems based on the viral non-specific, single-stranded RNA binding protein, ORF5.
- ORF5 the viral non-specific, single-stranded RNA binding protein
- the inventors therefore engineered HBEGF and b-defensin 3, two cationic human proteins, in the place of ORF5 in CIRTS-3 to generate CIRTS-5 and CIRTS- 6, respectively.
- HBEGF and b-defensin 3 two cationic human proteins
- the inventors used CIRTS to deliver the catalytic domain of human ADAR2 (hADAR2) to RNA transcripts to confirm CIRTS’ versatility in scope of functions with an additional effector protein.
- the inventors designed a dual luciferase reporter that contains a G- to-A mutation in the coding region of firefly luciferase resulting in a premature stop of translation and no measurable firefly luciferase activity (FIG. 15A and 22J).
- the inventors then deployed CIRTS to deliver wt ADAR2 (CIRTS-7) or hADAR 2 E488Q (CIRTS-8), a known hyperactive mutant of hADAR2 (Kuttan and Bass, 2012), to the mutated position, which resulted in gRNA-dependent rescue of luciferase activity in both cases (FIG. 15B).
- the hyperactive hADAR2 mutant showed higher editing efficiency and a higher background in the absence of an on-target gRNA based on luciferase assay.
- using the hyperactive mutant could be beneficial to allow targeting of a wider substrate scope as it has relaxed sequence constraints (Kuttan and Bass, 2012).
- the inventors next sought to assess whether the CIRTS system could deliver an effector protein to a target endogenous transcript, using the CIRTS- 1 programmable nuclease and CIRTS-3 programmable YTHDF2-mediated decay systems as exemplars.
- the inventors selected five RNA transcripts that have been previously validated as Casl3 targets, reasoning that these are accessible for RNA targeting by programmable RNA-binding systems.
- the inventors then designed gRNAs for each target, using the same binding sites on the targets that were previously used in Casl3 experiments (Abudayyeh et ak, 2017; Konermann et ah, 2018).
- the inventors assayed the effects of the CIRT system on RNA levels of each target by RT- qPCR.
- the inventors observed a significant decrease in RNA level by RT- qPCR for each of the five endogenous mRNA transcripts: RRP3, NFKB 1, NRAS, B4GALNT1, and SMARCA4 (FIG. 16A and 16B).
- the inventors also verified that the inventors can target other RNA species such as IncRNA by targeting CIRTS- 1 (Pin nuclease) to MALATl (FIG. 24A).
- the inventors set out to assess whether CIRTS-2 could trigger protein production of an endogenous transcript through a YTHDF1 -mediated epitranscriptomic pathways.
- the inventors selected an abundant transcript PPIB with a reported, reliable antibody for analysis of CypB (the protein product of PPIB) protein production by Western blotting. Indeed, cells transfected with CIRTS-2 and an on-target gRNA showed an increase in protein level (FIG. 16C and FIG. 24C) without a change in RNA levels (FIG. 24B), consistent with prior reported YTHDF1 effects on the transcript (Wang et al., 2015). As a control, the same experiment performed with CIRTS-3, which delivers YTHDF2, results in slight decrease in protein levels, which correlates with the decrease in mRNA levels (FIG. 16B and 24C).
- the inventors designed a series of experiments that address the sensitivity of the gRNA to mismatches, transcriptome-wide off-targets, and endogenous substrate targeting.
- the inventors designed a luciferase-based mismatch experiment that allows the inventors to assay targeting effects when introducing one, two, or three mismatches into the duplex formed between gRNA and target RNA.
- the inventors chose to fuse the disease relevant KRAS4b transcript to the luciferase reporter and asked whether the engineered system can differentiate between the cancer-associated G12D (target 1), the wild type (target 2), the G12C (target 3), and a G12W (target 4) KRAS4b variants (FIG. 25A).
- the inventors next assessed whether increasing the gRNA length could affect the mismatch tolerance of CIRTS, focusing on mismatches in the center region of the duplex formed between gRNA and target RNA as they showed the largest effect on knockdown efficiency in the assay. As observed with the shorter 20 nt gRNA, the inventors see no difference in knockdown efficiency when the inventors target a reporter with no or one mismatches. However, a longer 40 nt gRNA can rescue some of the effects when the two- mismatch variant was targeted, indicating that the gRNA length contributes to the specificity of the system (FIG. 25D).
- the inventors furthermore performed RNA immunoprecipitation followed by RT-qPCR.
- the inventors designed gRNAs to target two endogenous transcripts that were previously targeted by Casl3 systems, PPIB, and B4GALNT1 (Cox et ah, 2017; Konermann et ah, 2018).
- the inventors separately delivered each gRNA along with CIRTS-0 (dead nuclease CIRTS) fused to a 3x FLAG-tag.
- the inventors then subjected lysates to immunoprecipitation with an anti -Flag antibody, and quantified the relative amounts of each target RNA bound to the protein.
- both endogenous transcripts were enriched between 2.5- and 5-fold in a gRNA-dependent manner (FIG. 25H), confirming CIRTS function as a programmable RNA-guided RNA binding protein on endogenous transcripts.
- CIRTS complementary metal-oxide-semiconductor
- TBP hairpin binding domain and CIRTS built from the SLBP hairpin binding domain, which each use separately engineered gRNAs (FIG. 12C and 12D), should be orthogonal to one another, permitting selective targeting of multiple transcripts with either the same or even different CIRTS.
- the inventors co-transfected cells with CIRTS-6 along with three gRNAs targeting PPIB, SMARCA4, and NRAS and assessed changes in RNA level by RT- qPCR.
- the inventors observed a decrease in RNA levels for all three targeted transcripts (FIG. 18A and 18B).
- the inventors observed a slight decrease in efficiency when the inventors deploy several gRNAs or CIRTS in the same cells.
- CIRTS-9 a fully humanized version of the YTHDF1 construct
- CIRTS- 10 a fully humanized version of the YTHDF1 construct
- SMARCA4 a fully humanized version of the YTHDF1 construct
- the inventors find that both proteins are active and induce the anticipated increase in luciferase protein and decrease in RNA levels respectively.
- CIRTS-6 TBP6.7
- CIRTS-10 SMARCA4
- TBP6.7 and SLBP-based CIRTS can each simultaneously target endogenous transcripts in a gRNA-dependent manner.
- the inventors are currently working on engineering orthogonal CIRTS that have no endogenous binding partners by evolving human proteins toward new specificities, as was done with TBP6.7.
- CIRTS allow for multiple regulatory proteins to be simultaneously delivered, for example to target one transcript for degradation and another for translational activation, opening up possibilities for cell reprogramming by targeting multiple genes at once in multiple dimensions (Bao et al., 2017; Gao et al., 2016).
- Adenovirus-associated virus is a versatile delivery vehicle to deliver transgenes and gene therapies to different cell types due to wide range of serotypes available (Gao et al., 2005), low immune response stimulation (Vasileva and Jessberger, 2005), and low risk of genome insertion (Gao et al., 2005; Naso et al., 2017).
- serotypes available (Gao et al., 2005), low immune response stimulation (Vasileva and Jessberger, 2005), and low risk of genome insertion.
- CIRTS a versatile strategy for engineering programmable RNA effector proteins.
- CIRTS are small, can be fully humanized, can target endogenous RNAs in live cells, and can work for multidimensional transcriptome control.
- CIRTS should provide advantages to previous methods because of their smaller size.
- CIRTS-2 and CIRTS-3 are 65 and 36 kDa respectively, while the comparable Casl3b-based programmable YTHDF1 and YTHDF2 systems are 155 and 126 kDa, respectively (FIG. 20).
- CIRTS-1 is even smaller than the smallest DNA-targeting Cas protein found to date, Cas 14a and the smallest Casl2g RNA-targeting protein (Harrington et al., 2018; Yan et al., 2019).
- CIRTS should offer several key advantages and opportunities.
- the humanized nature of the CIRTS will provide a pathway toward avoiding immune responses, opening up the potential for continuously-delivered therapies. While the fusions between the human proteins in the CIRTS present potential limitations in the design where the immune system could respond to (Glaesner et al., 2010), this is a problem that can in principle be engineered around.
- the inventors computationally predicted the immunogenicity of the highest likelihood MHC I binding peptides in the inventors’ engineered constructs, the inventors find that the fully humanized CIRTS shows lower propensity to cause immune reactions (FIG. 27C), but further experimental testing is needed to discover where the limitations in the design emerge.
- the alternative hairpin binding protein SLBP in its current form has an endogenous RNA hairpin binding partner, which could influence stem loop RNA trafficking.
- the inventors only included the minimal RNA recognition motif (RRM) necessary for hairpin recognition in the system and omitted regions of potential interactions with other proteins or nucleic acids.
- RRM minimal RNA recognition motif
- the stem loop hairpin was already very short and could not be further truncated but the inventors chose to only use the minimally required region necessary for TBP6.7 binding to the TAR hairpin, resulting in a gRNA with less than half the original hairpin length.
- the cationic peptide, b-defensin 3 in its current form can theoretically still interact with its intracellular binding partners and elicit unwanted biological responses.
- human b- defensin 3 has been extensively studied and the structural bases for its function has been elucidated (Dhople et al., 2006; Kluver et al., 2005). The inventors can leverage this knowledge as the basis to engineer b-defensin 3 mutants that retain the highly charged nature required for CIRTS, but abolish endogenous functions, in order to engineer a human part-based, orthogonal RNA targeting system.
- the CIRTS platform demonstrates the potential of combining parts contained within the human protein toolbox to engineer proteins with new properties.
- the presented CIRTS were created through minimal protein engineering and optimization efforts, but function nearly as well as the naturally-evolved CRISPR/Cas systems.
- the inventors compared the Casl3b-based knockdown by its endogenous nuclease and by delivering YTHDF2 (FIG.
- RNA control properties there are a range of other regulatory proteins contained within the human proteome that likely house unique RNA control properties (Dominguez et ak, 2018), which can be coupled with CIRTS to create programmable versions of each protein for both functional characterization and potential translational applications.
- the CIRTS system provides a new approach for studying and exploiting RNA regulation and will open up future opportunities to intervene in cell regulation for disease treatment.
- C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573.
- Tethered function assays an adaptable approach to study RNA regulatory proteins. Methods Enzymol 429, 299-321.
- SMG6 promotes endonucleolytic cleavage of nonsense mRNA in human cells. Nat Struct Mol Biol 16, 49-55.
- TALENs a widely applicable technology for targeted genome editing. Nature Reviews Molecular Cell Biology 14, 49.
- CRISPR RNAs trigger innate immune responses in human cells. Genome Res.
- RNA Binding and HEPN-Nuclease Activation Are Decoupled in CRISPR-Casl3a. Cell Rep 24, 1025-1036.
- Casl3d Is a Compact RNA-Targeting Type VI CRISPR Effector Positively Modulated by a WYL-Domain-Containing Accessory Protein. Mol Cell 70, 327-339 e325.
- MHC class I presented peptides that enhance immunogenicity.
- YTHDF2 destabilizes m6A-containing RNA through direct recruitment of the CCR4-NOT deadenylase complex. Nat Commun 7, 12626.
- NetMHCpan-4.0 Improved Peptide-MHC Class I Interaction Predictions Integrating Eluted Ligand and Peptide Binding Affinity Data. J Immunol 199, 3360-3368.
- Adeno-Associated Virus as a Vector for Gene Therapy. BioDrugs 31, 317-334.
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Genetics & Genomics (AREA)
- Engineering & Computer Science (AREA)
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Zoology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Biomedical Technology (AREA)
- Wood Science & Technology (AREA)
- Molecular Biology (AREA)
- General Engineering & Computer Science (AREA)
- Biotechnology (AREA)
- General Health & Medical Sciences (AREA)
- Biochemistry (AREA)
- Microbiology (AREA)
- Biophysics (AREA)
- Medicinal Chemistry (AREA)
- Physics & Mathematics (AREA)
- Plant Pathology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Gastroenterology & Hepatology (AREA)
- Toxicology (AREA)
- Virology (AREA)
- Immunology (AREA)
- Pharmacology & Pharmacy (AREA)
- Epidemiology (AREA)
- Animal Behavior & Ethology (AREA)
- Public Health (AREA)
- Veterinary Medicine (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
- Medicines That Contain Protein Lipid Enzymes And Other Medicines (AREA)
- Pharmaceuticals Containing Other Organic And Inorganic Compounds (AREA)
- Medicinal Preparation (AREA)
- Medicines Containing Material From Animals Or Micro-Organisms (AREA)
- Peptides Or Proteins (AREA)
- Breeding Of Plants And Reproduction By Means Of Culturing (AREA)
- Enzymes And Modification Thereof (AREA)
Abstract
Description
Claims
Priority Applications (13)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| CA3125299A CA3125299A1 (en) | 2019-01-04 | 2020-01-03 | Systems and methods for modulating rna |
| SG11202107209WA SG11202107209WA (en) | 2019-01-04 | 2020-01-03 | Systems and methods for modulating rna |
| US17/309,936 US20220048962A1 (en) | 2019-01-04 | 2020-01-03 | Systems and methods for modulating rna |
| EP20736077.7A EP3906042A4 (en) | 2019-01-04 | 2020-01-03 | SYSTEMS AND METHODS FOR RNA MODULATION |
| AU2020204917A AU2020204917A1 (en) | 2019-01-04 | 2020-01-03 | Systems and methods for modulating RNA |
| KR1020217024731A KR20210124238A (en) | 2019-01-04 | 2020-01-03 | System and method for regulating RNA |
| MX2021008153A MX2021008153A (en) | 2019-01-04 | 2020-01-03 | Systems and methods for modulating rna. |
| JP2021538952A JP2022516919A (en) | 2019-01-04 | 2020-01-03 | Systems and methods for modulating RNA |
| CN202080018393.2A CN113543797A (en) | 2019-01-04 | 2020-01-03 | Systems and methods for regulating RNA |
| BR112021013173-6A BR112021013173A2 (en) | 2019-01-04 | 2020-01-03 | SYSTEMS AND METHODS TO MODULATE RNA |
| IL284516A IL284516A (en) | 2019-01-04 | 2021-06-30 | Systems and methods for modulating rna |
| JP2025030993A JP2025084877A (en) | 2019-01-04 | 2025-02-28 | Systems and methods for modulating RNA |
| AU2026200853A AU2026200853A1 (en) | 2019-01-04 | 2026-02-05 | Systems and methods for modulating RNA |
Applications Claiming Priority (8)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US201962788571P | 2019-01-04 | 2019-01-04 | |
| US62/788,571 | 2019-01-04 | ||
| US201962831342P | 2019-04-09 | 2019-04-09 | |
| US62/831,342 | 2019-04-09 | ||
| US201962903080P | 2019-09-20 | 2019-09-20 | |
| US62/903,080 | 2019-09-20 | ||
| US201962929339P | 2019-11-01 | 2019-11-01 | |
| US62/929,339 | 2019-11-01 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2020142676A1 true WO2020142676A1 (en) | 2020-07-09 |
Family
ID=71407144
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2020/012169 Ceased WO2020142676A1 (en) | 2019-01-04 | 2020-01-03 | Systems and methods for modulating rna |
Country Status (12)
| Country | Link |
|---|---|
| US (1) | US20220048962A1 (en) |
| EP (1) | EP3906042A4 (en) |
| JP (2) | JP2022516919A (en) |
| KR (1) | KR20210124238A (en) |
| CN (1) | CN113543797A (en) |
| AU (2) | AU2020204917A1 (en) |
| BR (1) | BR112021013173A2 (en) |
| CA (1) | CA3125299A1 (en) |
| IL (1) | IL284516A (en) |
| MX (1) | MX2021008153A (en) |
| SG (1) | SG11202107209WA (en) |
| WO (1) | WO2020142676A1 (en) |
Cited By (8)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20210095296A1 (en) * | 2019-09-29 | 2021-04-01 | Technion Research & Development Foundation Limited | Synthetic non-coding rnas |
| CN113583136A (en) * | 2021-04-25 | 2021-11-02 | 南方科技大学 | Phytohormone abscisic acid fluorescent molecular probe and application thereof |
| JP2023040984A (en) * | 2021-09-10 | 2023-03-23 | 国立大学法人信州大学 | Methods and kits for editing rna |
| WO2023057777A1 (en) * | 2021-10-08 | 2023-04-13 | Pencil Biosciences Limited | Synthetic genome editing system |
| WO2023150131A1 (en) * | 2022-02-01 | 2023-08-10 | The Regents Of The University Of California | Method of regulating alternative polyadenylation in rna |
| EP4110918A4 (en) * | 2020-02-28 | 2024-04-03 | The University of Chicago | METHODS AND COMPOSITIONS CONTAINING TRANSLATION ACTIVATORS |
| US20250051788A1 (en) * | 2021-05-24 | 2025-02-13 | Donald Danforth Plant Science Center | Compositions for rna-protein tethering and methods of using |
| WO2025168955A1 (en) | 2024-02-09 | 2025-08-14 | Ucl Business Ltd | Therapy for dravet syndrome by targeting scn1a antisense transcript |
Families Citing this family (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN114085837B (en) * | 2021-11-19 | 2024-04-12 | 中山大学 | Cell line for knocking out gene YTHDF1 and construction method thereof |
| WO2023164628A1 (en) * | 2022-02-25 | 2023-08-31 | The University Of Chicago | Methods and compositions for activating translation |
| CN114685685B (en) * | 2022-04-20 | 2023-11-07 | 上海科技大学 | Fusion protein for editing RNA and application thereof |
| WO2023215712A2 (en) * | 2022-05-02 | 2023-11-09 | The University Of Chicago | Systems and methods for modulating rna |
| CN121195069A (en) * | 2023-03-31 | 2025-12-23 | 杜克大学 | Compositions and methods for engineered transcriptomes |
| CN119899820B (en) * | 2025-01-23 | 2026-04-03 | 深圳开悦生命科技有限公司 | DHX8 fusion protein and its preparation method |
Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20170191082A1 (en) * | 2012-12-06 | 2017-07-06 | Sigma-Aldrich Co. Llc | Crispr-based genome modification and regulation |
| WO2017155408A1 (en) * | 2016-03-11 | 2017-09-14 | Erasmus University Medical Center Rotterdam | Improved crispr-cas9 genome editing tool |
| US20180291383A1 (en) * | 2013-04-04 | 2018-10-11 | President And Fellows Of Harvard College | THERAPEUTIC USES OF GENOME EDITING WITH CRISPR/Cas SYSTEMS |
Family Cites Families (6)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20040077082A1 (en) * | 2002-10-18 | 2004-04-22 | Koehn Richard K. | RNA-based inhibitory oligonucleotides |
| EP3341410B1 (en) * | 2015-08-24 | 2021-06-02 | Cellectis | Chimeric antigen receptors with integrated controllable functions |
| AU2017268842B2 (en) * | 2016-05-27 | 2022-08-11 | Aadigen, Llc | Peptides and nanoparticles for intracellular delivery of genome-editing molecules |
| GB2574769A (en) * | 2017-03-03 | 2019-12-18 | Univ California | RNA Targeting of mutations via suppressor tRNAs and deaminases |
| AU2018251801B2 (en) * | 2017-04-12 | 2024-11-07 | Massachusetts Institute Of Technology | Novel type VI CRISPR orthologs and systems |
| CN108559738A (en) * | 2018-02-12 | 2018-09-21 | 南昌大学 | A kind of system and method for plant RNA modification and editor |
-
2020
- 2020-01-03 WO PCT/US2020/012169 patent/WO2020142676A1/en not_active Ceased
- 2020-01-03 CN CN202080018393.2A patent/CN113543797A/en active Pending
- 2020-01-03 MX MX2021008153A patent/MX2021008153A/en unknown
- 2020-01-03 BR BR112021013173-6A patent/BR112021013173A2/en unknown
- 2020-01-03 US US17/309,936 patent/US20220048962A1/en active Pending
- 2020-01-03 EP EP20736077.7A patent/EP3906042A4/en active Pending
- 2020-01-03 JP JP2021538952A patent/JP2022516919A/en active Pending
- 2020-01-03 CA CA3125299A patent/CA3125299A1/en active Pending
- 2020-01-03 SG SG11202107209WA patent/SG11202107209WA/en unknown
- 2020-01-03 AU AU2020204917A patent/AU2020204917A1/en not_active Abandoned
- 2020-01-03 KR KR1020217024731A patent/KR20210124238A/en active Pending
-
2021
- 2021-06-30 IL IL284516A patent/IL284516A/en unknown
-
2025
- 2025-02-28 JP JP2025030993A patent/JP2025084877A/en active Pending
-
2026
- 2026-02-05 AU AU2026200853A patent/AU2026200853A1/en active Pending
Patent Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20170191082A1 (en) * | 2012-12-06 | 2017-07-06 | Sigma-Aldrich Co. Llc | Crispr-based genome modification and regulation |
| US20180291383A1 (en) * | 2013-04-04 | 2018-10-11 | President And Fellows Of Harvard College | THERAPEUTIC USES OF GENOME EDITING WITH CRISPR/Cas SYSTEMS |
| WO2017155408A1 (en) * | 2016-03-11 | 2017-09-14 | Erasmus University Medical Center Rotterdam | Improved crispr-cas9 genome editing tool |
Non-Patent Citations (1)
| Title |
|---|
| See also references of EP3906042A4 * |
Cited By (9)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US20210095296A1 (en) * | 2019-09-29 | 2021-04-01 | Technion Research & Development Foundation Limited | Synthetic non-coding rnas |
| EP4110918A4 (en) * | 2020-02-28 | 2024-04-03 | The University of Chicago | METHODS AND COMPOSITIONS CONTAINING TRANSLATION ACTIVATORS |
| CN113583136A (en) * | 2021-04-25 | 2021-11-02 | 南方科技大学 | Phytohormone abscisic acid fluorescent molecular probe and application thereof |
| US20250051788A1 (en) * | 2021-05-24 | 2025-02-13 | Donald Danforth Plant Science Center | Compositions for rna-protein tethering and methods of using |
| JP2023040984A (en) * | 2021-09-10 | 2023-03-23 | 国立大学法人信州大学 | Methods and kits for editing rna |
| WO2023057777A1 (en) * | 2021-10-08 | 2023-04-13 | Pencil Biosciences Limited | Synthetic genome editing system |
| EP4636084A3 (en) * | 2021-10-08 | 2025-12-24 | Pencil Biosciences Limited | Synthetic genome editing system |
| WO2023150131A1 (en) * | 2022-02-01 | 2023-08-10 | The Regents Of The University Of California | Method of regulating alternative polyadenylation in rna |
| WO2025168955A1 (en) | 2024-02-09 | 2025-08-14 | Ucl Business Ltd | Therapy for dravet syndrome by targeting scn1a antisense transcript |
Also Published As
| Publication number | Publication date |
|---|---|
| AU2026200853A1 (en) | 2026-04-02 |
| IL284516A (en) | 2021-08-31 |
| US20220048962A1 (en) | 2022-02-17 |
| BR112021013173A2 (en) | 2021-09-28 |
| JP2022516919A (en) | 2022-03-03 |
| SG11202107209WA (en) | 2021-07-29 |
| KR20210124238A (en) | 2021-10-14 |
| EP3906042A4 (en) | 2023-03-15 |
| JP2025084877A (en) | 2025-06-03 |
| CN113543797A (en) | 2021-10-22 |
| AU2020204917A1 (en) | 2021-08-19 |
| EP3906042A1 (en) | 2021-11-10 |
| CA3125299A1 (en) | 2020-07-09 |
| MX2021008153A (en) | 2021-08-11 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| AU2020204917A1 (en) | Systems and methods for modulating RNA | |
| JP7535142B2 (en) | Use of programmable DNA-binding proteins to enhance targeted genome modification | |
| KR102523217B1 (en) | Use of nucleosome interacting protein domains to improve targeted genomic modification | |
| JP6914274B2 (en) | Crystal structure of CRISPRCPF1 | |
| AU2014361834B2 (en) | CRISPR-Cas systems and methods for altering expression of gene products, structural information and inducible modular Cas enzymes | |
| AU2012284365B2 (en) | Methods of transcription activator like effector assembly | |
| JP7275043B2 (en) | Enhanced hAT Family Transposon-Mediated Gene Transfer and Related Compositions, Systems and Methods | |
| JP7449646B2 (en) | Vector-free delivery of gene editing proteins and compositions to cells and tissues | |
| JP2024041866A (en) | Enhanced hAT family transposon-mediated gene transfer and related compositions, systems, and methods | |
| CA3026055A1 (en) | Novel crispr enzymes and systems | |
| CN107922949A (en) | Compounds and methods for for the genome editor based on CRISPR/CAS by homologous recombination | |
| JP2021519101A (en) | Modified nucleic acid editing system for ligating donor DNA | |
| US20250283062A1 (en) | Systems and methods for modulating rna | |
| HK40060450A (en) | Systems and methods for modulating rna | |
| Mistry et al. | Nucleic acid editing | |
| WO2026044291A1 (en) | Conversion of iscb and cas9 into rna-guided rna-editors | |
| EP4720304A2 (en) | Improved modular prime editing with modified effectors and templates | |
| HK40093157A (en) | Using programmable dna binding proteins to enhance targeted genome modification | |
| CN117916372A (en) | CAS12F1 without cleavage activity, fusion protein based on CAS12F1 without cleavage activity, CRISPR gene editing system including the same, and preparation method and use thereof | |
| CN121227666A (en) | A nuclease polypeptide, composition and its application |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 20736077 Country of ref document: EP Kind code of ref document: A1 |
|
| ENP | Entry into the national phase |
Ref document number: 3125299 Country of ref document: CA |
|
| ENP | Entry into the national phase |
Ref document number: 2021538952 Country of ref document: JP Kind code of ref document: A |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| REG | Reference to national code |
Ref country code: BR Ref legal event code: B01A Ref document number: 112021013173 Country of ref document: BR |
|
| ENP | Entry into the national phase |
Ref document number: 2020736077 Country of ref document: EP Effective date: 20210804 |
|
| ENP | Entry into the national phase |
Ref document number: 2020204917 Country of ref document: AU Date of ref document: 20200103 Kind code of ref document: A |
|
| ENP | Entry into the national phase |
Ref document number: 112021013173 Country of ref document: BR Kind code of ref document: A2 Effective date: 20210702 |