WO2014148557A1 - Egfr阻害剤感受性予測方法 - Google Patents
Egfr阻害剤感受性予測方法 Download PDFInfo
- Publication number
- WO2014148557A1 WO2014148557A1 PCT/JP2014/057556 JP2014057556W WO2014148557A1 WO 2014148557 A1 WO2014148557 A1 WO 2014148557A1 JP 2014057556 W JP2014057556 W JP 2014057556W WO 2014148557 A1 WO2014148557 A1 WO 2014148557A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- egfr inhibitor
- tumor
- cancer
- subject
- blood sample
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q1/00—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions
- C12Q1/68—Measuring or testing processes involving enzymes, nucleic acids or microorganisms; Compositions therefor; Processes of preparing such compositions involving nucleic acids
- C12Q1/6876—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes
- C12Q1/6883—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material
- C12Q1/6886—Nucleic acid products used in the analysis of nucleic acids, e.g. primers or probes for diseases caused by alterations of genetic material for cancer
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N33/00—Investigating or analysing materials by specific methods not covered by groups G01N1/00 - G01N31/00
- G01N33/48—Biological material, e.g. blood, urine; Haemocytometers
- G01N33/50—Chemical analysis of biological material, e.g. blood, urine; Testing involving biospecific ligand binding methods; Immunological testing
- G01N33/53—Immunoassay; Biospecific binding assay; Materials therefor
- G01N33/575—Immunoassay; Biospecific binding assay; Materials therefor for cancer
- G01N33/5758—Immunoassay; Biospecific binding assay; Materials therefor for cancer involving compounds serving as markers for tumours, cancers or neoplasias, e.g. cellular determinants, receptors, heat shock/stress proteins, A-protein, oligosaccharides or metabolites
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/106—Pharmacogenomics, i.e. genetic variability in individual responses to drugs and drug metabolism
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Q—MEASURING OR TESTING PROCESSES INVOLVING ENZYMES, NUCLEIC ACIDS OR MICROORGANISMS; COMPOSITIONS OR TEST PAPERS THEREFOR; PROCESSES OF PREPARING SUCH COMPOSITIONS; CONDITION-RESPONSIVE CONTROL IN MICROBIOLOGICAL OR ENZYMOLOGICAL PROCESSES
- C12Q2600/00—Oligonucleotides characterized by their use
- C12Q2600/156—Polymorphic or mutational markers
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N2333/00—Assays involving biological materials from specific organisms or of a specific nature
- G01N2333/90—Enzymes; Proenzymes
- G01N2333/914—Hydrolases (3)
-
- G—PHYSICS
- G01—MEASURING; TESTING
- G01N—INVESTIGATING OR ANALYSING MATERIALS BY DETERMINING THEIR CHEMICAL OR PHYSICAL PROPERTIES
- G01N2800/00—Detection or diagnosis of diseases
- G01N2800/52—Predicting or monitoring the response to treatment, e.g. for selection of therapy based on assay results in personalised medicine; Prognosis
Definitions
- the present invention relates to a method for predicting the sensitivity of a subject to an EGFR inhibitor by examining the genotype of a KRAS protein in a biological sample collected minimally invasively from the subject.
- Epidermal growth factor receptor is a very well-related receptor ErbB including EGFR (ErbB-1), Her2 / neu (ErbB-2), Her3 (ErbB-3) and Her4 (ErbB-4).
- ErbB-1 EGFR
- Her2 / neu ErbB-2
- Her3 ErbB-3
- Her4 ErbB-4
- the ErbB family is a type 1 tyrosine kinase family of growth factor receptors that play an important role in cell growth, differentiation, and survival. Activation of these receptors typically occurs through specific ligand binding, forming hetero- or homodimers between receptor family members. Subsequently, autophosphorylation of the tyrosine kinase domain occurs.
- EGFR activation in addition to inhibiting apoptosis, which is an important process for the continued growth and survival of cancer cells, activates receptor tyrosine kinases as well as cell proliferation, motility, attachment, invasion and chemotherapy Guides a series of downstream signaling events that mediate resistance to.
- members of this family including EGFR and HER2, are directly involved in cell transformation.
- molecular target drugs targeting EGFR have been developed as anticancer agents.
- clinical trials of two main types of EGFR inhibitors have been conducted: anti-EGFR antibodies and small molecule EGFR tyrosine kinase inhibitors (TKIs).
- Anti-EGFR antibodies such as cetuximab were designed to bind to the extracellular domain of EGFR and block the activation of signaling downstream of EGFR.
- Cetuximab also known as antibody 225, see US Pat. No. 6,057,059
- small molecule TKIs such as gefitinib (compound ZD1839, Iressa) and erlotinib (compound OSI-774, Tarceva) compete for ATP to bind to the intracellular catalytic domain of EGFR tyrosine kinase. As a result, it inhibits EGFR autophosphorylation and downstream signaling.
- cetuximab and panitumumab which are molecular target drugs for colorectal cancer
- the relationship with the mutated KRAS gene or its protein the therapeutic effect of gefitinib, erlotinib, which are molecular target drugs for lung cancer, and the mutated EGFR gene
- the relationship with the protein, and the therapeutic response of crizotinib, an ALK inhibitor, and the relationship with the fusion gene with translocation or the protein have been clinically clarified.
- companion diagnostics have been noted that involve testing for KRAS mutations.
- KRAS wild type patients had a response rate of 59.3% when combined with cetuximab
- KRAS mutant patients combined with cetuximab
- the time response rate was 36.2%
- the cetuximab response rate for KRAS mutant patients was dramatically low (see, for example, Non-Patent Document 1).
- cetuximab response efficiency was extremely low in KRAS mutant patients, and it was proved that cetuximab was effective in wild-type patients.
- Non-Patent Document 2 the mutated KRAS gene in colorectal cancer patients was a predictor of therapeutic effect of EGFR inhibitors.
- Non-Patent Document 2 results of clinical trials in which mutations exist in BRAF and PIK3CA have also been reported.
- the first problem is that in cases where surgical resection is not performed, biopsy specimens are used for KRAS genetic diagnosis, but their reliability has not been confirmed.
- taking a biopsy is invasive to the patient, and after resection of the primary lesion, recurrent tumors cannot often be resected. Therefore, a highly reliable diagnostic marker that can replace the method of directly examining the status of KRAS (KRAS genotype) in tumor tissue is desired.
- the second problem is that the mutation status does not always match between the metastatic lesion and the primary lesion.
- Watanabe et al. Reported that 15 out of 43 colorectal cancer patients were mutant patients, and the coincidence rate between the primary lesion and the metastatic lesion was 88.4% (see Non-Patent Document 3). That is, about 10% of the primary lesions and metastatic lesions do not have the same KRAS mutation status.
- Gatttenlohner et al As a result of examining the status before and after the treatment of 21 cases of metastatic colorectal cancer, there was no change in 20 cases (95.2%).
- One example in which there was a change was heterogeneity and was a frequent occurrence (see Non-Patent Document 4).
- the third problem is the possibility that mutant clones will increase during EGFR inhibitor treatment, that is, acquisition of resistance to EGFR inhibitors.
- Diaz Jr et al Detected the KRAS gene in which plasma DNA was mutated after EGFR inhibitor treatment was started in 24 cases of wild-type colorectal cancer. As a result, mutated KRAS gene was detected in 9 cases (37.5%), and mutations were observed in all cases by at least 26 weeks after administration, and at the same time as or after observation of mutations in extracellular DNA. Reported that they had acquired resistance.
- Non-Patent Document 5 Misale et al. Also predicts sensitivity to EGFR inhibitors by detecting KRAS gene mutations in plasma with high sensitivity, and if a KRAS gene mutation in plasma is detected, EGFR It has been proposed to switch to other drugs instead of inhibitors (see Non-Patent Document 6).
- Non-Patent Documents 5 and 6 describe that the status of KRAS in blood is examined by utilizing the fact that KRAS present in a metastasis of a subject is also detected in circulating DNA. Moreover, it is described that the sensitivity of the metastasis to the EGFR inhibitor can be examined and the response when the EGFR inhibitor is administered to the subject can be predicted.
- the present invention relates to a method for predicting the sensitivity of an EGFR-mediated tumor (cancer) to an EGFR inhibitor using the KRAS status in the peripheral blood of the patient as an index regardless of the KRAS status in the tumor tissue of the patient.
- cancer EGFR-mediated tumor
- the present inventors have conducted intensive research to solve the above problems. As a result, in patients with EGFR-mediated tumors (cancers), there are cases where the KRAS status in the tumor tissue does not match the KRAS status in the peripheral blood, and the KRAS mutant type is not detected in the tumor tissue.
- the present inventors completed the present invention by discovering that EGFR inhibitors are less effective in patients in which mutant KRAS is detected in peripheral blood.
- the EGFR inhibitor sensitivity prediction method includes (a) whether or not a KRAS gene-derived nucleic acid or a protein thereof is present in a blood sample collected from a subject, and said blood sample in the blood sample.
- the mutant KRAS gene-derived nucleic acid or protein thereof is not detected, it is determined that the subject's tumor is likely to be sensitive to EGFR inhibitor, and the mutant KRAS is contained in the blood sample.
- the subject's tumor may not be sensitive to EGFR inhibitors
- the KRAS gene-derived nucleic acid or protein thereof detected from a tissue sample or cell sample collected from a tumor of the subject is the KRAS gene-derived nucleic acid or protein genotype detected from the blood sample.
- the subject's it may be determined that the tumor is likely not sensitive to EGFR inhibitors.
- the subject may have undergone surgical resection of a tumor portion in the past.
- the subject may have been administered an EGFR inhibitor in the past.
- the subject may have shown drug resistance to the EGFR inhibitor in the past.
- the blood sample may be collected from a subject who has passed 60 days after the administration of the EGFR inhibitor.
- the subject is a tumor patient who has received another EGFR inhibitor treatment after receiving the EGFR inhibitor medication, and the blood sample is the tumor It may be collected before the patient is about to receive the EGFR inhibitor medication again.
- the other anti-tumor therapy may be a chemotherapeutic drug treatment.
- the chemotherapeutic agent is fluorouracil, folinic acid, oxaliplatin, irinotecan, cytarabine, fludarabine, gemcitabine, hydroxyurea, methotrexate, bleomycin, chlorambucil, cisplatin, cyclophosphamide, doxorubicin, mitoxantrone, It may be one or more selected from the group consisting of camptothecin, topotecan, teniposide, colcemid, colchicine, paclitaxel, vinblastine, vincristine, and tamoxifen.
- the other anti-tumor therapy may be radiation therapy.
- the other anti-tumor therapy may be a medication treatment of a different type of molecular target drug than the EGFR inhibitor already administered to the subject.
- the molecular target drug is selected from the group consisting of 2 or a species selected from the group consisting of cetuximab, panitumumab, bevacizumab, gefitinib, erlotinib, regorafenib, crizotinib, sunitinib, sorafenib, everolimus, trastuzumab, lapatinib, and rituximab It may be the above.
- the other anti-tumor therapy may be a combination therapy of the molecular target drug and the chemotherapeutic drug.
- the tumor may be a recurrent tumor.
- the tumor may be a metastatic focus.
- the tumor may be a primary lesion.
- the tumor comprises colon cancer, colon cancer, rectal cancer, lung cancer, liver cancer, breast cancer, ovarian cancer, prostate cancer, renal cancer, esophageal cancer, head and neck cancer, uterine cancer, and cervical cancer. 1 type or 2 types or more selected from a group may be sufficient.
- the tumor may be present at a plurality of locations in the body of the subject.
- the mutant forms are GRAS proteins G12A, G12C, G12D, G12R, G12S, G12V, G13D, G12S2, G13A, G13S, G13V, G13R, G13C, Q61H, Q61L, Q61R, A146T, and A146V. 1 or 2 or more selected from the group consisting of may be sufficient.
- the presence or absence of KRAS in the blood sample and the determination of whether it is a wild type or a mutant type are determined based on whether or not a wild type KRAS gene-derived nucleic acid is detected from circulating DNA in the blood sample. And whether or not a mutant KRAS gene-derived nucleic acid is detected.
- the blood sample may be peripheral blood, serum, or plasma.
- CEA in the blood sample may be 5 ng / mL or less, or the value of CA19-9 in the blood sample may be 37.0 U / mL or less.
- the sensitivity of the subject to the EGFR inhibitor can be accurately predicted from a biological sample that is less invasive to the subject.
- the EGFR inhibitor sensitivity prediction method according to an embodiment of the present invention may be used to determine the KRAS status in blood regardless of the KRAS status in the tumor tissue. Using the status as an index, the sensitivity of the EGFR inhibitor is predicted. A subject whose blood KRAS is wild type and no mutant is detected is predicted to be highly sensitive to EGFR inhibitors regardless of whether KRAS in the tumor tissue is wild type or mutant. Conversely, subjects in whom mutant KRAS has been detected in blood are expected to be less sensitive to EGFR inhibitors regardless of whether KRAS in the tumor tissue is wild-type or mutant.
- a mutated KRAS gene or a protein thereof protein derived from a mutated KRAS gene
- the EGFR inhibitor does not work.
- mRNA of KRAS protein or KRAS gene was detected not only in tumor tissue but also in blood.
- the status of KRAS in this blood is as follows: It was thought to be consistent with the KRAS status in the tumor tissue.
- the status of KRAS in blood may not match the status of KRAS in tumor tissue.
- the efficacy of the EGFR inhibitor in the tumor tissue is based on the status of KRAS in the blood rather than the status of KRAS in the tumor tissue.
- Example 1 in the analysis of clinical results for patients with recurrent colorectal cancer, the KRAS gene mutated in both primary and metastatic lesions was not observed, but circulating in the blood In patients with recurrent colorectal cancer in which a mutated KRAS gene-derived nucleic acid was detected from DNA, no tumor reduction effect was observed with cetuximab, an EGFR inhibitor. In addition, in patients with recurrent colorectal cancer in which mutant KRAS was detected in blood despite the absence of mutant KRAS in the tumor tissue in either the primary lesion or metastatic lesion, The tumor did not respond.
- the subject in which the mutated KRAS gene is detected from circulating DNA in peripheral blood has a low sensitivity to the EGFR inhibitor, and the EGFR inhibitor It can be predicted that the efficacy of the is low.
- the tumor is EGFR It is a tumor that does not respond to an inhibitor, and it can be predicted that re-administration of an EGFR inhibitor will not work.
- the findings of the present inventors for the first time indicate that the sensitivity to EGFR inhibitors can be accurately predicted by using the status of KRAS in blood instead of the status of KRAS in tumor tissue as an index. is there.
- the sensitivity prediction method is a method for predicting the EGFR inhibitor sensitivity of a subject, and has the following steps (a) and (b).
- the step (a) when a wild-type KRAS gene-derived nucleic acid or a protein thereof is detected in the blood sample and a mutant KRAS gene-derived nucleic acid or a protein thereof is not detected, the subject Is determined to be highly sensitive to EGFR inhibitor, and when a mutant KRAS gene-derived nucleic acid or protein thereof is detected in the blood sample, the subject tumor is sensitive to EGFR inhibitor.
- the mutant KRAS detected in the sensitivity prediction method according to the present invention includes all forms of mutation such as insertion, inversion, deletion, and / or point mutation. These mutated KRAS genes can be found in somatic or germline, unlike wild-type KRAS found in one allele (heterozygous) or both alleles (homozygous), respectively. Mutant KRAS. Somatic mutations occur only in certain tissues, such as tumor tissues, and are not inherited in the germline. Germline mutations can be found in any body tissue.
- the mutant KRAS detected in the method for predicting sensitivity according to the present invention is mutated with one or more amino acid substitutions in codons 12, 13, 61, and 146 on exons 2 to 4 of the KRAS gene.
- KRAS is preferred.
- KRAS proteins G12A, G12C, G12D, G12R, G12S, G12V, G13D, G13A, G12S2, G13S, G13V, G13R, G13C, Q61H, Q61L, Q61R, A146T, and A146V
- Examples include mutant KRAS having one or more selected mutations. Table 1 shows embodiments of nucleic acid (gene) mutation of each mutant type.
- amino acid sequence of KRAS is SEQ ID NO: 1
- the gene sequence containing codon 12 on exon 2 of KRAS is shown in SEQ ID NO: 2
- the gene sequence containing codon 61 on exon 3 of KRAS is shown in SEQ ID NO: 3
- KRAS The gene sequences containing codon 146 on exon 4 are shown in SEQ ID NO: 4, respectively.
- the sensitivity prediction method according to the present invention in the step (a), not the KRAS status in the tumor tissue but the KRAS status in the blood sample is examined.
- the blood sample may be peripheral blood itself, or serum or plasma. Since blood samples can be collected from subjects less invasively than tumor tissues, subjects with difficulty in collecting biological samples of tumor tissues, such as patients with recurrent tumors, are expected to be sensitive to EGFR inhibitors. be able to. Moreover, since a blood sample can also be collected from a subject over time, the sensitivity prediction method according to the present invention is also suitable for monitoring whether a tumor has recurred.
- the blood sample is preferably a blood sample collected from a subject at an early stage regardless of whether it is a primary tumor, a metastatic tumor, or a recurrent tumor.
- the blood sample used in the sensitivity prediction method according to the present invention preferably has a CEA of 5 ng / mL or less, or a CA19-9 value of 37.0 U / mL or less.
- the EGFR inhibitor whose sensitivity is predicted is not particularly limited as long as it is a substance having an EGFR inhibitory action in animals including humans, and may be an anti-EGFR antibody, It may be TKI.
- the anti-EGFR antibody include cetuximab (product name: Erbitutux (registered trademark), Imclone Systems Inc.) and panitumumab (product name: ABX-EGF, Abgenix, Inc.).
- TKI small molecules that compete with ATP, such as erlotinib (product name: Tarceva (registered trademark), OSI Pharmaceuticals), gefitinib (product name: Iressa (registered trademark), Astra-Zeneca) , Dvir et al., Journal of Cell Biology, vol. 113, pp. 857-865 (1991); triphospyrimidine compounds disclosed in US Pat. No.
- the tumor that is the target for predicting the sensitivity of the EGFR inhibitor is not particularly limited as long as it is an EGFR-mediated tumor (cancer), that is, a tumor that plays a role of EGFR in tumor formation.
- the tumor includes brain, liver, kidney, bladder, breast, stomach, ovary, colorectal, prostate, pancreas, breast, lung, vulva, thyroid, colorectal, esophagus, liver cancer, sarcoma, glioblastoma , Head and neck, leukemia and lymphoid malignancies.
- neuroblastoma intestinal cancer (eg rectal cancer, colon cancer, familial colorectal polyposis cancer and hereditary non-polyposis colorectal cancer), esophageal cancer, lip cancer, laryngeal cancer, hypopharyngeal cancer, tongue cancer , Salivary gland cancer, stomach cancer, adenocarcinoma, medullary thyroid cancer, papillary thyroid cancer, kidney cancer, renal parenchymal cancer, ovarian cancer, cervical cancer, endometrial cancer, endometrial cancer, choriocarcinoma, pancreatic cancer, prostate cancer, Testicular cancer, breast cancer, ureteral cancer, melanoma, brain tumor (eg glioblastoma, astrocytoma, meningioma, medulloblastoma and peripheral neuroectodermal tumor), Hodgkin lymphoma, non-Hodgkin lymphoma, bar Kit lymphoma, acute lymphoblastic leukemia
- tumors to be predicted in the sensitivity prediction method according to the present invention include colon cancer, colon cancer, rectal cancer, lung cancer, liver cancer, breast cancer, ovarian cancer, prostate cancer, kidney cancer, esophageal cancer, head and neck cancer, uterine cancer, And one or more selected from the group consisting of cervical cancer.
- the tumor that is a target for predicting the sensitivity of the EGFR inhibitor may be a primary lesion (primary tumor) or a metastatic lesion (metastatic tumor). It may also be a recurrent tumor. Furthermore, tumors may be present at a plurality of locations in the subject's body.
- a recurrent tumor it is preferable to predict the sensitivity of an EGFR inhibitor for a recurrent tumor. For example, by using a blood sample collected from a subject who has undergone surgical excision of a tumor part in the past or a subject who has been administered an EGFR inhibitor in the past, recurrent tumors or metastases can be obtained. EGFR inhibitor sensitivity can be predicted. In addition, when a subject has been administered an EGFR inhibitor in the past, it is preferable to use a blood sample collected after 60 days have passed since the administration of the EGFR inhibitor.
- the sensitivity prediction method according to the present invention is carried out every 60 days because of the balance between blood sampling for tumor marker testing about once a month and CT testing about once every three months. This is because it is considered preferable.
- the blood sample used in the sensitivity prediction method according to the present invention may be collected from a subject who has shown drug resistance to an EGFR inhibitor in the past. Even if a subject has previously shown drug resistance to an EGFR inhibitor, if the mutant KRAS gene-derived nucleic acid or protein is not detected from the blood sample, the subject will collect the blood sample. At that time, it can be determined that there is sensitivity to the EGFR inhibitor. Therefore, it can be determined that taking an EGFR inhibitor has a high possibility of obtaining a tumor reduction effect.
- the subject when a mutant KRAS gene-derived nucleic acid or a protein thereof is detected from a blood sample, the subject is less sensitive to the EGFR inhibitor at the time the blood sample is collected, and is taking the EGFR inhibitor. However, it can be determined that there is a high possibility that the tumor reduction effect is not obtained.
- the sensitivity prediction method according to the present invention is also preferably performed on a blood sample collected from a subject who re-challenges an EGFR inhibitor.
- the “subject who re-challenges the EGFR inhibitor” refers to another anti-tumor therapy different from the EGFR inhibitor dosage treatment after the EGFR inhibitor dosage treatment, It means a subject who intends to receive EGFR inhibitor medication.
- tolerance is obtained by administration of an EGFR inhibitor, but by examining the status of KRAS in blood before re-challenge, the response of the EGFR inhibitor in re-challenge can be predicted.
- anti-tumor therapies received before re-challenge with the EGFR inhibitor can be appropriately selected from known anti-tumor therapies according to the condition of the subject.
- Specific examples of such other anti-tumor therapies include radiation therapy, chemotherapeutic drug dosing treatment, and dosing treatment of a molecular target drug of a type different from the already administered EGFR inhibitor.
- the other anti-tumor therapy may be a combination therapy of one or two or more anti-tumor therapies.
- a combination therapy of a molecular target drug and a chemotherapeutic drug is preferable.
- the chemotherapeutic agent is not limited and may be a compound having cytotoxicity or cell division inhibitory property.
- an antimetabolite such as fluorouracil, cytarabine, fludarabine, 5-fluoro-2′-deoxyuridine, gemcitabine, hydroxyurea, or methotrexate
- a DNA fragmenting agent such as bleomycin
- DNA cross-linking agents such as chlorambucil, cisplatin, cyclophosphamide, or nitrogen mustard
- intercalating agents such as adriamycin (doxorubicin), or mitoxantrone
- protein synthesis inhibitors such as , L-asparaginase, cycloheximide, puromycin, or diphtheria toxin
- topoisomerase I toxin such as camptothecin, or topotecan
- folinic acid oxaliplatin
- irinotecan daunarubicin
- taxotere adenomycin C
- mitomycin C adenomycin C
- chemotherapeutic agents include fluorouracil, folinic acid, oxaliplatin, irinotecan, cytarabine, fludarabine, gemcitabine, hydroxyurea, methotrexate, bleomycin, chlorambucil, cisplatin, cyclophosphamide, doxorubicin, mitoxantrone, camptothecin, topotecan, tenposide 1 or 2 or more selected from the group consisting of colcemid, colchicine, paclitaxel, vinblastine, vincristine, and tamoxifen.
- the molecular target agent is selected from the group consisting of cetuximab, panitumumab, bevacizumab, gefitinib, erlotinib, regorafenib, crizotinib, sunitinib, sorafenib, everolimus, trastuzumab, lapatinib, and one selected from the group consisting of rituximab The above is mentioned.
- the KRAS status in the blood sample may be determined at the protein level or at the nucleic acid level (genomic DNA or mRNA).
- a KRAS gene-derived nucleic acid as a measurement target because it can be detected with high sensitivity.
- the presence or absence of KRAS in the blood sample and the determination of whether it is a wild type or a mutant type are detected from the circulating DNA in the blood sample, and nucleic acids derived from wild type and mutant KRAS genes are detected. It is preferable to check by checking whether or not.
- the KRAS gene-derived nucleic acid is a full-length or part of the KRAS gene genomic DNA, a full-length or part of the mRNA of the KRAS gene, a cDNA obtained using the full-length or part of the mRNA as a template, or a polymerase chain reaction (PCR) ) Etc., and amplification products artificially amplified.
- PCR polymerase chain reaction
- the detection of the KRAS gene-derived nucleic acid in the blood sample and the determination of the genotype of the detected KRAS gene-derived nucleic acid can be performed by conventional methods.
- the presence and status of KRAS in a blood sample can be determined by detecting the KRAS gene-derived nucleic acid contained in the blood sample using digital PCR.
- Highly sensitive detection especially by using Biorad's droplet digital PCR (ddPCR) technology (Hindson, et.al., Analytical Chemistry, 2011, vol. 83 (22), pp. 8604-8610) can do.
- the greater the number of droplets the higher the analysis accuracy.
- 10,000 droplets are required to detect one mutation. Therefore, it is preferable to define the surfactant concentration in the PCR master mix.
- ethylene glycol or glycerol used as a storage solution for DNA extender or the like has a final concentration of 0.15% or less, or Triton-X has a final concentration of 0.0003% or less.
- the surfactant is at the final concentration or more, the number of emulsions due to droplets is drastically reduced, making it difficult to detect mutations with high sensitivity.
- the absolute amount of the allelic copy number of mutant KRAS that would be contained in the nucleic acid was increased by performing the first PCR for 15 to 50 cycles using the nucleic acid and nucleic acid fragment obtained from the blood sample. It is also preferable to carry out a known mutation detection method after dilution to about 106 copies. According to this method, the total number of mutants present in the reaction system is increased, so that even if the number of mutations increases, the possibility that the mutant allele does not physically exist and becomes undetectable is reduced. be able to. Furthermore, the method may be combined with the digital PCR described above.
- a specific KRAS specific product is amplified with respect to the amplified product.
- the probe is detectably labeled with, for example, a radioisotope ( 3 H, 32 P, 33 P, etc.), a fluorescent agent (rhodamine, fluorescene, etc.) or a color former.
- the probe may also be an antisense oligomer such as PNA, morpholino-phosphoramidates, or LNA.
- the base length of the probe may be about 8 to about 100 nucleotides, or about 10 to about 75, or about 15 to about 50, or about 20 to about 30.
- the presence and status of KRAS in a blood sample can be analyzed using the invader method (Michael Olivier, Mutation Research 573: 103-110, 2005).
- the invader method an allele probe and an invader oligo are hybridized so as to partially form a triple base with respect to double-stranded DNA or mRNA prepared by PCR or the like.
- a part of the 5 ′ end of the allele probe is designed to have a sequence (flap portion) that is non-complementary to the double-stranded DNA or mRNA.
- Invader oligos on the other hand, have sequences that are completely complementary to them.
- the two allele probes are designed to be complementary to the wild type and the mutant type, respectively, and hybridize competitively with the double-stranded DNA or mRNA as described above, thereby completely hybridizing.
- a part of the flap endonuclease that is a triple base is recognized, and the flap part of the allele probe hybridizes to a self-complementary FRET cassette existing in the reaction system.
- the target mutation is cleaved by the flap endonuclease, the fluorescently modified DNA fragment in the FRET cassette is released, and dissociates from the quencher in FRET.
- the flap part of the allele probe once cleaved can be re-hybridized to another FRET cassette, and the mutation can be detected with a very high sensitivity that the signal is amplified.
- a ligase chain reaction known in the art can be used to amplify a fragment containing a region encoding a mutation site in the KRAS gene (eg, Wu, et.al., Genomics, 1989, vol. .4, pp. 560-569).
- a technique known as allele-specific PCR can also be used (see, for example, Ruano and Kidd, Nucleic Acids Research, 1989, vol. 17, pp. 8392). According to the technique, a primer is used that hybridizes at its 3 'end to a particular KRAS mutation. If a particular KRAS mutation is not present, no amplification product is observed.
- Amplification Reference Mutation System (ARMS) (see, for example, European Patent Application Publication No. 0332435 and Newton et.al., Nucleic Acids Research, 1989, vol. 17, pp. 7) can also be used.
- a sequence analysis method based on the Sanger method is used to directly determine the base sequence of the KRAS gene genomic DNA or mRNA in the blood sample, or the amplification product thereof. A method is mentioned.
- the base sequence can also be determined via PCR.
- restriction fragment length polymorphism (RFLP) probes for the gene or surrounding marker genes can be used to score allelic alterations or insertions in the polymorphic fragment.
- SSCP Single-stranded DNA conformation polymorphism
- the sensitivity prediction method according to the present invention can be performed more easily by kitting reagents used for the detection of the KRAS gene-derived nucleic acid in the blood sample and the determination of the genotype of the detected KRAS gene-derived nucleic acid.
- the reagent include a reagent for extracting a nucleic acid from a blood sample, an enzyme such as polymerase or ligase, a probe or primer that can specifically hybridize to a specific KRAS genotype (a mutation site of the KRAS gene). Or an oligonucleotide that specifically hybridizes to the adjacent site).
- the kit also includes a protocol for detecting a KRAS gene-derived nucleic acid in a blood sample and a method for determining the genotype thereof, and a criterion for determining EGFR inhibitor sensitivity based on the result of the obtained KRAS genotype (status). It may contain a document or the like that includes instructions for.
- the sensitivity prediction method according to the present invention can provide important information when determining whether or not an EGFR inhibitor administration treatment is applied. That is, the sensitivity prediction method according to the present invention can provide useful information to the clinician, and the clinician can determine an appropriate treatment method based on the information obtained from the method.
- Example 1 Mutations with non-synonymous amino acid substitutions in 23 patients with recurrent colorectal cancer with surgical resection of primary lesions, and KRAS contained in serum prepared from primary lesions, metastatic lesions, and blood collected after confirmation of metastatic lesions I investigated.
- “None” means patients who have not yet been treated and may be administered in the future
- “Before treatment” means that administration is scheduled
- “Ongoing” means chemotherapy. It means that the patient is responding and continues to be administered.
- “After administration” means that chemotherapy has been effective, but at that time point, administration has been completed. It means a state in which is no longer effective (ie, a state in which palliative care is entered).
- Anti-EGFR antibody drug treatment was performed on patients with ID numbers 8 and 15 whose primary lesions were KRAS wild type. In both cases, the primary lesion was excised after anti-EGFR antibody treatment before the primary lesion was excised.
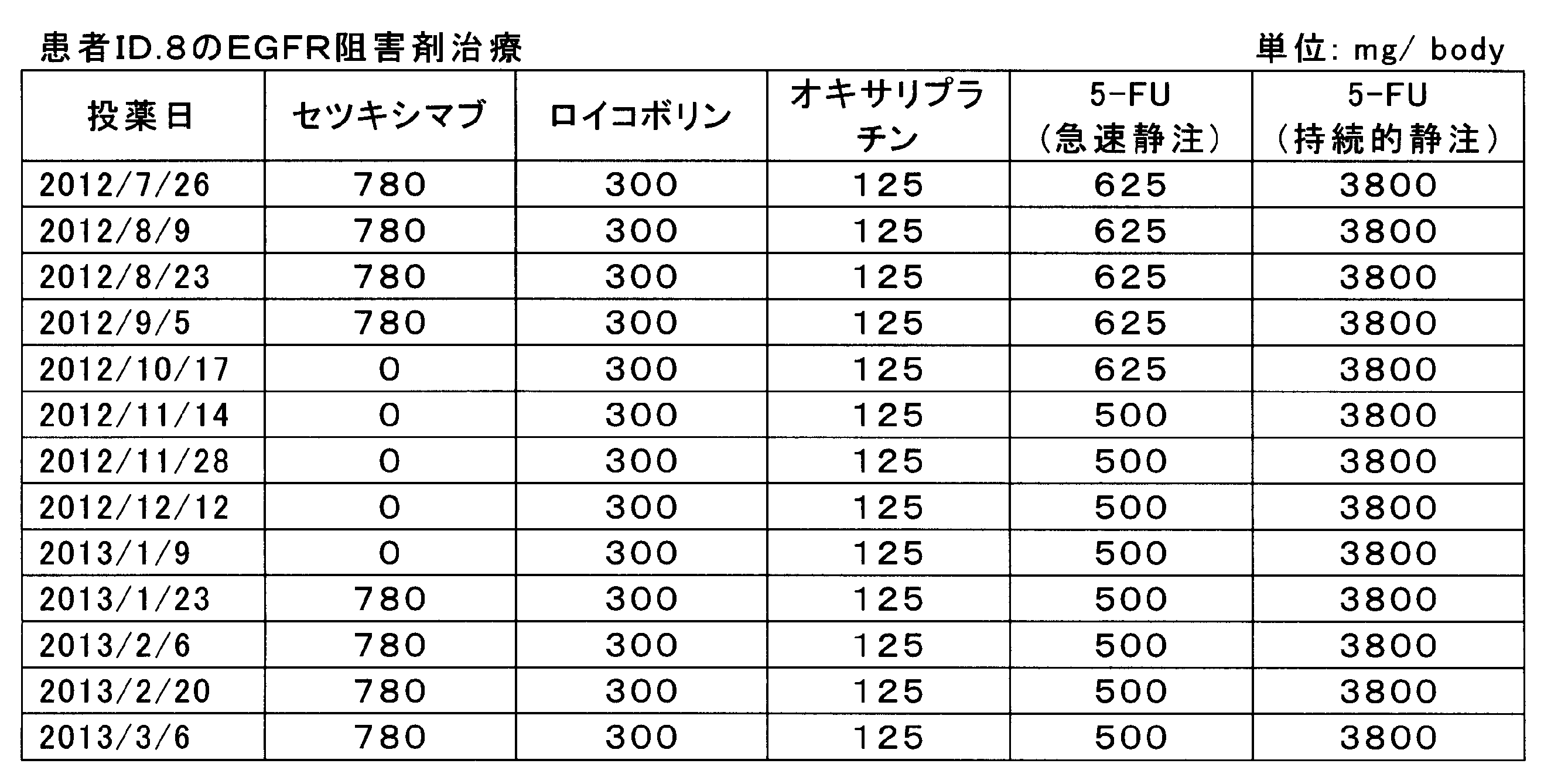
- the patient with ID number 8 was treated with cetuximab and FOLOX chemotherapy (fluorouracil, folinic acid, oxaliplatin). Cetuximab was administered by a 2-week interval administration method, and 780 mg / body was administered intravenously over 1 hour.
- FOLFOX chemotherapy was administered by intravenous infusion of 300 mg / body folinic acid (leucovorin) and 125 mg / body oxaliplatin over 2 hours, and fluorouracil (5-FU) was rapidly infused intravenously at 625 or 500 mg / body. After administration, 3800 mg / body was further intravenously administered for 22 hours. The administration history is shown in Table 3.
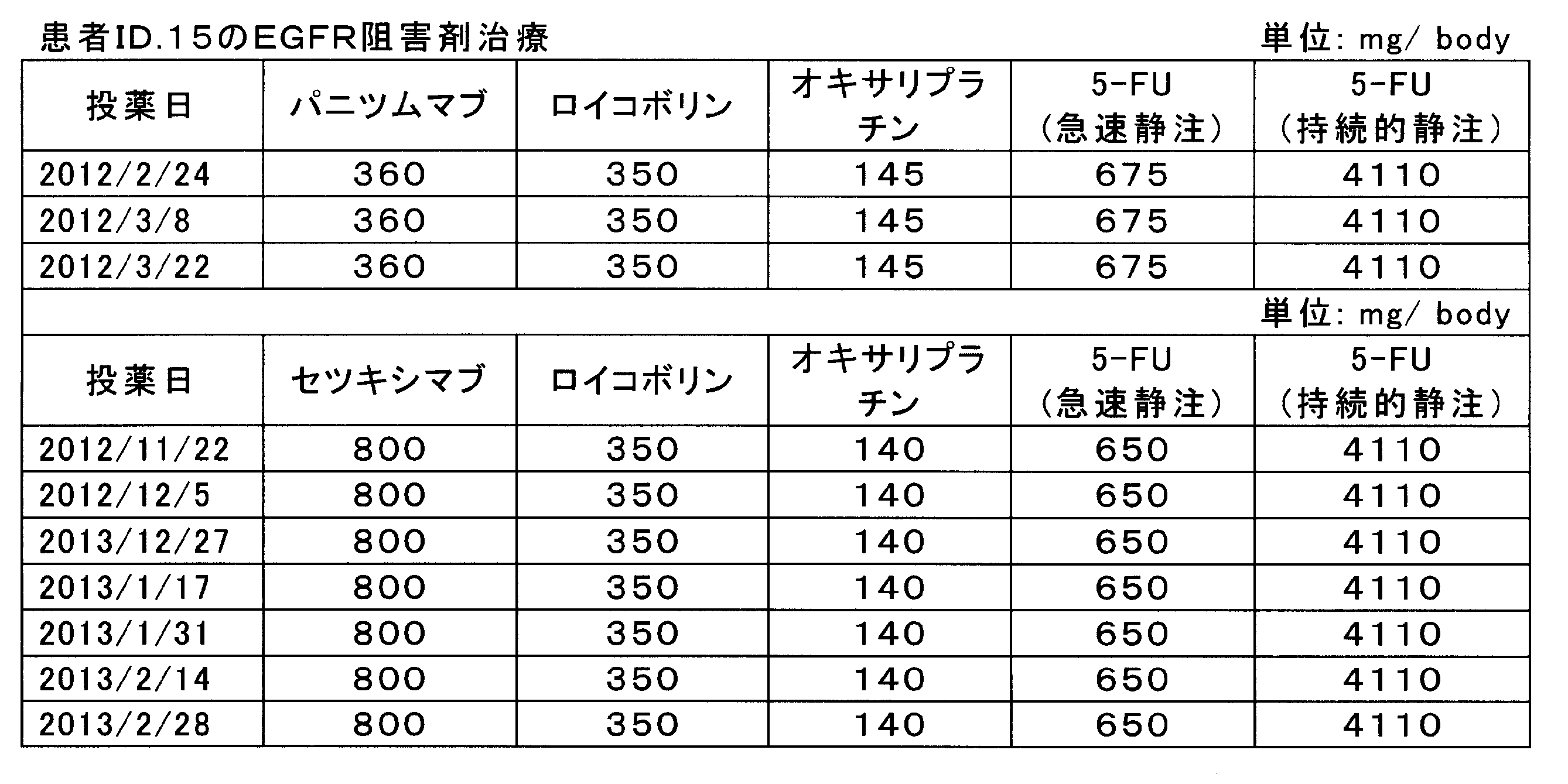
- panitumumab or cetuximab and FOLOX chemotherapy fluorouracil, folinic acid, oxaliplatin.
- Panitumumab or cetuximab was administered by a 2-week interval administration method, panitumumab was administered intravenously at 360 mg / body, and cetuximab was administered intravenously at 800 mg / body over 1 hour.
- FOLFOX chemotherapy is 350 mg / body folinic acid (leucovorin) and 145 or 140 mg / body oxaliplatin administered intravenously over 2 hours, and fluorouracil (5-FU) is administered at 675 or 650 mg / body.
- fluorouracil 5-FU
- 4110 mg / body was further intravenously administered for 22 hours.
- the administration history is shown in Table 4.
- CEA and CA-19-9 in serum were measured by the CLEIA method (chemiluminescence enzyme immunoassay). Table 5 shows the measurement results.
- Isolation and purification of cell-free (cf) DNA from serum Isolation and purification of cfDNA from serum was performed using QIAamp Circulating Nucleic Acid Kit (Qiagen). The amount of serum sample provided to this kit varied from patient to patient and was between 2 mL and 4 mL. The DNA isolation / purification process followed the instructions attached to the kit. Final elution from the spin column was performed using 50 ⁇ L of TE buffer.
- DNA isolation and purification from FFPE sections were performed using QIAamp DNA FFPE Tissue Kit (Qiagen). For each sample, three FFPE slices sliced to 10 ⁇ m were used. The DNA isolation / purification process followed the instructions attached to the kit. Final elution from the spin column was performed using 100 ⁇ L of TE buffer.
- Quantification of DNA Quantification of DNA isolated and purified from cfDNA and FFPE sections was performed using Quant-iT (registered trademark) PicoGreen (registered trademark) dsDNA Reagent and Kits (Invitrogen). All samples to be measured were obtained by diluting the isolated DNA 20 times with TE buffer. SAFIRA (TECAN) was used as the fluorescence measuring apparatus.
- KRAS forward: 5′-GAATGGTCCTGCACCAGTAA-3 ′ (SEQ ID NO: 5) and KRAS (reverse): 5′-GTGTGACATGTTCTAATATAGTCA-3 ′ (SEQ ID NO: 6) were used as primer sequences for direct sequencing of KRAS.
- the length of each PCR product is 214 bp, and the PCR conditions are 40 cycles of 94 ° C. for 20 seconds, 60 ° C. for 20 seconds, and 72 ° C. for 30 seconds after predenaturation at 95 ° C.
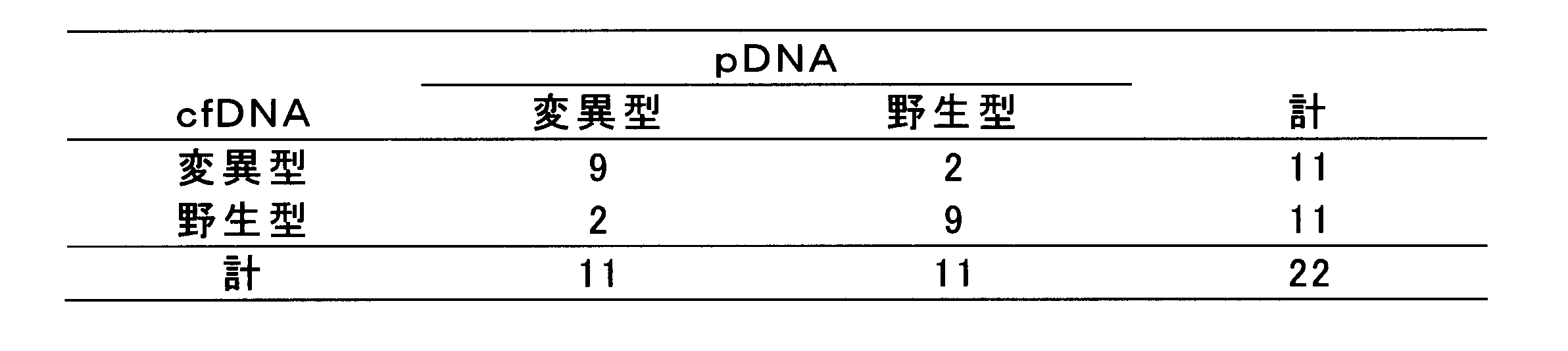
- the concordance rate between the KRAS gene mutation (codons 12 and 13) in the surgical specimen of the primary lesion and the KRAS gene mutation in cfDNA was 81.8%.
- Table 6 shows a correlation table between KRAS gene mutations in the primary lesion and cfDNA.
- pDNA means DNA extracted from the primary lesion.
- G13D KRAS protein in which the 13th glycine is replaced by aspartic acid
- G13D KRAS protein in which the 13th glycine is replaced by aspartic acid
- the mutated KRAS gene was also observed in the serum collected at the time of recurrence diagnosis in all patients in which the KRAS gene mutated in the primary lesion was observed. It was. From the results, it is useful to identify a mutated KRAS gene in the circulation in the diagnosis of recurrence after resection of the primary lesion. For subjects who have received treatment for the primary lesion, By examining the KRAS status, the presence of a recurrent tumor can be confirmed early (in some patients, earlier than existing biomarkers such as CEA and CA19-9).
- the EGFR inhibitor sensitivity prediction method according to the present invention uses a peripheral blood sample. For this reason, the sensitivity to the EGFR inhibitor and the response of the EGFR inhibitor can be predicted with high accuracy in a minimally invasive manner without excision of the primary lesion or collection of a biopsy such as biopsy. From the viewpoint of this low invasiveness and good accuracy, it is considered that the EGFR inhibitor sensitivity prediction method according to the present invention is widely used as a test method that changes to fecal occult blood or the like in cancer screening or the like.
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Immunology (AREA)
- Engineering & Computer Science (AREA)
- Organic Chemistry (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Analytical Chemistry (AREA)
- Pathology (AREA)
- Molecular Biology (AREA)
- Genetics & Genomics (AREA)
- Wood Science & Technology (AREA)
- Zoology (AREA)
- Biochemistry (AREA)
- Biotechnology (AREA)
- Physics & Mathematics (AREA)
- Microbiology (AREA)
- General Health & Medical Sciences (AREA)
- Hematology (AREA)
- Urology & Nephrology (AREA)
- Biomedical Technology (AREA)
- Oncology (AREA)
- Biophysics (AREA)
- Hospice & Palliative Care (AREA)
- Bioinformatics & Cheminformatics (AREA)
- General Engineering & Computer Science (AREA)
- Food Science & Technology (AREA)
- General Physics & Mathematics (AREA)
- Cell Biology (AREA)
- Medicinal Chemistry (AREA)
- Measuring Or Testing Involving Enzymes Or Micro-Organisms (AREA)
- Investigating Or Analysing Biological Materials (AREA)
Abstract
Description
本願は、2013年3月19日に日本に出願された特願2013-057033号に基づき優先権を主張し、その内容をここに援用する。
また、EGFR及びHER2を含むこのファミリーのメンバーは細胞形質転換に直接関係している。このため、抗癌剤として、EGFRを標的とする分子標的薬が開発されている。現在までに、抗EGFR抗体及び低分子EGFRチロシンキナーゼ阻害剤(TKI)の2つの主なタイプのEGFR阻害剤の臨床試験が行われた。セツキシマブなどの抗EGFR抗体は、EGFRの細胞外ドメインに結合し、EGFR下流のシグナリングの活性化をブロックするようにデザインされた。セツキシマブ(抗体225としてもまた公知である、特許文献1参照。)は、高レベルの野生型EGFRを発現するA431細胞に対して作製された。これとは対照的に、ゲフィチニブ(化合物ZD1839、イレッサ)やエルロチニブ(化合物OSI-774、タルセバ)などの低分子TKIは、EGFRチロシンキナーゼの細胞内触媒ドメインへの結合するためのATPを競合する。その結果、EGFR自己リン酸化及び下流のシグナリングを阻害する。
渡邉らは、大腸癌患者43症例のうち15例が変異型患者で、原発巣と転移巣の一致率は88.4%と報告している(非特許文献3参照。)。つまり、約1割は、原発巣と転移巣のKRAS変異ステイタスが一致しないことになる。一方、EGFR阻害剤治療中の原発巣と転移巣の変化については、S.Gattenlohnerらが報告している。転移性大腸癌21症例の治療前後のステイタスを検討した結果、20例(95.2%)で変化なしだった。変化のあった1例はヘテロジェナイティーがあり、かつ多発例であった(非特許文献4参照。)。
前記第一態様において、前記被験者の腫瘍から採取された組織検体又は細胞検体から検出されたKRAS遺伝子由来核酸又はそのタンパク質が、前記血液サンプルから検出された前記KRAS遺伝子由来核酸又はそのタンパク質の遺伝子型と相違する場合であっても、前記工程(b)において、前記被験者から採取された前記血液サンプル中に、前記変異型のKRAS遺伝子由来核酸又はそのタンパク質が検出された場合には、前記被験者の前記腫瘍はEGFR阻害剤感受性ではない可能性が高いと判定してもよい。
前記第一態様において、前記被験者が、過去に腫瘍部分の外科的切除処置を受けたことがあってもよい。
前記第一態様において、前記被験者が、過去にEGFR阻害剤を投与されたことがあってもよい。
前記第一態様において、前記被験者が、過去に前記EGFR阻害剤に対して薬剤耐性を示していてもよい。
前記第一態様において、前記血液サンプルが、前記EGFR阻害剤を投薬された後60日間が経過した被験者から採取されたものであってもよい。
前記第一態様において、前記被験者が、EGFR阻害剤の投薬処置を受けた後に、当該EGFR阻害剤投薬処置とは異なる他の抗腫瘍療法を受けた腫瘍患者であり、前記血液サンプルが、前記腫瘍患者が再び前記EGFR阻害剤投薬処置を受けようとする前に採取されたものであってもよい。
前記第一態様において、前記他の抗腫瘍療法が化学療法剤の投薬治療であってもよい。
前記第一態様において、前記化学療法剤が、フルオロウラシル、フォリン酸、オキサリプラチン、イリノテカン、シタラビン、フルダラビン、ゲムシタビン、ヒドロキシウレア、メトトレキセート、ブレオマイシン、クロラムブシル、シスプラチン、シクロフォスファミド、ドキソルビシン、ミトキサントロン、カンプトセシン、トポテカン、テニポシド、コルセミド、コルヒチン、パクリタキセル、ビンブラスチン、ビンクリスチン、及びタモキシフェンからなる群より選択される1種又は2種以上であってもよい。
前記第一態様において、前記他の抗腫瘍療法が放射線治療であってもよい。
前記第一態様において、前記他の抗腫瘍療法が、前記被験者に既に投薬されたEGFR阻害剤とは異なる種類の分子標的薬剤の投薬治療であってもよい。
前記第一態様において、前記分子標的薬剤が、セツキシマブ、パニツムマブ、ベバシヅマブ、ゲフィニチブ、エルロチニブ、レゴラフェニブ、クリゾチニブ、スニチニブ、ソラフェニブ、エベロリムス、トラスツズマブ、ラパチニブ、及びリツキシマブからなる群より選択される1種又は2種以上であってもよい。
前記第一態様において、前記他の抗腫瘍療法が、前記分子標的薬剤の投薬治療と、化学療法剤の投薬治療との併用療法であってもよい。
前記第一態様において、前記腫瘍が、再発性腫瘍であってもよい。
前記第一態様において、前記腫瘍が、転移巣であってもよい。
前記第一態様において、前記腫瘍が、原発巣であってもよい。
前記第一態様において、前記腫瘍が、大腸癌、結腸癌、直腸癌、肺癌、肝癌、乳癌、卵巣癌、前立腺癌、腎癌、食道癌、頭頸部癌、子宮癌、及び子宮頸癌からなる群より選択される1種又は2種以上であってもよい。
前記第一態様において、前記腫瘍が、前記被験者の体内の複数個所に存在していてもよい。
前記第一態様において、前記変異型が、KRASタンパク質のG12A、G12C、G12D、G12R、G12S、G12V、G13D、G12S2、G13A、G13S、G13V、G13R、G13C、Q61H、Q61L、Q61R、A146T、及びA146Vからなる群より選択される1又は2以上であってもよい。
前記第一態様において、前記血液サンプル中のKRASの存在の有無、及び野生型か変異型かの決定を、当該血液サンプル中の循環DNAから、野生型のKRAS遺伝子由来核酸が検出されるか否か、及び変異型のKRAS遺伝子由来核酸が検出されるか否かを調べることにより行ってもよい。
前記第一態様において、前記血液サンプルが、末梢血液、血清、血漿であってもよい。
前記第一態様において、前記血液サンプル中のCEAが5ng/mL以下であってもよいし、又は前記血液サンプル中のCA19-9の値が37.0U/mL以下であってもよい。
血中のKRASが野生型であり、変異型が検出されなかった被験者は、腫瘍組織中のKRASが野生型か変異型かにかかわらず、EGFR阻害剤に対する感受性は高いと予測される。逆に、血中から変異型のKRASが検出された被験者は、腫瘍組織中のKRASが野生型か変異型かにかかわらず、EGFR阻害剤に対する感受性は低いと予測される。
(a)被験者から採取された血液サンプル中に、KRAS遺伝子由来核酸又はそのタンパク質が存在するか否か、及び当該血液サンプル中のKRASが、野生型か変異型かを決定する工程と、(b)前記工程(a)において、前記血液サンプル中に、野生型のKRAS遺伝子由来核酸又はそのタンパク質が検出され、かつ変異型のKRAS遺伝子由来核酸又はそのタンパク質が検出されなかった場合には、前記被験者の腫瘍はEGFR阻害剤感受性である可能性が高いと判定し、前記血液サンプル中に、変異型のKRAS遺伝子由来核酸又はそのタンパク質が検出された場合には、前記被験者の腫瘍はEGFR阻害剤感受性ではない可能性が高いと判定する工程。
また、再発性腫瘍であってもよい。さらに、腫瘍が、被験者の体内の複数個所に存在していてもよい。
この時に、前記の一部三重塩基になる部分ができ、目的の突然変異をフラップエンドヌクレアーゼが切断し、FRETカセット内の蛍光修飾されたDNA断片が遊離して、FRET内の消光物質と乖離するために蛍光を発する。理論上、一度切断されたアレルプローブのフラップ部は、別のFRETカセットに再度ハイブリダイゼーションすることができ、シグナルが増幅される非常に高感度に突然変異を検出できる手法である。
原発巣を外科的切除した再発性大腸癌患者23名について、原発巣、転移巣、及び転移巣確認後に採取された血液から調製された血清に含まれるKRASについて、ノンシノニマスなアミノ酸置換を伴う突然変異を調べた。
原発巣の外科的切除手術の術前又は術後に、再発性大腸癌患者の末梢血6mLを採血後、遠心分離処理(3,000rpm、10分間)を行い、血清成分を得た。さらに、ID番号9の患者については、転移巣の外科的切除手術の術後にも、末梢血6mLを採血し、同様にして血清成分を得た(サンプル番号16)。また、一部の患者の原発巣、転移巣のホルマリン固定パラフィン包埋(FFPE)切片も実験サンプルとした。なお、本試験は、日本医科大学付属病院内の倫理審査委員会で承認されており、全ての患者からは本研究を包含するインフォームドコンセントを得て行った。本試験における患者情報について表2に示す。表2中、「転移巣」の欄の「-」は、転移巣が確認される前の患者であることを意味する。また、表2中、「セツキシマブ」の欄の「あり」は、転移巣の外科的切除手術後(但し、サンプル番号8とサンプル番号15は、原発巣の外科的切除手術前)にセツキシマブの投与処置を行ったことを、同欄の「なし」は、同投与処置を行わなかったことを、それぞれ意味する。表2中、「化学療法」の欄は、転移巣の外科的切除手術後のセツキシマブ投与処置時点(但し、サンプル番号20と21では、原発巣の外科的切除手術後のセツキシマブ投与処置時点、サンプル番号16では血液サンプル採取時点)における、化学療法の施行状況を示す。より詳細には、「なし」は投与未定であり、今後投与の可能性がある患者を意味し、「施行前」は施行が予定されていることを意味し、「施行中」は化学療法が奏効しており、投与を続けている状態を意味し、「施行後」は化学療法が奏効していたが、当該時点では投与を終了していることを意味し、「終了後」は化学療法が効かなくなった状態(すなわち、緩和ケアに入っている状態)を意味する。
血清中のCEA及びCA-19-9を、CLEIA法(化学発光酵素免疫測定法)により測定した。測定結果を表5に示す。
血清からのcfDNAの単離精製は、QIAamp Circulating Nucleic Acid Kit(キアゲン社)を用いて行った。本キットに供した血清サンプルの量は、患者によって異なり、2mL~4mlであった。DNAの単離精製工程は、キットに付属されているインストラクションに従った。スピンカラムからの最終溶出は、TE緩衝液50μLを用いて行った。
FFPE切片からのDNA単離精製は、QIAamp DNA FFPE Tissue Kit(キアゲン社)を用いて行った。1サンプルにつき、FFPE切片は10μmにスライスされたものを3枚用いた。DNAの単離精製工程は、キットに付属されているインストラクションに従った。スピンカラムからの最終溶出は、TE緩衝液100μLを用いて行った。
cfDNA及びFFPE切片から単離精製したDNAの定量は、Quant-iT(登録商標)PicoGreen(登録商標)dsDNA Reagent and Kits(invitorogen社)を用いて行った。測定するサンプルは全て、単離したDNAをTE緩衝液で20倍に希釈したものを用いた。蛍光測定装置は、SAFIRA(TECAN社)を用いた。
原発巣や転移巣の手術標本、並びに血清中のKRAS塩基配列解析は、ダイレクトシークエンシングにて行った。KRASのダイレクトシークエンス用のプライマー配列として、KRAS(フォワード):5’-GAATGGTCCTGCACCAGTAA-3’(配列番号5)、KRAS(リバース):5’-GTGTGACATGTTCTAATATAGTCA-3’(配列番号6)を用いた。各PCR産物の長さは214bpであり、PCR条件は、95℃で10分間の前変性後、94℃で20秒間、60℃で20秒間、72℃で30秒間を1サイクルとして40サイクル行い、その後72℃で10分間の伸長反応を行った。シークエンス解析はABI3730(アプライドバイオシステムズ、Foster City、CA)を用いて、ビッグダイターミネーター法によるサイクルシークエンシングを行った。結果を表5に示す。表5の「血清」欄中、遺伝子型の語尾の「(術前)」と「(術後)」は、それぞれ原発巣(サンプル番号16のみ、転移巣)の外科的切除手術の術前又は術後に採取された血清中の遺伝子型であることを意味する。また、表5の「転移巣」の欄中、「-」は、転移巣中のKRASの遺伝子型を解析していないことを意味する。
Claims (22)
- (a)被験者から採取された血液サンプル中に、KRAS遺伝子由来核酸又はそのタンパク質が存在するか否か、及び当該血液サンプル中の前記KRAS遺伝子由来核酸又はそのタンパク質が、野生型か変異型かを決定する工程と、
(b)前記工程(a)において、前記血液サンプル中に、野生型のKRAS遺伝子由来核酸又はそのタンパク質が検出され、かつ変異型のKRAS遺伝子由来核酸又はそのタンパク質が検出されなかった場合には、前記被験者の腫瘍はEGFR阻害剤感受性である可能性が高いと判定し、前記血液サンプル中に、変異型のKRAS遺伝子由来核酸又はそのタンパク質が検出された場合には、前記被験者の腫瘍はEGFR阻害剤感受性ではない可能性が高いと判定する工程と、
を有するEGFR阻害剤感受性予測方法。 - 前記被験者の腫瘍から採取された組織検体又は細胞検体から検出されたKRAS遺伝子由来核酸又はそのタンパク質が、前記血液サンプルから検出された前記KRAS遺伝子由来核酸又はそのタンパク質の遺伝子型と相違する場合であっても、
前記工程(b)において、前記被験者から採取された前記血液サンプル中に、前記変異型のKRAS遺伝子由来核酸又はそのタンパク質が検出された場合には、前記被験者の前記腫瘍はEGFR阻害剤感受性ではない可能性が高いと判定する、請求項1に記載のEGFR阻害剤感受性予測方法。 - 前記被験者が、過去に腫瘍部分の外科的切除処置を受けたことがある、請求項1又は2に記載のEGFR阻害剤感受性予測方法。
- 前記被験者が、過去にEGFR阻害剤を投与されたことがある、請求項1~3のいずれか一項に記載のEGFR阻害剤感受性予測方法。
- 前記被験者が、過去に前記EGFR阻害剤に対して薬剤耐性を示していた、請求項4に記載のEGFR阻害剤感受性予測方法。
- 前記血液サンプルが、前記EGFR阻害剤を投薬された後60日間が経過した被験者から採取されたものである、請求項4又は5に記載のEGFR阻害剤感受性予測方法。
- 前記被験者が、EGFR阻害剤の投薬処置を受けた後に、当該EGFR阻害剤投薬処置とは異なる他の抗腫瘍療法を受けた腫瘍患者であり、
前記血液サンプルが、前記腫瘍患者が再び前記EGFR阻害剤投薬処置を受けようとする前に採取されたものである、請求項1又は2に記載のEGFR阻害剤感受性予測方法。 - 前記他の抗腫瘍療法が化学療法剤の投薬治療である、請求項7に記載のEGFR阻害剤感受性予測方法。
- 前記化学療法剤が、フルオロウラシル、フォリン酸、オキサリプラチン、イリノテカン、シタラビン、フルダラビン、ゲムシタビン、ヒドロキシウレア、メトトレキセート、ブレオマイシン、クロラムブシル、シスプラチン、シクロフォスファミド、ドキソルビシン、ミトキサントロン、カンプトセシン、トポテカン、テニポシド、コルセミド、コルヒチン、パクリタキセル、ビンブラスチン、ビンクリスチン、及びタモキシフェンからなる群より選択される1種又は2種以上である、請求項8に記載のEGFR阻害剤感受性予測方法。
- 前記他の抗腫瘍療法が放射線治療である、請求項7に記載のEGFR阻害剤感受性予測方法。
- 前記他の抗腫瘍療法が、前記被験者に既に投薬されたEGFR阻害剤とは異なる種類の分子標的薬剤の投薬治療である、請求項7に記載のEGFR阻害剤感受性予測方法。
- 前記分子標的薬剤が、セツキシマブ、パニツムマブ、ベバシヅマブ、ゲフィニチブ、エルロチニブ、レゴラフェニブ、クリゾチニブ、スニチニブ、ソラフェニブ、エベロリムス、トラスツズマブ、ラパチニブ、及びリツキシマブからなる群より選択される1種又は2種以上である、請求項11に記載のEGFR阻害剤感受性予測方法。
- 前記他の抗腫瘍療法が、前記分子標的薬剤の投薬治療と、化学療法剤の投薬治療との併用療法である、請求項11又は12に記載のEGFR阻害剤感受性予測方法。
- 前記腫瘍が、再発性腫瘍である、請求項1~13のいずれか一項に記載のEGFR阻害剤感受性予測方法。
- 前記腫瘍が、転移巣である、請求項1~13のいずれか一項に記載のEGFR阻害剤感受性予測方法。
- 前記腫瘍が、原発巣である、請求項1~12のいずれか一項に記載のEGFR阻害剤感受性予測方法。
- 前記腫瘍が、大腸癌、結腸癌、直腸癌、肺癌、肝癌、乳癌、卵巣癌、前立腺癌、腎癌、食道癌、頭頸部癌、子宮癌、及び子宮頸癌からなる群より選択される1種又は2種以上である、1~16のいずれか一項に記載のEGFR阻害剤感受性予測方法。
- 前記腫瘍が、前記被験者の体内の複数個所に存在している、請求項1~17のいずれか一項に記載のEGFR阻害剤感受性予測方法。
- 前記変異型が、KRASタンパク質のG12A、G12C、G12D、G12R、G12S、G12V、G13D、G12S2、G13A、G13S、G13V、G13R、G13C、Q61H、Q61L、Q61R、A146T、及びA146Vからなる群より選択される1又は2以上である、請求項1~18のいずれか一項に記載のEGFR阻害剤感受性予測方法。
- 前記血液サンプル中のKRASの存在の有無、及び野生型か変異型かの決定を、当該血液サンプル中の循環DNAから、野生型のKRAS遺伝子由来核酸が検出されるか否か、及び変異型のKRAS遺伝子由来核酸が検出されるか否かを調べることにより行う、請求項1~19のいずれか一項に記載のEGFR阻害剤感受性予測方法。
- 前記血液サンプルが、末梢血液、血清、血漿である、請求項1~20のいずれか一項に記載のEGFR阻害剤感受性予測方法。
- 前記血液サンプル中のCEAが5ng/mL以下である、又は前記血液サンプル中のCA19-9の値が37.0U/mL以下である、請求項1~21のいずれか一項に記載のEGFR阻害剤感受性予測方法。
Priority Applications (5)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| EP14767593.8A EP2977464A4 (en) | 2013-03-19 | 2014-03-19 | PROCEDURE FOR PREDICTING SENSITIVITY TO EGFR HEMMER |
| SG11201507739TA SG11201507739TA (en) | 2013-03-19 | 2014-03-19 | Method for predicting sensitivity to egfr inhibitor |
| JP2015506829A JP6520705B2 (ja) | 2013-03-19 | 2014-03-19 | Egfr阻害剤感受性予測方法 |
| CN201480017000.0A CN105074009B (zh) | 2013-03-19 | 2014-03-19 | Egfr抑制剂的敏感性的预测方法 |
| US14/857,187 US20160002741A1 (en) | 2013-03-19 | 2015-09-17 | Method for predicting sensitivity to egfr inhibitor |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2013-057033 | 2013-03-19 | ||
| JP2013057033 | 2013-03-19 |
Related Child Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| US14/857,187 Continuation US20160002741A1 (en) | 2013-03-19 | 2015-09-17 | Method for predicting sensitivity to egfr inhibitor |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2014148557A1 true WO2014148557A1 (ja) | 2014-09-25 |
Family
ID=51580231
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/JP2014/057556 Ceased WO2014148557A1 (ja) | 2013-03-19 | 2014-03-19 | Egfr阻害剤感受性予測方法 |
Country Status (6)
| Country | Link |
|---|---|
| US (1) | US20160002741A1 (ja) |
| EP (1) | EP2977464A4 (ja) |
| JP (1) | JP6520705B2 (ja) |
| CN (1) | CN105074009B (ja) |
| SG (2) | SG10201707548RA (ja) |
| WO (1) | WO2014148557A1 (ja) |
Cited By (8)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2015072555A1 (ja) * | 2013-11-15 | 2015-05-21 | 凸版印刷株式会社 | Vegf阻害剤長期奏功性予測方法 |
| WO2016103727A1 (ja) * | 2014-12-25 | 2016-06-30 | 凸版印刷株式会社 | 特異的核酸配列の増幅促進方法 |
| WO2017110764A1 (ja) * | 2015-12-24 | 2017-06-29 | 公益財団法人がん研究会 | マルチキナーゼ阻害剤の有効性と安全性を予測する検査方法、検査キット、及びバイオマーカー |
| WO2018131705A1 (ja) * | 2017-01-16 | 2018-07-19 | 凸版印刷株式会社 | 腫瘍マーカー遺伝子の体細胞変異の検出方法及び腫瘍状態の評価方法 |
| KR20180136515A (ko) | 2016-04-28 | 2018-12-24 | 덴카 주식회사 | 상피성 세포 성장 인자 수용체 저해제에 대한 암세포의 내성을 판정하는 방법 |
| JP2020532282A (ja) * | 2017-06-30 | 2020-11-12 | コリア アドバンスト インスティチュート オブ サイエンス アンド テクノロジー | Vegf−grabタンパク質と薬物の結合体及びその用途 |
| US11725010B2 (en) | 2019-12-02 | 2023-08-15 | Storm Therapeutics Limited | Polyheterocyclic compounds as METTL3 inhibitors |
| JP2023176123A (ja) * | 2022-05-31 | 2023-12-13 | 株式会社 光バイオ | ウイルスベクター及びそれを含むがん細胞増殖抑制剤 |
Families Citing this family (13)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2012129363A2 (en) | 2011-03-24 | 2012-09-27 | President And Fellows Of Harvard College | Single cell nucleic acid detection and analysis |
| US10876152B2 (en) | 2012-09-04 | 2020-12-29 | Guardant Health, Inc. | Systems and methods to detect rare mutations and copy number variation |
| US11913065B2 (en) | 2012-09-04 | 2024-02-27 | Guardent Health, Inc. | Systems and methods to detect rare mutations and copy number variation |
| GB2533006B (en) | 2012-09-04 | 2017-06-07 | Guardant Health Inc | Systems and methods to detect copy number variation |
| US20160040229A1 (en) | 2013-08-16 | 2016-02-11 | Guardant Health, Inc. | Systems and methods to detect rare mutations and copy number variation |
| EP3771745A1 (en) | 2013-12-28 | 2021-02-03 | Guardant Health, Inc. | Methods and systems for detecting genetic variants |
| EP3390668A4 (en) | 2015-12-17 | 2020-04-01 | Guardant Health, Inc. | METHOD FOR DETERMINING THE TUMORGEN COPY NUMBER BY ANALYSIS OF CELL-FREE DNA |
| JP2019202936A (ja) * | 2016-08-17 | 2019-11-28 | 中外製薬株式会社 | 抗Epiregulin抗体と抗EGFR抗体との併用医薬 |
| US20200347140A1 (en) * | 2017-08-30 | 2020-11-05 | Symphogen A/S | Compositions and methods for treating cancer with anti-egfr antibodies |
| CN108998535B (zh) * | 2018-09-11 | 2021-09-10 | 广州市宝创生物技术有限公司 | 一种kras基因多重突变位点检测试剂盒 |
| CN112162042B (zh) * | 2020-09-14 | 2022-04-26 | 首都医科大学附属北京朝阳医院 | 超高效液相色谱串联质谱测定血浆中amg510浓度的方法 |
| AU2022233518A1 (en) * | 2021-03-09 | 2023-10-26 | Janssen Biotech, Inc. | Treatment of cancers lacking egfr-activating mutations |
| WO2026041666A1 (en) * | 2024-08-20 | 2026-02-26 | Genome Research Limited | Mitogen signalling pathway protein variants |
Citations (7)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| EP0332435A2 (en) | 1988-03-10 | 1989-09-13 | Zeneca Limited | Method of detecting nucleotide sequences |
| US4943533A (en) | 1984-03-01 | 1990-07-24 | The Regents Of The University Of California | Hybrid cell lines that produce monoclonal antibodies to epidermal growth factor receptor |
| US5679683A (en) | 1994-01-25 | 1997-10-21 | Warner-Lambert Company | Tricyclic compounds capable of inhibiting tyrosine kinases of the epidermal growth factor receptor family |
| JP2010521154A (ja) * | 2007-03-13 | 2010-06-24 | アムゲン インコーポレイティッド | K‐ras変異およびB‐raf変異ならびに抗EGFr抗体療法 |
| JP2010521153A (ja) * | 2007-03-13 | 2010-06-24 | アムゲン インコーポレイティッド | K‐ras変異および抗EGFr抗体療法 |
| US20110320392A1 (en) * | 2009-03-02 | 2011-12-29 | Black Esther P | Methods for predicting cancer response to egfr inhibitors |
| US20140134158A1 (en) * | 2012-05-22 | 2014-05-15 | Alberto Bardelli | Kras mutations and resistance to anti-egfr treatment |
Family Cites Families (1)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JP2012135290A (ja) * | 2010-12-28 | 2012-07-19 | Fuaruko Bio Syst:Kk | 高感度K−ras遺伝子変異解析法による分子標的薬の治療感受性の評価法 |
-
2014
- 2014-03-19 EP EP14767593.8A patent/EP2977464A4/en not_active Withdrawn
- 2014-03-19 WO PCT/JP2014/057556 patent/WO2014148557A1/ja not_active Ceased
- 2014-03-19 CN CN201480017000.0A patent/CN105074009B/zh not_active Expired - Fee Related
- 2014-03-19 SG SG10201707548RA patent/SG10201707548RA/en unknown
- 2014-03-19 JP JP2015506829A patent/JP6520705B2/ja active Active
- 2014-03-19 SG SG11201507739TA patent/SG11201507739TA/en unknown
-
2015
- 2015-09-17 US US14/857,187 patent/US20160002741A1/en not_active Abandoned
Patent Citations (7)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US4943533A (en) | 1984-03-01 | 1990-07-24 | The Regents Of The University Of California | Hybrid cell lines that produce monoclonal antibodies to epidermal growth factor receptor |
| EP0332435A2 (en) | 1988-03-10 | 1989-09-13 | Zeneca Limited | Method of detecting nucleotide sequences |
| US5679683A (en) | 1994-01-25 | 1997-10-21 | Warner-Lambert Company | Tricyclic compounds capable of inhibiting tyrosine kinases of the epidermal growth factor receptor family |
| JP2010521154A (ja) * | 2007-03-13 | 2010-06-24 | アムゲン インコーポレイティッド | K‐ras変異およびB‐raf変異ならびに抗EGFr抗体療法 |
| JP2010521153A (ja) * | 2007-03-13 | 2010-06-24 | アムゲン インコーポレイティッド | K‐ras変異および抗EGFr抗体療法 |
| US20110320392A1 (en) * | 2009-03-02 | 2011-12-29 | Black Esther P | Methods for predicting cancer response to egfr inhibitors |
| US20140134158A1 (en) * | 2012-05-22 | 2014-05-15 | Alberto Bardelli | Kras mutations and resistance to anti-egfr treatment |
Non-Patent Citations (19)
| Title |
|---|
| BOKEMEYER ET AL., ANNALS OF ONCOLOGY, vol. 22, no. 7, 2011, pages 1535 - 1546 |
| CUTSEM ET AL., THE NEW ENGLAND JOURNAL OF MEDICINE, vol. 360, 2009, pages 1408 - 1417 |
| DIAZ JR ET AL., NATURE, vol. 486, 2012, pages 537 - 540 |
| DIAZ LA JR. ET AL.: "The molecular evolution of acquired resistance to targeted EGFR blockade in colorectal cancers", NATURE, vol. 486, no. 7404, 2012, pages 537 - 540, XP055287050 * |
| DVIR ET AL., JOURNAL OF CELL BIOLOGY, vol. 113, 1991, pages 857 - 865 |
| GATTENLOHNER ET AL., NEW ENGLAND JOURNAL OF MEDICINE, vol. 360, no. 8, 2009, pages 835 |
| GENOMICS, vol. 5, 1989, pages 874 - 879 |
| HINDSON ET AL., ANALYTICAL CHEMISTRY, vol. 83, no. 22, 2011, pages 8604 - 8610 |
| MICHAEL OLIVIER, MUTATION RESEARCH, vol. 573, 2005, pages 103 - 110 |
| MISALE ET AL., NATURE, vol. 486, no. 7404, 2012, pages 532 - 536 |
| MISALE S. ET AL.: "Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer", NATURE, vol. 486, no. 7404, 2012, pages 532 - 536, XP055287051 * |
| NEWTON ET AL., NUCLEIC ACIDS RESEARCH, vol. 17, no. 7, 1989 |
| ORITA, PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES, USA, vol. 86, 1989, pages 2766 - 2770 |
| PANEK ET AL., JOURNAL OF PHARMACOLOGY AND EXPERIMENTRAL THERAPEUTICS, vol. 283, 1997, pages 1433 - 1444 |
| RUANO; KIDD, NUCLEIC ACIDS RESEARCH, vol. 17, 1989, pages 8392 |
| SANTINI ET AL., ANNALS OF ONCOLOGY, vol. 23, 2012, pages 2313 - 2318 |
| See also references of EP2977464A4 |
| WATANABE ET AL., DISEASES OF THE COLON AND RECT, vol. 54, 2011, pages 1170 - 1178 |
| WU ET AL., GENOMICS, vol. 4, 1989, pages 560 - 569 |
Cited By (15)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2015072555A1 (ja) * | 2013-11-15 | 2015-05-21 | 凸版印刷株式会社 | Vegf阻害剤長期奏功性予測方法 |
| WO2016103727A1 (ja) * | 2014-12-25 | 2016-06-30 | 凸版印刷株式会社 | 特異的核酸配列の増幅促進方法 |
| WO2017110764A1 (ja) * | 2015-12-24 | 2017-06-29 | 公益財団法人がん研究会 | マルチキナーゼ阻害剤の有効性と安全性を予測する検査方法、検査キット、及びバイオマーカー |
| JPWO2017110764A1 (ja) * | 2015-12-24 | 2018-11-01 | 公益財団法人がん研究会 | マルチキナーゼ阻害剤の有効性と安全性を予測する検査方法、検査キット、及びバイオマーカー |
| KR20180136515A (ko) | 2016-04-28 | 2018-12-24 | 덴카 주식회사 | 상피성 세포 성장 인자 수용체 저해제에 대한 암세포의 내성을 판정하는 방법 |
| JPWO2018131705A1 (ja) * | 2017-01-16 | 2019-11-07 | 凸版印刷株式会社 | 腫瘍マーカー遺伝子の体細胞変異の検出方法及び腫瘍状態の評価方法 |
| WO2018131705A1 (ja) * | 2017-01-16 | 2018-07-19 | 凸版印刷株式会社 | 腫瘍マーカー遺伝子の体細胞変異の検出方法及び腫瘍状態の評価方法 |
| JP7215675B2 (ja) | 2017-01-16 | 2023-01-31 | 凸版印刷株式会社 | 腫瘍状態の評価方法 |
| US11746382B2 (en) | 2017-01-16 | 2023-09-05 | Toppan Printing Co., Ltd. | Method of detecting somatic mutations in tumor marker genes, and tumor status evaluation method |
| JP2020532282A (ja) * | 2017-06-30 | 2020-11-12 | コリア アドバンスト インスティチュート オブ サイエンス アンド テクノロジー | Vegf−grabタンパク質と薬物の結合体及びその用途 |
| JP2022028654A (ja) * | 2017-06-30 | 2022-02-16 | コリア アドバンスト インスティチュート オブ サイエンス アンド テクノロジー | Vegf-grabタンパク質と薬物の結合体及びその用途 |
| JP7348249B2 (ja) | 2017-06-30 | 2023-09-20 | コリア アドバンスト インスティチュート オブ サイエンス アンド テクノロジー | Vegf-grabタンパク質と薬物の結合体及びその用途 |
| US11725010B2 (en) | 2019-12-02 | 2023-08-15 | Storm Therapeutics Limited | Polyheterocyclic compounds as METTL3 inhibitors |
| US12195458B2 (en) | 2019-12-02 | 2025-01-14 | Storm Therapeutics Limited | Polyheterocyclic compounds as METTL3 inhibitors |
| JP2023176123A (ja) * | 2022-05-31 | 2023-12-13 | 株式会社 光バイオ | ウイルスベクター及びそれを含むがん細胞増殖抑制剤 |
Also Published As
| Publication number | Publication date |
|---|---|
| US20160002741A1 (en) | 2016-01-07 |
| JPWO2014148557A1 (ja) | 2017-02-16 |
| CN105074009B (zh) | 2017-09-26 |
| EP2977464A4 (en) | 2016-10-19 |
| CN105074009A (zh) | 2015-11-18 |
| SG11201507739TA (en) | 2015-10-29 |
| EP2977464A1 (en) | 2016-01-27 |
| SG10201707548RA (en) | 2017-10-30 |
| JP6520705B2 (ja) | 2019-05-29 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP6520705B2 (ja) | Egfr阻害剤感受性予測方法 | |
| Guo et al. | MET-dependent solid tumours—molecular diagnosis and targeted therapy | |
| JP5827935B2 (ja) | Egfrおよびkras変異 | |
| EP2592155B2 (en) | EGFR mutations | |
| US20170002425A1 (en) | Method for predicting long-term efficacy of vegf inhibitor | |
| JP7215675B2 (ja) | 腫瘍状態の評価方法 | |
| JP6837632B2 (ja) | 体液試料の単位量当たりのセルフリーdna量を健康状態の評価のための指標とする方法 | |
| AU2013200810B2 (en) | EGFR Mutations | |
| WO2013177705A1 (en) | Genotyping assay for identifying chromosomal rearrangements | |
| AU2015203691B2 (en) | EGFR Mutations | |
| Pechanska | A novel approach to develop predictive biomarkers | |
| Richter-Pechańska | A novel approach to develop predictive biomarkers: prediction of response to anti-EGFR therapy in a large panel of patient-derived colorectal cancer xenograft models |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| WWE | Wipo information: entry into national phase |
Ref document number: 201480017000.0 Country of ref document: CN |
|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 14767593 Country of ref document: EP Kind code of ref document: A1 |
|
| ENP | Entry into the national phase |
Ref document number: 2015506829 Country of ref document: JP Kind code of ref document: A |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 2014767593 Country of ref document: EP |