WO2024254530A1 - Pleckstrin holomogy domains for delivery of protein payloads to a target cell - Google Patents
Pleckstrin holomogy domains for delivery of protein payloads to a target cell Download PDFInfo
- Publication number
- WO2024254530A1 WO2024254530A1 PCT/US2024/033111 US2024033111W WO2024254530A1 WO 2024254530 A1 WO2024254530 A1 WO 2024254530A1 US 2024033111 W US2024033111 W US 2024033111W WO 2024254530 A1 WO2024254530 A1 WO 2024254530A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- protein
- chimeric protein
- cases
- sequence
- domain
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/113—Non-coding nucleic acids modulating the expression of genes, e.g. antisense oligonucleotides; Antisense DNA or RNA; Triplex- forming oligonucleotides; Catalytic nucleic acids, e.g. ribozymes; Nucleic acids used in co-suppression or gene silencing
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K14/00—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof
- C07K14/435—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans
- C07K14/46—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates
- C07K14/47—Peptides having more than 20 amino acids; Gastrins; Somatostatins; Melanotropins; Derivatives thereof from animals; from humans from vertebrates from mammals
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/14—Hydrolases (3)
- C12N9/16—Hydrolases (3) acting on ester bonds (3.1)
- C12N9/22—Ribonucleases [RNase]; Deoxyribonucleases [DNase]
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/01—Fusion polypeptide containing a localisation/targetting motif
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/60—Fusion polypeptide containing spectroscopic/fluorescent detection, e.g. green fluorescent protein [GFP]
-
- C—CHEMISTRY; METALLURGY
- C07—ORGANIC CHEMISTRY
- C07K—PEPTIDES
- C07K2319/00—Fusion polypeptide
- C07K2319/61—Fusion polypeptide containing an enzyme fusion for detection (lacZ, luciferase)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2310/00—Structure or type of the nucleic acid
- C12N2310/10—Type of nucleic acid
- C12N2310/20—Type of nucleic acid involving clustered regularly interspaced short palindromic repeats [CRISPR]
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N2740/00—Reverse transcribing RNA viruses
- C12N2740/00011—Details
- C12N2740/10011—Retroviridae
- C12N2740/10022—New viral proteins or individual genes, new structural or functional aspects of known viral proteins or genes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12Y—ENZYMES
- C12Y302/00—Hydrolases acting on glycosyl compounds, i.e. glycosylases (3.2)
- C12Y302/01—Glycosidases, i.e. enzymes hydrolysing O- and S-glycosyl compounds (3.2.1)
- C12Y302/01052—Beta-N-acetylhexosaminidase (3.2.1.52)
Definitions
- a chimeric protein comprising a Pleckstrin Homology domain comprising: (i) at least 60% at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99.1%, at least 99.2%, at least 99.3%, at least 99.4%, at least 99.5%, at least 99.6%, at least 99.7%, at least 99.8%, or at least 99.9% sequence identity to the sequence of SEQ ID NO: 569; and (ii) at least two selected from the group of amino acid substitutions consisting of E17K, R25C, T81Y, T101C, and a combination of K142A, Hl 43 A, R144A (142-144A) relative to the sequence of SEQ ID NO: 569
- the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K and R25C; E17K and T81Y; E17K and T101C; E17K and 142-144A; R25C and T81Y; R25C and T101C; R25C and 142-144A; T81Y and T101C; T81Y and 142-144A; and T101C and 142-144A.
- the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 572, 573, 574, 575, 576, 577, 578, 579, or 580.
- the chimeric protein comprises at least three mutations selected from the group consisting of E17K, R25C, T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569.
- the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K, R25C, and T81Y; E17K, R25C, and T101C; E17K, R25C, and 142-144A; E17K, T81Y, and T101C; E17K, T81Y, and 142-144A; E17K, T101C, and 142-144A; R25C, T81Y, and T101C; R25C, T81Y, and 142-144A; R25C, T101C, and 142-144A; and T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569.
- the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 581, 582, 583, 584, 585,586, 587, 588, 589, or 590. In some cases, the chimeric protein comprises at least four selected from the group consisting of E17K, R25C, T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569.
- the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K, R25C, T81Y, and T101C; E17K, R25C, T81Y, and 142- 144A; R25C, T81Y, T101C, and 142-144A; R25C, T81Y, T101C, and 142-144A; and E17K, T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569.
- the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 591, 592, 593, 594, or 595.
- the chimeric protein comprises at least the mutations E17K, R25C, T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569. In some cases, the chimeric protein comprises the sequence of SEQ ID NO: 596.
- the chimeric protein further comprises one or more nuclear export sequences.
- the one or more nuclear export sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 5.
- the chimeric protein further comprises one or more nuclear localization sequences.
- the one or more nuclear localization sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 6.
- the Pleckstrin Homology domain is coupled to the protein payload. In some cases, the Pleckstrin Homology domain is reversibly coupled to the protein payload. In some cases, the protein payload is reversibly coupled to the Pleckstrin Homology domain by a cleavable linker. In some cases, the cleavable linker is cleavable by a protease. In some cases, the protein payload is coupled to a C- terminal of the Pleckstrin Homology domain.
- the protein payload is coupled to an N-terminal of the Pleckstrin Homology domain.
- the protein payload comprises one or more viral proteins.
- the one or more viral proteins comprise one or more retroviral proteins.
- the one or more retroviral proteins comprises a retroviral Gag protein.
- the protein payload comprises one or more non-viral proteins.
- the one or more non-viral proteins comprises one or more mammalian proteins.
- the one or more mammalian proteins comprises an Arc protein.
- the protein payload comprises a gene editing protein.
- the gene editing protein comprises a prime editing protein.
- the prime editing protein is coupled to a target-specific prime editing guide RNA.
- the gene editing protein comprises a CRISPR system.
- the CRISPR system comprises a Cas domain.
- the gene editing protein is coupled to a target-specific guide RNA.
- the protein payload comprises an epigenetic editing protein.
- the epigenetic editing protein is coupled to a target-specific guide RNA.
- the protein payload comprises a recombinase protein.
- the protein payload comprises an integrase protein.
- nucleic acid molecule encoding a chimeric protein.
- a lipid delivery particle comprising a chimeric protein.
- the lipid delivery particle further comprises an envelope, wherein the envelope comprises a lipid bilayer encasing a lumen, wherein the chimeric protein is located in the lumen.
- the lipid delivery particle further comprises a target ligand.
- the target ligand reversibly couples to a target receptor.
- the target receptor comprises a cell surface protein on a target cell.
- the envelope couples to the target cell at least partially through coupling of the target ligand and the target receptor on the target cell.
- coupling of the target ligand and the target receptor on the target cell induces release of the protein payload in the target cell.
- release of the protein payload in the target cell follows fusion of the envelope with cell membrane of the target cell.
- a method to deliver a payload to a target cell comprising delivering the lipid delivery particle to a target cell.
- a chimeric protein comprising: a Pleckstrin Homology domain comprising at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99.1%, at least 99.2%, at least 99.3%, at least 99.4%, at least 99.5%, at least 99.6%, at least 99.7%, at least 99.8%, or at least 99.9% sequence identity to any one of the sequences set forth in Table 4B; and a protein payload.
- a Pleckstrin Homology domain comprising at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at
- the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 597, 598, 599, 600, 601, 602, 603, 604, 605, 606, 607, 608, 609, 610, 611, 612, 613, 614, 615, 616, 617, 618, 619, 620, 621, 622, 623, 624, 625, 626, 627, 628, 629,
- the chimeric protein further comprises one or more nuclear export sequences.
- the one or more nuclear export sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 5.
- the chimeric protein further comprises one or more nuclear localization sequences.
- the one or more nuclear localization sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 6.
- the Pleckstrin Homology domain is coupled to the protein payload. In some cases, the Pleckstrin Homology domain is reversibly coupled to the protein payload. In some cases, the protein payload is reversibly coupled to the Pleckstrin Homology domain by a cleavable linker. In some cases, the cleavable linker is cleavable by a protease. In some cases, the protein payload is coupled to C-terminal of the Pleckstrin Homology domain. In some cases, the protein payload is coupled to N-terminal of the Pleckstrin Homology domain.
- the protein payload comprises one or more viral proteins.
- the one or more viral proteins comprise one or more retroviral proteins.
- the one or more retroviral proteins comprises a retroviral Gag protein.
- the protein payload comprises one or more non-viral proteins.
- the one or more non-viral proteins comprises one or more mammalian proteins.
- the one or more mammalian proteins comprises an Arc protein.
- the protein payload comprises a gene editing protein.
- the gene editing protein comprises a prime editing protein.
- the prime editing protein is coupled to a target-specific prime editing guide RNA.
- the gene editing protein comprises a CRISPR system.
- the CRISPR system comprises a Cas domain.
- the gene editing protein is coupled to a target-specific guide RNA.
- the protein payload comprises an epigenetic editing protein.
- the epigenetic editing protein is coupled to a target-specific guide RNA.
- the protein payload comprises a recombinase protein.
- the protein payload comprises an integrase protein.
- nucleic acid molecule encoding a chimeric protein.
- a lipid delivery particle comprising a chimeric protein.
- the lipid delivery particle further comprises an envelope, wherein the envelope comprises a lipid bilayer encasing a lumen, wherein the chimeric protein is located in the lumen.
- the lipid delivery particle further comprises a target ligand.
- the target ligand reversibly couples to a target receptor.
- the target receptor comprises a cell surface protein on a target cell.
- the envelope couples to the target cell through coupling of the target ligand and the target receptor on the target cell.
- coupling of the target ligand and the target receptor on the target cell induces release of the protein payload in the target cell.
- release of the protein payload in the target cell follows endocytosis of the lipid delivery particle.
- a method to deliver a payload to a target cell comprising: delivering the lipid delivery particle to a target cell.
- a chimeric protein comprising: a Pleckstrin Homology domain comprising: (i) at least 60% at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99.1%, at least 99.2%, at least 99.3%, at least 99.4%, at least 99.5%, at least 99.6%, at least 99.7%, at least 99.8%, or at least 99.9% sequence identity to the sequence of SEQ ID NO: 613; and (ii) at least one amino acid substitution selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K relative to the sequence of SEQ ID NO: 613; and a protein payload.
- the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 603, 604, 605, 606, or 607. In some cases, the chimeric protein comprises at least two mutations selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613.
- the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K and E58K; E17K and E52K; E17K and E53K; E17K and E55K; E17K and E65K; E58K and E52K; E58K and E53K; E58K and E55K; E58K and E65K; E52K and E53K; E52K and E55K; E52K and E65K; E53K and E55K; and E53K and E55K; and E53K and E65K, relative to the sequence of SEQ ID NO: 613.
- the chimeric protein comprises the sequence of SEQ ID NO: 608.
- the chimeric protein comprises at least three mutations selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613. [0026] In some cases, the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K, E58K, and E52K; E17K, E58K, and E53K; E17K, E58K, and E55K; E17K, E58K, and E65K; E17K, E52K, and E53K; E17K, E52K, and E55K; E17K, E52K, and E65K; E17K, E52K, and E65K; E17K, E52K, and E65K; E17K, E17K, and E65K; E17K, E17K, and E65K; E17K, E17K, and E65K; E17K, E17K, and E65K; E17
- E53K, and E55K E17K, E53K, and E65K; E58K, E52K, and E53K; E58K, E52K, and E55K; E58K,
- E52K, and E65K E58K, E53K, and E55K; E58K, E53K, and E65K; E52K, E53K, and E65K; and E52K,
- the chimeric protein comprises the sequence of SEQ ID NO: 609. In some cases, the chimeric protein comprises at least four mutations selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613.
- the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K, E58K, E52K, and E53K; E17K, E58K, E52K, and E55K; E17K, E58K, E52K, and E65K; E58K, E52K, E53K, and E55K; E58K, E52K, E53K, and E65K; E58K, E52K, E53K, and E65K; E58K, E52K, E53K, and E65K, relative to the sequence of SEQ ID NO: 613.
- the chimeric protein comprises the sequence of SEQ ID NO: 610.
- the chimeric protein comprises at least five selected from the group consisting of: E17K, E58K, E52K, E53K, and E55K; E17K, E58K, E52K, E53K, and E65K; E17K, E58K, E52K, E55K, and E65K; E17K, E58K, E53K, E55K, and E65K; E17K, E52K, E53K, E55K, and E65K; and E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613.
- the chimeric protein comprises the sequence of SEQ ID NO: 611.
- the chimeric protein comprises the mutations E17K, E58K, E52K, E53K, E55K, and E65K , relative to the sequence of SEQ ID NO: 569. In some cases, the chimeric protein comprises the sequence of SEQ ID NO: 612.
- the chimeric protein further comprises one or more nuclear export sequences.
- the one or more nuclear export sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 5.
- the chimeric protein further comprises one or more nuclear localization sequences.
- the one or more nuclear localization sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 6.
- the Pleckstrin Homology domain is coupled to the protein payload. In some cases, the Pleckstrin Homology domain is reversibly coupled to the protein payload. In some cases, the protein payload is reversibly coupled to the Pleckstrin Homology domain by a cleavable linker. In some cases, the cleavable linker is cleavable by a protease. In some cases, the protein payload is coupled to C-terminal of the Pleckstrin Homology domain. In some cases, the protein payload is coupled to N-terminal of the Pleckstrin Homology domain.
- the protein payload comprises one or more viral proteins.
- the one or more viral proteins comprise one or more retroviral proteins.
- the one or more retroviral proteins comprises a retroviral Gag protein.
- the protein payload comprises one or more non-viral proteins.
- the one or more non-viral proteins comprises one or more mammalian proteins.
- the one or more mammalian proteins comprises an Arc protein.
- the protein payload comprises a gene editing protein.
- the gene editing protein comprises a prime editing protein.
- the prime editing protein is coupled to a target-specific prime editing guide RNA.
- the gene editing protein comprises a CRISPR system.
- the CRISPR system comprises a Cas domain.
- the gene editing protein is coupled to a target-specific guide RNA.
- the protein payload comprises an epigenetic editing protein.
- the epigenetic editing protein is coupled to a target-specific guide RNA.
- the protein payload comprises a recombinase protein.
- the protein payload comprises an integrase protein.
- nucleic acid molecule encoding a chimeric protein.
- a lipid delivery particle comprising a chimeric protein.
- the lipid delivery particle further comprises an envelope, wherein the envelope comprises a lipid bilayer encasing a lumen, wherein the chimeric protein is located in the lumen.
- the lipid delivery particle further comprises a target ligand.
- the target ligand reversibly couples to a target receptor.
- the target receptor comprises a cell surface protein on a target cell.
- the envelope couples to the target cell through coupling of the target ligand and the target receptor on the target cell.
- coupling of the target ligand and the target receptor on the target cell induces release of the protein payload in the target cell.
- release of the protein payload in the target cell follows endocytosis of the lipid delivery particle.
- a method to deliver a payload to a target cell comprising delivering the lipid delivery particle to a target cell.
- FIG. 1 is a schematic showing nonlimiting examples of potential architecture for the chimeric protein disclosed herein.
- the black box designates the plasma membrane recruitment element
- the checkered box designates the [n * nuclear export signal]
- the black line designates the cleavable linker
- the gray box designates a payload, which can further comprise a nuclear localization sequence.
- FIGs. 2A-2D are graphs depicting the editing percentage in HEK293T and K5662 cells using Cas9 genetic editors fused to various PH domains and encapsulated in lipid delivery particles at high and low doses.
- FIG. 1 is a schematic showing nonlimiting examples of potential architecture for the chimeric protein disclosed herein.
- the black box designates the plasma membrane recruitment element
- the checkered box designates the [n * nuclear export signal]
- the black line designates the cleavable linker
- the gray box designates a payload, which can further comprise a nuclear localization sequence.
- FIGs. 2A-2D are graphs
- FIG. 2A is a graph depicting editing percentages in HEK293T cells treated with a high dose of lipid delivery particle encapsulated with Cas9 genetic editors fused to various PH domains.
- FIG. 2B is a graph depicting editing percentages in HEK293T cells treated with a low dose of lipid delivery particle encapsulated with Cas9 genetic editors fused to various PH domains.
- FIG. 2C is a graph depicting editing percentages in K562 cells treated with a high dose of lipid delivery particle encapsulated with Cas9 genetic editors fused to various PH domains.
- FIG. 2D is a graph depicting editing percentages in K562 cells treated with a low dose of lipid delivery particle encapsulated with Cas9 genetic editors fused to various PH domains.
- the present disclosure relates to the use of Pleckstrin Homology (PH) domains for delivery of protein payloads to target cells.
- PH domains are used to facilitate recruitment and packaging of a payload in a lipid delivery particle.
- PH domains have a range of attributes and functions that can be modified for particular applications.
- the PH domain can be selected or engineered to for one or more improved attributes and functions for a particular application.
- the plasma membrane recruitment element is modified for more efficient recruitment of protein payloads into a lipid delivery particle.
- the plasma membrane recruitment element is modified so that the total number of active protein payload that can be encapsulated in a lipid delivery particle is increased.
- the PH domain can be modified for more efficient release of the protein payload from a lipid delivery particle into a target cell.
- the plasma membrane recruitment element is modified for more efficient trafficking of the protein payload to a particular target cell.
- the plasma membrane recruitment element can be modified for more efficient trafficking of the protein payload to a particular subcellular locale in the target cell.
- the plasma membrane recruitment element is modified for suitability for combination with other proteins, protein domains, protein components, or a combination thereof.
- chimeric proteins comprising engineered PH domains.
- chimeric proteins comprising PH domains coupled to payloads.
- nucleic acid molecules encoding the chimeric proteins and protein payloads.
- lipid delivery particles comprising the chimeric proteins with PH domains.
- provided herein also includes cells comprising nucleic acid molecules encoding the chimeric proteins coupled to protein payloads.
- cells comprising the chimeric proteins.
- methods of making lipid delivery vehicles comprising chimeric proteins comprising PH domains coupled to protein payloads.
- kits comprising nucleic acid molecules encoding chimeric proteins and protein payloads.
- the lipid delivery particle provided herein comprises an envelope protein.
- the envelope protein can be associated with the outside boundary or the surface of the lipid delivery particle, for example, the membrane or envelope of the lipid delivery particle.
- the membrane of the lipid delivery particle can comprise a lipid layer, such as a single layer or a lipid bilayer.
- the membrane of the lipid delivery particle is from plasma membrane, endoplasmic reticulum, or a combination thereof.
- the membrane of the lipid delivery particle is from Golgi complex, ER Golgi intermediate compartment, or nuclear envelope.
- the membrane of the lipid delivery particle is from plasma membrane.
- the membrane of the lipid delivery particle is a phospholipid bilayer.
- the envelope protein can be associated with the membrane of the lipid delivery particle in various manners.
- the envelope protein can be anchored or attached to the external membrane of the particle or anchored or attached to the internal membrane of the particle.
- the envelope protein can be embedded or inserted in the membrane, spanning through the membrane, with certain portions located at the outside of the membrane, or certain portions extending to the inside of the particle, or both.
- the envelope protein within the lipid delivery particle described herein can be overexpressed from an exogenous source, such as plasmids or stably integrated transgenes, in the production cells.
- the envelope protein can play a role in the delivery of the lipid delivery particle to a target cell and release of the components of the lipid delivery particle within the target cell.
- the envelope protein can contact with the surface of a target cell and participate in the fusion of the lipid delivery particle and the membrane of the target cell.
- the envelope protein can participate in the fusion of the lipid delivery particle with the membrane of the target cell via any appropriate mechanism, such as those described in White et al. Crit Rev Biochem Mol Biol. 2008; 43(3): 189-219.
- One example of the fusion mechanisms is unifying Trimer-of-Hairpins Fusion Mechanism.
- Membrane fusion can occur after allosteric priming by binding to a target receptor. In some cases, membrane fusion occurs after proteolysis. In some cases, membrane fusion occurs after isomerization of disulfide bridges.
- membrane fusion occurs by internalization and then priming of fusion via (i) cathepsin-mediated proteolysis, or (ii) low pH/acidification.
- the cathepsin-mediated proteolysis can be pH dependent or pH independent.
- Other fusion triggering mechanisms can include low PH, binding to target cell receptors, and a receptor followed by low pH.
- the envelope protein can also play a role in the formation of the lipid delivery particle.
- the envelope protein can interact with another component within the lipid delivery particle and participate in the assembly of the lipid delivery particle, for example, in a producer cell.
- the envelope protein can make contact with another envelope protein and form an oligomer embedded within the membrane.
- the envelope protein can be a glycoprotein, for example, a transmembrane glycoprotein.
- envelope protein comprises multiple membrane -spanning regions. These multiple membranespanning regions can oligomerize and form channels in the membrane.
- the envelope protein is fused with a targeting moiety.
- the targeting moiety recognizes a specific molecule (e.g, antigen, receptor, or other membrane protein) on the surface of a target cell to allow targeted cell entry with more specificity.
- the targeting moiety is specific for a certain cell type or is specific for a certain target cell.
- the targeting moiety can be fused to the envelope protein at a position that is located at an outside of the lipid delivery particle.
- the targeting moiety includes scFvs, antibody variable regions, nanobodies, T-cell receptor variable regions, other antigen-binding fragments or their mimetics, such as DARPins.
- the targeting moiety is a protein ligand from the human ligandome.
- the targeting moiety can be a natural peptide or a synthetic peptide.
- the targeting moiety is not fused with the envelope protein and is attached to the membrane of the lipid delivery particle from the outside, for example, via a transmembrane domain.
- a targeting moiety can include, e.g., an antibody or an antigen-binding fragment thereof (e.g., Fab, Fab', F(ab')2, Fv fragments, scFv antibody fragments, disulfide-linked Fvs (sdFv), a Fd fragment consisting of the VH and CHI domains, linear antibodies, single domain antibodies such as sdAb (either VL or VH), nanobodies, or camelid VHH domains), an antigen-binding fibronectin type III (Fn3) scaffold such as a fibronectin polypeptide minibody, a ligand, a cytokine, a chemokine, or a T cell receptor (TCRs).
- an antibody or an antigen-binding fragment thereof e.g., Fab, Fab', F(ab')2, Fv fragments, scFv antibody fragments, disulfide-linked Fvs (sdFv), a Fd fragment consisting
- Membrane-fusion proteins can be re-targeted by non-covalently conjugating a targeting moiety to the membrane-fusion protein or targeting protein (e.g. the hemagglutinin protein).
- the membrane -fusion protein can be engineered to bind the Fc region of an antibody that targets an antigen on a target cell, redirecting the membrane fusion activity towards cells that display the antibody’s target.
- the targeting moiety linked to the membrane -fusion protein binds a cell surface marker on the target cell, e.g, a protein, glycoprotein, receptor, cell surface ligand, agonist, lipid, sugar, class I transmembrane protein, class II transmembrane protein, or class III transmembrane protein.
- the lipid delivery particles disclosed herein display targeting moieties that are not conjugated to the membrane -fusion protein or other proteins in order to redirect the fusion activity of the lipid delivery particle towards a cell that is bound by the targeting moiety, or to affect tropism of the lipid delivery particle toward the target cell.
- an envelope protein has a viral origin.
- a suitable envelope protein is from a DNA virus, an RNA virus, or a retrovirus.
- the envelope protein can be envelope protein from Herpesviruses, Avian sarcoma leukosis virus, Poxviruses, Hepadnaviruses, Asfarviridae, Flaviviruses, Alphaviruses, Togaviruses, Coronaviruses, Hepatitis D, Orthomyxoviruses, Rhabdovirus, Bunyaviruses, Filoviruses, Oncoretroviruses, lentiviruses, Spumaviruses.
- envelope protein can be envelope protein from lentiviruses, for example, human immunodeficiency virus (HIV), simian immunodeficiency virus (SIV), feline immunodeficiency virus (FIV) and equine infectious anemia virus (EIAV).
- HIV human immunodeficiency virus
- SIV simian immunodeficiency virus
- FV feline immunodeficiency virus
- EIAV equine infectious anemia virus
- an envelope protein is a fusion of two different envelope proteins, wherein each comes from a different virus. Additional suitable envelope proteins that are from viral origins and their functions are described in White JM et al., Crit Rev Biochem Mol Biol. 2008 May-Jun;43(3): 189-219.
- the envelope protein is a vesicular stomatitis virus glycoprotein (VSVG) or a biologically active mutant thereof.
- VSVG vesicular stomatitis virus glycoprotein
- a “biologically active mutant” disclosed herein in connection with a reference protein can refer to a mutant of the reference protein that remains displaying one or more biological activities that are of same nature as the reference protein, which are relevant to the context in which the reference protein is used in the lipid delivery particle disclosed herein, while the level of the one or more biological activities of the biologically active mutant can be either similar as or different than the reference protein.
- the biologically active mutant of a VSVG in the context of an envelope protein remains displaying the biological activities of an envelope protein, e.g., mediating membrane fusion, tropism of the lipid delivery particle toward a target cell, or both.
- a mutant as described in the present disclosure is equivalent to a biologically active mutant.
- the envelope protein is a Human immunodeficiency virus GP160 or a biologically active mutant thereof.
- the envelope protein is a Baboon Endogenous Retrovirus (BaEVTR) glycoprotein or a biologically active mutant thereof.
- the envelope protein is a modified Baboon Endogenous Retrovirus (BaEVTRless) glycoprotein or a biologically active mutant thereof.
- the envelope protein is the fusion protein of Vesicular stomatitis Indiana virus and Rabies virus Glycoproteins (FuG-E) or a biologically active mutant thereof.
- the envelope protein pantropic murine leukemia virus envelope protein (MLV) or a biologically active mutant thereof.
- the envelope protein is a modified Fusion protein of Vesicular stomatitis Indiana virus and Rabies virus Glycoproteins (FuG-E P440E) or a biologically active mutant thereof.
- the envelope protein is an ecotropic Murine Leukemia Virus envelope protein (MLV ENV ecotropic) or a biologically active mutant thereof.
- the envelope protein is an amphotrophic Murine Leukemia Virus envelope protein (MLV ENV amphotropic) or a biologically active mutant thereof.
- the envelope protein is a Moloney murine leukemia virus envelope protein (MMLV) or a biologically active mutant thereof.
- the envelope protein is a Moloney murine sarcoma virus envelope protein (MoMSVg) or a biologically active mutant thereof.
- the envelope protein is a moloney murine leukemia virus 10A1 strain Glycoprotein (MLV 10A1) or a biologically active mutant thereof.
- the envelope protein is a xenotropic murine leukemia virus envelope protein (MLV ENV xenotropic) or a biologically active mutant thereof.
- the envelope protein is a xenotropic murine leukemia virus-related envelope protein (XMRV) or a biologically active mutant thereof.
- the envelope protein is a Baculovirus envelope glycoprotein (GP64) or a biologically active mutant thereof. In some cases, the envelope protein is an endogenous feline virus envelope protein (RD114 ENV) or a biologically active mutant thereof. In some cases, the envelope protein is a mammalian endogenous retrovirus protein, or a biologically active mutant thereof.
- the mammalian endogenous retrovirus protein can be a koala retrovirus protein (KoRV) or a Jaagsiekte sheep retrovirus protein (enJSRV), or a biologically active mutant thereof.
- the envelope protein is a simian endogenous type D retrovirus protein (RD-114) or a biologically active mutant thereof.
- the envelope protein is a gibbon ape leukemia virus envelope protein (GALV) or a biologically active mutant thereof.
- the envelope protein is a feline leukemia virus envelope protein (FLV) or a biologically active mutant thereof.
- the envelope protein is a mouse mammary tumor virus envelope protein (MMTV) or a biologically active mutant thereof.
- the envelope protein is an avian leukosis virus envelope protein or a biologically active mutant thereof.
- the envelope protein is a rous sarcoma virus envelope protein or a biologically active mutant thereof.
- the envelope protein can direct the lipid delivery particles to fuse with a certain type of target cells rather than other cells.
- the lipid delivery particle can preferentially target different cell types (z.e., tropisms of the lipid delivery particles), such as liver cells, ocular cells, nerve cells, lung cells, immune cells, muscle cells, and any other cell types of interest.
- the envelope protein can be an Ebola virus glycoprotein or a biologically active mutant thereof, a Marburg virus glycoprotein or a biologically active mutant thereof, or a VSV-G or a biologically active mutant thereof.
- a target immune cell for example, CD8+ T cell, an HTLV-1 glycoprotein or a biologically active mutant thereof, or a VSV- G glycoprotein or a biologically active mutant thereof.
- the envelope protein can be a HIV-1 envelope or a biologically active mutant thereof, a HTLV-1 glycoprotein or a biologically active mutant thereof, or a VSV-G glycoprotein or a biologically active mutant thereof.
- the envelope protein can be a respiratory syncytial virus glycoprotein or a biologically active mutant thereof, or a SARS-CoV glycoprotein or a biologically active mutant thereof.
- the envelope protein can be a rabies glycoprotein or a biologically active mutant thereof, a Mokola virus glycoprotein or a biologically active mutant thereof, a Semliki Forest virus glycoprotein or a biologically active mutant thereof, a Venezuelan equine encephalitis virus glycoprotein or a biologically active mutant thereof, or a VSV-G or a biologically active mutant thereof.
- the envelope protein can be an Ebola virus glycoprotein or a biologically active mutant thereof, a Marburg virus glycoprotein or a biologically active mutant thereof, or a VSV-G or a biologically active mutant thereof.
- the envelope protein comprises the sequences set forth in Table 1.
- the envelope protein comprises the sequences set forth in Table 1 with at least one amino acid substitution, deletion, or insertion.
- N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein.
- the envelope protein comprises the sequences set forth in Table 1 and a heterologous peptide sequence fused to the N-terminal or C-terminal.
- the envelope protein comprises one or more of the sequences set forth in Table 1 with at least one amino acid substitution, deletion, or insertion.
- N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein.
- the envelope protein comprises one or more of the sequences set forth in Table 1 and a heterologous peptide sequence fused to the N-terminal or C-terminal.
- the envelope protein comprises any one of the sequences set forth in Table 1 with at least one amino acid substitution, deletion, or insertion.
- N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein.
- the envelope protein comprises any one of the sequences set forth in Table 1 and a heterologous peptide sequence fused to the N-terminal or C-terminal.
- the envelope protein comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in Table 1.
- the envelope protein comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104.
- the envelope protein comprises an amino acid sequence that has at least about 50% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 60% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104 In some cases, the envelope protein comprises an amino acid sequence that has at least about 70% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 75% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104.
- the envelope protein comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104.
- the envelope protein comprises an amino acid sequence that has at least about 80% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104
- the envelope protein comprises an amino acid sequence that has at least about 85% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104.
- the envelope protein comprises an amino acid sequence that has at least about 90% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 95% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104 In some cases, the envelope protein comprises an amino acid sequence that has at least about 96% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 97% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104.
- the envelope protein comprises an amino acid sequence that has at least about 98% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 99% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104.
- the envelope protein in the lipid delivery particle described herein has a human origin, e.g., has significant sequence similarity to a human wild-type protein, such as at least 90%, at least 95%, at least 98%, or at least 99%.
- Using an envelope protein of a human origin can have benefits such as providing a minimized immunogenicity and better tolerance in a human subject receiving the lipid delivery particles.
- the lipid delivery particle comprising an envelope protein of a human origin can comprise another component that is from human origin or from non-human origin (e.g. , a payload or a plasma membrane recruitment element).
- An envelope protein that is from human origin can include, example, envelope proteins or glycoproteins of human endogenous retroviruses (HERVs), other human endogenous envelope proteins, or other human endogenous proteins that serve a similar function of recognizing and/or fusing with membrane of a target cell (e.g., clathrin adaptor protein complex- 1, CHMP4C, Proteolipid protein 1, TSAP6, immunoglobulin variable domains, or a biologically active mutant thereof).
- the envelope protein is a HERV envelope protein such as any one of those listed in Table 2.
- the envelope protein is a hENVHl or a biologically active mutant thereof.
- the envelope protein is a hENVH2 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVH3 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVKl or a biologically active mutant thereof. In some cases, the envelope protein is a hENVK2 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVK3 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVK4 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVK5 or a biologically active mutant thereof.
- the envelope protein is a hENVK6 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVT or a biologically active mutant thereof. In some cases, the envelope protein is a hENVW or a biologically active mutant thereof. In some cases, the envelope protein is a hENVFRD or a biologically active mutant thereof. In some cases, the envelope protein is a hENVR or a biologically active mutant thereof. In some cases, the envelope protein is a hENVR(b) or a biologically active mutant thereof. In some cases, the envelope protein is a hENVR(c)2 or a biologically active mutant thereof.
- the envelope protein is a hENVR(c) 1 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVKcon or a biologically active mutant thereof. In some cases, the envelope protein is a truncated HERV protein.
- the envelope protein comprises the sequences set forth in Table 3.
- the envelope protein comprises the sequences set forth in Table 3 with at least one amino acid substitution, deletion, or insertion.
- the N-terminal methionine can be absent.
- the envelope protein comprises the sequences set forth in Table 3 and a heterologous peptide sequence fused to the N-terminal or C-terminal.
- the envelope protein comprises one or more of the sequences set forth in Table 3 with at least one amino acid substitution, deletion, or insertion.
- N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein.
- the envelope protein comprises one or more of the sequences set forth in Table 3 and a heterologous peptide sequence fused to the N-terminal or C-terminal.
- the envelope protein comprises any one of the sequences set forth in Table 3 with at least one amino acid substitution, deletion, or insertion.
- N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein.
- the envelope protein comprises any one of the sequences set forth in Table 3 and a heterologous peptide sequence fused to the N-terminal or C-terminal.
- the envelope protein comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 50% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 60% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82.
- the envelope protein comprises an amino acid sequence that has at least about 70% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 75% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82.
- the envelope protein comprises an amino acid sequence that has at least about 80% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 85% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 90% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 95% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82.
- the envelope protein comprises an amino acid sequence that has at least about 96% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 97% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82.In some cases, the envelope protein comprises an amino acid sequence that has at least about 98% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 99% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. Table 3. Exemplary sequences for human HERV envelope proteins PLASMA MEMBRANE RECRUITMENT ELEMENT
- the lipid delivery particle provided herein comprises a plasma membrane recruitment element.
- the lipid delivery particle disclosed herein can comprise a membrane.
- the membrane encapsulates a payload.
- the lipid delivery particle comprises a plasma membrane recruitment element, for example, inside the cavity of the lipid delivery particle.
- the plasma membrane recruitment element can localize itself to the membrane of the lipid delivery particles.
- the plasma membrane recruitment element can be utilized to recruit a component (e.g., a payload) to the membrane of the lipid delivery particles via forming a chimeric protein of the plasma membrane recruitment element and a component to be localized to the membrane or other mechanisms of attachment.
- the membrane encapsulates a protein core.
- At least a portion of the plasma membrane recruitment element forms the basic structure of the lipid delivery particle, such as a portion of the protein core inside the lipid delivery particle. In some cases, at least a portion of the plasma membrane recruitment element binds to the membrane of the lipid delivery particle from the inside.
- the plasma membrane recruitment element can play a role in the assembly of the lipid delivery particle, such as packing various components (e.g., a payload) into the lipid delivery particles.
- the plasma membrane recruitment element can direct budding of the lipid delivery particles from a producer cell.
- expressing plasma membrane recruitment element alone or together with an envelope protein disclosed herein in a producer cell can lead to formation of the lipid delivery particle.
- the plasma membrane recruitment element has a viral origin.
- the plasma membrane recruitment element comprises a retroviral gag protein, e.g., a retroviral polyprotein that comprises one or more of a matrix (MA) polypeptide, an RNA-binding phosphoprotein polypeptide, a capsid (CA) polypeptide, or a nucleocapsid (NC) polypeptide.
- the plasma membrane recruitment element can comprise HIV gag or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a gag from murine leukemia virus (MLV) or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a gag from Moloney murine leukemia virus (MMLV) or a biologically active mutant thereof.
- the plasma membrane recruitment element forms structural protein that forms the protein core of the lipid delivery particles described herein.
- the plasma membrane recruitment element can comprise Respiratory syncytial virus (RSV) M or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Human Papillomavirus (HPV) LI protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise HPV L2 protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Hepatitis B virus (HBV) core protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Hepatitis C virus (HCV) core protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise hepatitis E virus (HeV) M protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Chikungunya virus (CHIKV) C-E3-E2-6k-El or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise RSV NP or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Human metapneumovirus (HMPV) M or a biologically active mutant thereof.
- the plasma membrane can comprise a glycoprotein from a flavivirus.
- the flavivirus can comprise Chikungunya virus, Zika virus, Dengue virus, or West Niles virus.
- the plasma membrane recruitment element can comprise Zika virus (ZIKV) C or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise ZIKV prM/M or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Dengaue virus (DENV) C-prM or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise West Nile Virus (WNV) prME protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise WNV CprME protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Filovirus VP40 or Z protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Baculovirus Pl protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Rotavirus VP7 or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Rotavirus VP2 protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Rotavirus VP6 protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Porcine Circovirus Type 2 (PCV2) capsid or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise baculovirus VP2 protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise baculovirus VP5 protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise baculovirus VP3 protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise or baculovirus VP7 protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Ebola nucleocapsid or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Parovirus VP1 protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Parovirus VP2 protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Newcastle disease virus (NDV) M protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Human polyomavirus 2 (JCPyV) VP1 protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise Human parainfluenza virus type 3 (HPIV3) M protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise HPIV3N protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise or Mumps virus (MuV) M proteins or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise SARS M protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise SARS E protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise SARS N protein or a biologically active mutant thereof.
- the plasma membrane recruitment element is a mammalian protein or part thereof.
- the plasma membrane recruitment element can include a pleckstrin homology (PH) domain or a transmembrane domain of a mammalian protein, such as a mouse protein or a human protein.
- the plasma membrane recruitment element has a human origin.
- the plasma membrane recruitment element can include a gag from human endogenous retrovirus, such as Human Endogenous Retrovirus K (e.g., HERV- K113, HERV-K101, HERV-K102, HERV-K104, HERV-K107, HERV-K108, HERV-K109, HERV- K115, HERV- KI lp22, and HERV-K12ql3) and Human Endogenous Retrovirus-W (HERV-W) or a biologically active mutant thereof.
- the plasma membrane recruitment element can include a hGAGK con or a biologically active mutant thereof.
- the plasma membrane recruitment element can include an endogenous gag of a mammal (e.g, human) from retrotransposons (e.g., Arc from vertebrate lineage of Ty3/gypsy retrotransposon), which are also ancestral to retroviruses.
- the plasma membrane recruitment element comprises a portion from human Arc.
- the plasma membrane recruitment element can include a pleckstrin homology (PH) domain from a mammalian protein or a biologically active mutant thereof.
- the plasma membrane recruitment element can include a pleckstrin homology (PH) domain from a human protein or a biologically active mutant thereof.
- the PH domains can play a role in protein-membrane interactions via binding to phosphatidylinositol phosphate (PIP), for example PIP2 or PIP3, or other lipids or proteins within the membrane of the lipid delivery particles.
- PIP phosphatidylinositol phosphate
- PH domains with different sequences can have varied affinities and selectivity when binding different PIPs.
- the plasma membrane recruitment element can include a pleckstrin homology (PH) domain from a human protein or a mutant thereof.
- the PH domains can play a role in protein-membrane interactions via binding to phosphatidylinositol phosphate (PIP), for example PIP2 or PIP3, or other lipids or proteins within the membrane of the lipid delivery particles.
- PIP phosphatidylinositol phosphate
- the plasma membrane recruitment element interacts with one or more anionic lipids within the membrane of the lipid delivery particles.
- the plasma membrane recruitment element interacts with one or more monophosphorylated derivatives of phosphatidylinositol.
- the plasma membrane recruitment element interacts with one or more bisphosphorylated derivatives of phosphatidylinositol. In some cases, the plasma membrane recruitment element interacts with one or more trisphosphorylated derivatives of phosphatidylinositol. In some cases, the plasma membrane recruitment element interacts with one or more phosphatidylinositol 4,5-bisphosphate (PI(4,5)P2) molecules or phosphatidylinositol 3,5-bisphosphate (PI(3,5)P2) molecules within a lipid layer. In some cases, the plasma membrane recruitment element interacts with one or more phosphatidylinositol(3,4,5)- trisphosphate (PIP3) molecules within a lipid layer. PH domains with different sequences can have varied affinities and selectivity when binding different PIPs.
- the plasma membrane recruitment element interacts with one or more proteins within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more G-protein coupled receptors (GPRC) within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more bg-subunits of GPRC within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more kinase proteins within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more protein kinase C (PKC) proteins within the membrane of the lipid delivery particles.
- GPRC G-protein coupled receptors
- PLC protein kinase C
- the plasma membrane recruitment element interacts with one or more members of an Arf family small G- proteins within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more GDP-bound form of ADP-ribosylation factor (ARF) within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with other proteins, protein domains, protein components, or a combination thereof within or tethered to the membrane of the lipid delivery particles.
- ADP-ribosylation factor ADP-ribosylation factor
- the PH domain is selected or engineered for enhanced or altered properties.
- the plasma membrane recruitment element comprises an enhancement or alteration of the recruitment of a payload into a lipid delivery particle, the amount of a payload that can be encapsulated in a lipid delivery particle, release of a payload from the lipid delivery particle into a target cell, trafficking of a lipid delivery particle to a subcellular locale in a target cell, suitability of the PH for combination with other protein domains, or a combination thereof.
- the plasma membrane recruitment element is engineered to enhance or alter the delivery of a payload to a target cell or target cellular compartment.
- the plasma membrane recruitment element is engineered to enhance or alter the delivery of protein payload to a target cell or target cellular compartment. In some cases, the plasma membrane recruitment element is engineered to enhance or alter the delivery of a homogenous protein payload to a target cell or target cellular compartment. In some cases, the plasma membrane recruitment element is engineered to enhance or alter the delivery of a heterogenous protein payload to a target cell or target cellular compartment. In some cases, the plasma membrane recruitment element is engineered to enhance, alter, or decrease other properties and/or functions of the PH domain.
- the plasma membrane recruitment element is engineered to comprise mutations in particular structurally and functionally important regions of the PH domain.
- PH domain comprises one or more mutations that affect interaction and/or binding between the PH domains and a lipid.
- the plasma membrane recruitment element comprises one or more mutations that affect the lipid binding surface.
- the plasma membrane recruitment element comprises one or more mutations that affect interaction and/or binding between the PH domains and a phosphatidylinositol or derivative thereof.
- the plasma membrane recruitment element comprises one or more mutations that affect the binding surface for phosphatidylinositol or derivative thereof.
- the plasma membrane recruitment element comprises one or more mutations that affect interaction between the PH domains and a protein. In some cases, the plasma membrane recruitment element comprises one or more mutations that affect the protein-interacting surfaces. In some cases, the plasma membrane recruitment element comprises one or more mutations that affect the other structural and functional features of the PH domains, nonlimiting examples of which can include tertiary structure, hydrophobicity, a helix architecture and configuration, [3-barrel architecture and configuration, electrostatic interactions, hydrogen-bonding, dimerization, oligomerization, any other structural and/or functional features, of a combination thereof. In some cases, the plasma membrane recruitment elements can be screened for particular functional properties. In some cases, the plasma membrane recruitment elements can be screened for particular functional properties in a high throughput manner.
- the PH is selected from the genera: Homo, Mus, Rattus, Bos, Sus, Gallus, Danio, Canis, Xenopus, Macaca, Pan, Felis, Pongo, Dasypus, Anser, Cavia, Capra, Ovis, Meriones, Mustela, Columba, Mauremys, Callorhinchus, Pteropus, Cricetulus, Scophthalmus, Protopterus, Lates, Thunnus, Callithrix, Equus, Mesocricetus, Oryzias, Mya, Polypterus, Gallus, Drosophila, and other genera belonging to the Chordata phylum.
- the plasma membrane recruitment element is selected from birds, turtles, alligators, lizards, snakes, mammals, amphibians, lungfishes, bony fishes, cartilaginous fishes, othered jawed vertebrates, or a combination thereof.
- the plasma membrane recruitment element is selected from human proteins.
- the plasma membrane recruitment element can include a PH domain of phospholipase C51 (e.g., human phospholipase C51) or a biologically active mutant thereof.
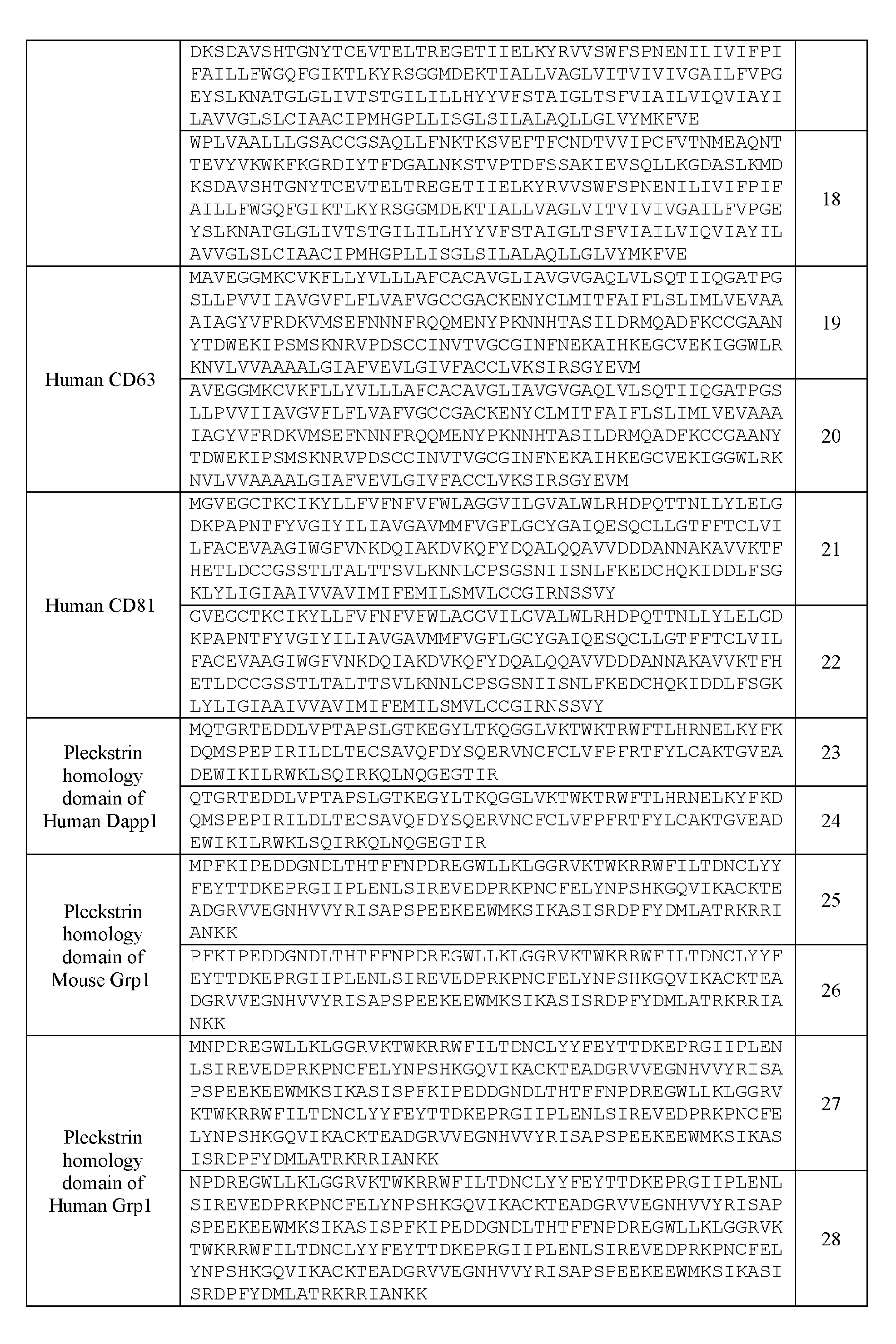
- the plasma membrane recruitment element can comprise a PH domain of Aktl (e.g, human Aktl) or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a mutant PH domain of human Aktl with E17K substitution or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a PH domain of 3 -phosphoinositide-dependent protein kinase 1 (e.g., human 3- phosphoinositide-dependent protein kinase 1) or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a PH domain of Dappl (e.g., human Dappl) or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a PH domain of Grp 1 (e.g. , mouse Grp 1) or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a PH domain of human Grpl or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a PH domain of OSBP (e.g., human OSBP) or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a PH domain of Btkl (e.g. , human Btkl) or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a PH domain of FAPP1 (e.g., human FAPP1) or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a PH domain of CERT (e.g. , human CERT) or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a PH domain of PKD (e.g. , human PKD) or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a PH domain of PHLPP1 (e.g., human PHLPP1) or a biologically active mutant thereof.
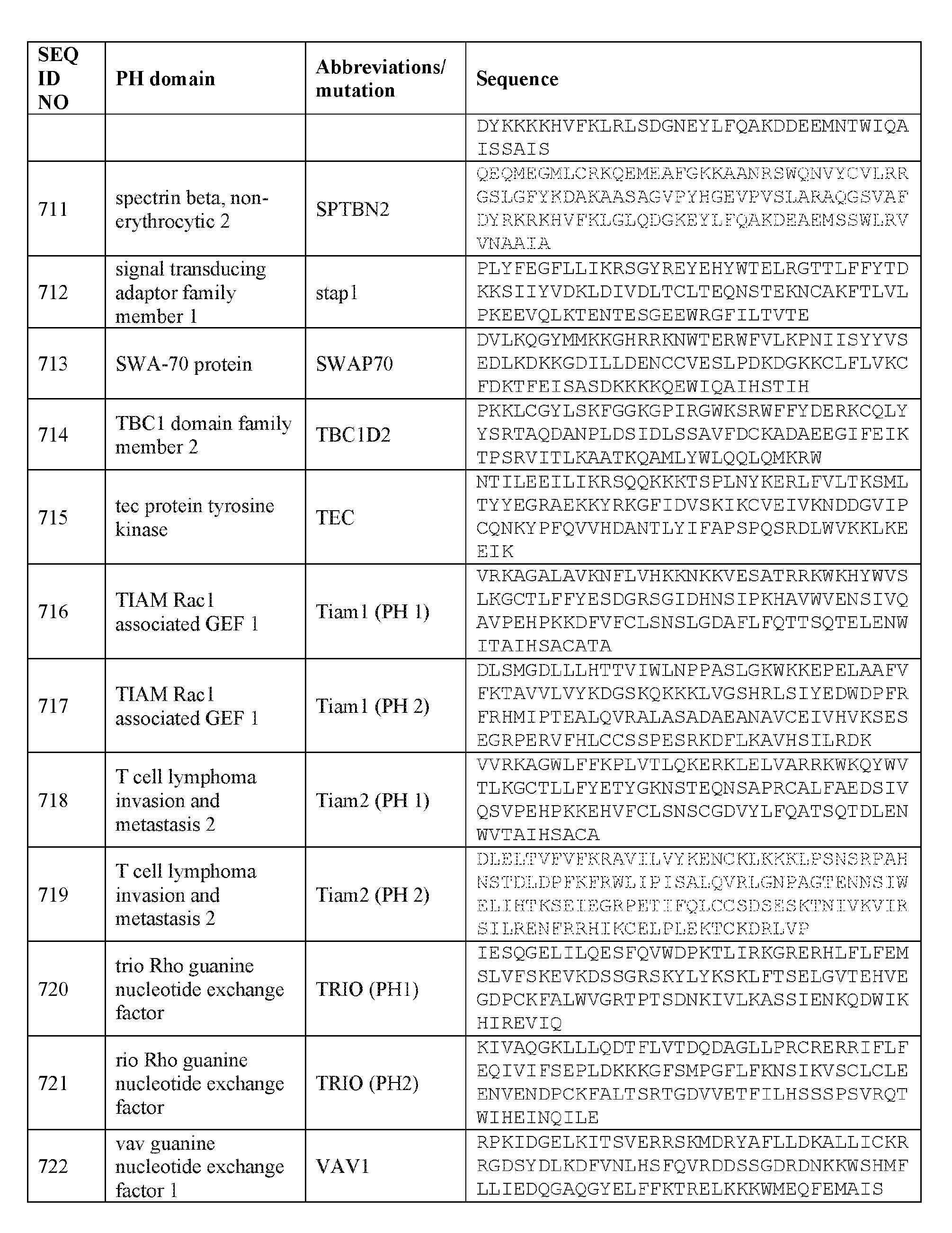
- the plasma membrane recruitment element can comprise a PH domain of SWAP70 (e.g., human SWAP70) or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a PH domain of MAPKAP1 (e.g., human MAPKAP1) or a biologically active mutant thereof.
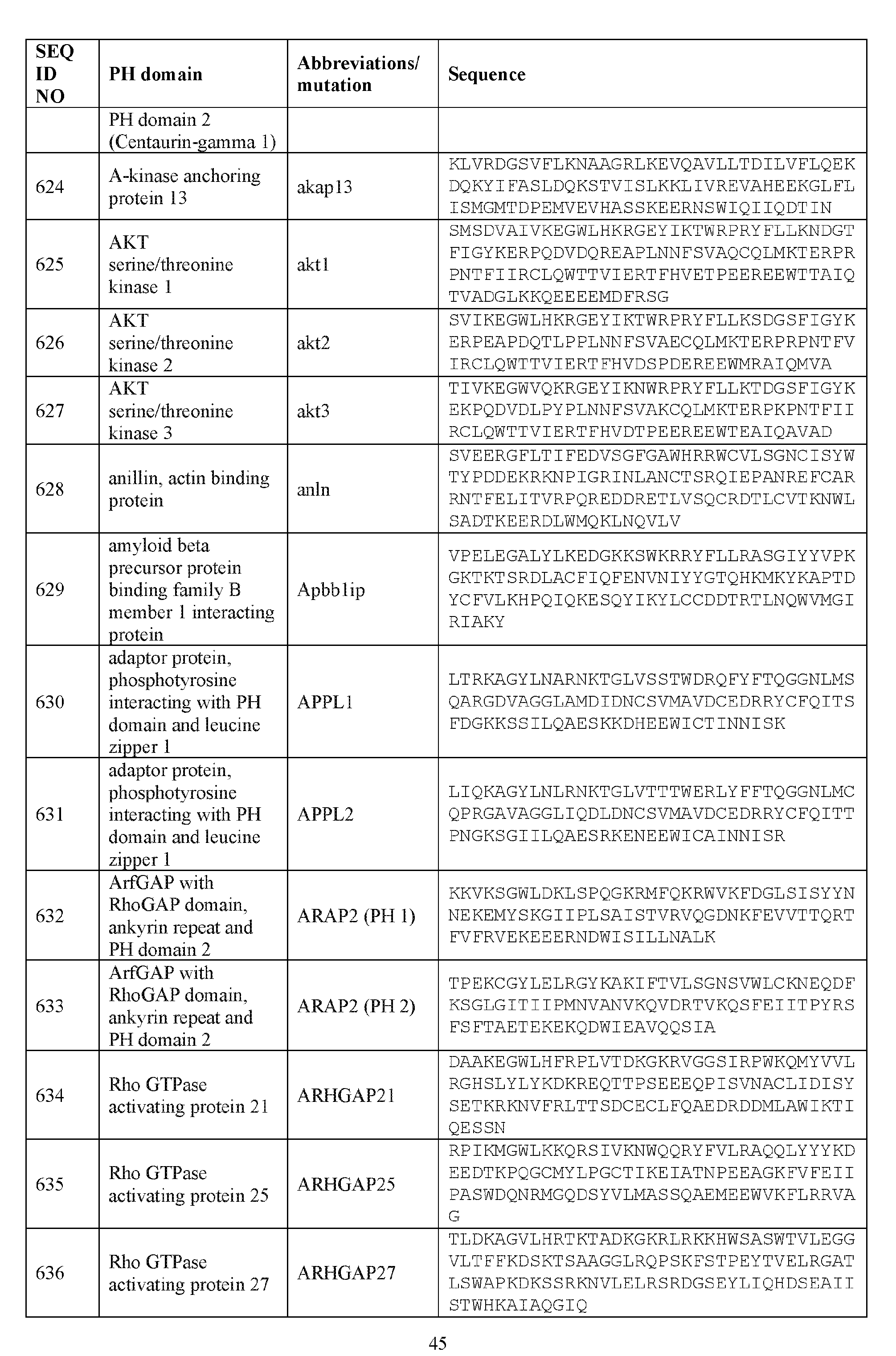
- the plasma membrane recruitment element comprises a PH domain of SH3 domain binding protein 2 (3BP2), Rho guanine nucleotide exchange factor 28-like (AKP13), Oxysterol binding protein like 7 (OSBPL7), FYVE, RhoGEF And PH Domain Containing 6 (FGD6), Bruton agammaglobulinemia tyrosine kinase (BTK), connector enhancer of kinase suppressor of Ras 2 (CNKSR2), Rho associated coiled-coil containing protein kinase 1 (Rock 1), Rho associated coiled-coil containing protein kinase 2 (Rock 2), ceramide kinase like (CERKL), ceramide kinase (CERK), SAPK- interacting protein 1 (SIN1), pleckstrin homology like domain family A member 2 (PHLDA2), tubby bipartite transcription factor (TUB), PH domain and leucine rich
- AFAP1L2 ArfGAP with GTPase domain, ankyrin repeat and PH domain 2
- AGAP2 ArfGAP with GTPase domain, ankyrin repeat and PH domain 2
- Akapl3 A-kinase anchoring protein 13
- AKT2 AKT serine/threonine kinase 2
- AKT serine/threonine kinase AKT serine/threonine kinase
- AKT3 anillin, actin binding protein (anln), amyloid beta precursor protein binding family B member
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to the sequence of SEQ ID NO: 569.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to the sequence of SEQ ID NO: 569.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to the sequence set forth in any one of SEQ ID NOs: 570-596.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to the sequence set forth in any one of SEQ ID NOs: 570-596.
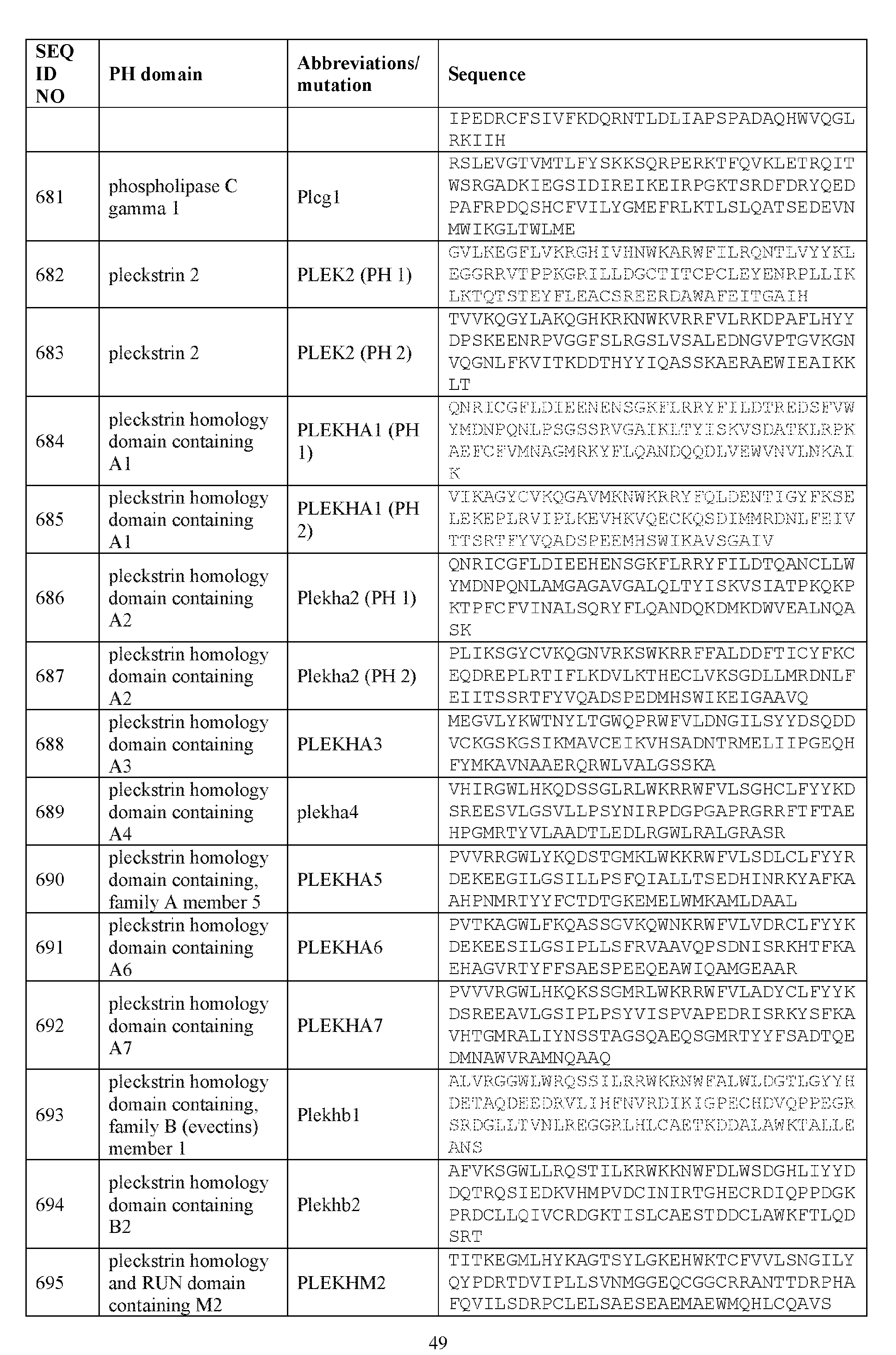
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to any one of the sequences listed in Table 4A.
- the plasma membrane recruitment element comprises any one of the sequences listed in Table 4A without the N- terminal Methionine. In some cases, the plasma membrane recruitment element comprises the sequence set forth in any one of SEQ ID NOs: 569-596 without the N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises any one of the sequences listed in Table 4A with the N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises the sequence set forth in any one of SEQ ID NOs: 569-596 with the N-terminal Methionine.
- the plasma membrane recruitment element comprises any one of the sequences described in Table 4A with a further truncation on the N-terminus.
- the N-terminal methionine can be absent.
- the plasma membrane recruitment element comprises any one of the sequences described in Table 4A with atruncation (e.g., truncation of about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 12, 14, 16, 18, 20, or more amino acids) on the C-terminus.
- the plasma membrane recruitment element includes those described in Table 4A with one amino acid substitution.
- the plasma membrane recruitment element comprises any one of the sequences described in Table 4A with two or more amino acid substitutions. In some cases, the plasma membrane recruitment element comprises one or more of the sequences described in Table 4A and further comprises a heterologous peptide sequence fused to the N- terminus or C-terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to one or more of the sequences listed in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4A fused to a heterologous peptide sequence on the N terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to one or more of the sequences listed in Table 4A fused to a heterologous peptide sequence on the N terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4A fused to a heterologous peptide sequence on the C terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to one or more of the sequences listed in Table 4A fused to a heterologous peptide sequence on the C terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A fused to a heterologous peptide sequence on the N terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A fused to a heterologous peptide sequence on the N terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A fused to a heterologous peptide sequence on the C terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A fused to a heterologous peptide sequence on the C terminus.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50% sequence identity to any one of the sequences set forth in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 60% sequence identity to any one of the sequences set forth in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 70% sequence identity to any one of the sequences set forth in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 75% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 85% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 90% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 95% sequence identity to any one of the sequences set forth in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 96% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 97% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 98% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 99% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has about 100% sequence identity to any one of the sequences set forth in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4A.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences set forth in SEQ ID NOs: 597-722.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to any one of the sequences set forth in SEQ ID NOs: 597-722.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, or about 99% or more sequence identity to any one of the sequences listed in Table 4B.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4B.
- the plasma membrane recruitment element comprises any one of the sequences listed in Table 4B without the N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in SEQ ID NOs: 597-722 without the N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises any one of the sequences listed in Table 4B with an N- terminal Methionine. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in SEQ ID NOs: 597-722 with an N-terminal Methionine.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences set forth in SEQ ID NOs: 597-722.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to a one or more of the sequences set forth in SEQ ID NOs: 597-722.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, or about 99% or more sequence identity to one or more of the sequences listed in Table 4B.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4B.
- the plasma membrane recruitment element comprises one or more of the sequences listed in Table 4B without the N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in SEQ ID NOs: 597-722 without the N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises one or more of the sequences listed in Table 4B with an N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in SEQ ID NOs: 597-722 with an N-terminal Methionine.
- the plasma membrane recruitment element comprises any one of the sequences described in Table 4B with a further truncation on the N-terminus.
- the plasma membrane recruitment element comprises any one of the sequences described in Table 4B with a truncation on the C-terminus.
- the plasma membrane recruitment element comprises any one of the sequences described in Table 4B with one amino acid substitution.
- the plasma membrane recruitment element comprises any one of the sequences described in Table 4B with two or more amino acid substitutions.
- the plasma membrane recruitment element comprises any one of the sequences described in Table 4B and a heterologous peptide sequence fused to the N-terminus or C-terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more sequences listed in Table 4B.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more sequences listed in Table 4B fused to a heterologous peptide sequence on the N terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4B fused to a heterologous peptide sequence on the C terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4B.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4B fused to a heterologous peptide sequence on the N terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4B fused to a heterologous peptide sequence on the C terminus.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4B.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50% sequence identity to any one of the sequences set forth in Table 4B.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 60% sequence identity to any one of the sequences set forth in Table 4B.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 70% sequence identity to any one of the sequences set forth in Table 4B.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 75% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80% sequence identity to any one of the sequences set forth in Table 4B.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 85% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 90% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 95% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 96% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 97% sequence identity to any one of the sequences set forth in Table 4B.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 98% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 99% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has about 100% sequence identity to any one of the sequences set forth in Table 4B. [0085] In some cases, the plasma membrane recruitment element is engineered to comprise one or more mutations to yield PH domain variants.
- the plasma membrane recruitment element variants comprises about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to the sequence set forth in any one of SEQ ID NOs: 569-722.
- the plasma membrane recruitment element is engineered to comprise one or more mutations to yield PH domain variants.
- the plasma membrane recruitment element variants comprises about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to the sequence set forth in one or more of SEQ ID NOs: 569-722.
- the plasma membrane recruitment element comprises the PH domain of any one of the proteins or protein fragments set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises the PH domain of one or more of the proteins or protein fragments set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a PH domain that comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to any one of the sequences set forth in any one of SEQ ID NOs: 161-290.
- the plasma membrane recruitment element comprises a PH domain that comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to one or more of the sequences set forth in any one of SEQ ID NOs: 161-290.
- the plasma membrane recruitment element comprises a PH domain that comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to any one of the sequences listed in Table 4C.
- the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to the PH domain of one or more of the sequences listed in Table 4C.
- the plasma membrane recruitment element comprises the protein or protein fragment of any one of the proteins or protein fragments set forth in Table 4C with a further truncation on the N-terminus.
- the N- terminal methionine can be absent.
- the plasma membrane recruitment element comprises the protein or protein fragment of any one of the proteins or protein fragments set forth in Table 4C with a further truncation on the C-terminus.
- the plasma membrane recruitment element comprises the protein or protein fragment of any one of the proteins or protein fragments set forth in Table 4C with one amino acid substitution.
- the plasma membrane recruitment element comprises the protein or protein fragment of any one of the proteins or protein fragments set forth in Table 4C with two or more amino acid substitutions. In some cases, the plasma membrane recruitment element comprises the protein or protein fragment of any one of the proteins or protein fragments set forth in Table 4C and a heterologous peptide sequence fused to the N-terminus or C-terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more sequences listed in Table 4C.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more sequences listed in Table 4C fused to a heterologous peptide sequence on the N terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4C fused to a heterologous peptide sequence on the C terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C fused to a heterologous peptide sequence on the N terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C fused to a heterologous peptide sequence on the C terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C fused to a heterologous peptide sequence on the N terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C fused to a heterologous peptide sequence on the C terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4C.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4C fused to a heterologous peptide sequence on the N terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4C fused to a heterologous peptide sequence on the C terminus.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 50% sequence identity to any one of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 60% sequence identity to any one of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 70% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 75% sequence identity to any one of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragments that comprises an amino acid sequence that has at least about 80% sequence identity to any one of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 85% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 90% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 95% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 96% sequence identity to any one of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 97% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 98% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 99% sequence identity to any one of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 50% sequence identity to one or more of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 60% sequence identity to one or more of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 70% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 75% sequence identity to one or more of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to one or more of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragments that comprises an amino acid sequence that has at least about 80% sequence identity to one or more of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 85% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 90% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 95% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 96% sequence identity to one or more of the sequences set forth in Table 4C.
- the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 97% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 98% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 99% sequence identity to one or more of the sequences set forth in Table 4C.
- the plasma membrane recruitment element is identified and/or isolated.
- the plasma membrane recruitment element comprises at least about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to the sequence set forth in any one of SEQ ID NOs: 161-290.
- the plasma membrane recruitment element is engineered to comprise one or more mutations to yield PH domain variants.
- the plasma membrane recruitment element variants comprises at least about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to the sequence set forth in any one of SEQ ID NOs: 161-290.
- the plasma membrane recruitment element can also include a membrane protein (e.g. , a human membrane protein), a transmembrane domain thereof, or a biologically active mutant thereof.
- a membrane protein e.g. , a human membrane protein
- the transmembrane domain of a human protein can be a tetraspanin or a biologically active mutant thereof.
- the plasma membrane recruitment element comprises a transmembrane domain of human CD9 or a biologically active mutant thereof.
- the plasma membrane recruitment element comprises a transmembrane domain of human CD47 or a biologically active mutant thereof.
- the plasma membrane recruitment element comprises a transmembrane domain of human CD63 or a biologically active mutant thereof.
- the plasma membrane recruitment element comprises a transmembrane domain of human CD81, or a biologically active mutant thereof.
- the plasma membrane recruitment element can comprise a retroviral gag or a biologically active mutant thereof.
- the mutant of a retroviral gag can include only a portion of the retroviral gag.
- the plasma membrane recruitment element can include a gag of an alpha retrovirus or a biologically active mutant thereof.
- the plasma membrane recruitment element can a beta retrovirus or biologically active mutant thereof.
- the plasma membrane recruitment element can include a gamma retrovirus or biologically active mutant thereof.
- the plasma membrane recruitment element can include a delta retrovirus or biologically active mutant thereof.
- the plasma membrane recruitment element can include or biologically active mutant thereof.
- the plasma membrane recruitment element can include an epsilon retrovirus or biologically active mutant thereof.
- the plasma membrane recruitment element can include a spumavirus or biologically active mutant thereof.
- the retroviral gag can include a gag of HIV (e.g., HIV-1), a gag of murine leukemia virus (MLV), a gag of Moloney murine leukemia virus (MMLV), a gag of Simian immunodeficiency virus (SIV), a gag of Rous sarcoma virus (RSV), a gag of human T-cell leukemia virus type-1 (HTLV), or a gag of bovine leukemia virus (BLV), or a biologically active mutant thereof.
- HIV e.g., HIV-1
- MMV murine leukemia virus
- MMLV gag of Moloney murine leukemia virus
- SIV Simian immunodeficiency virus
- RSV Rous sarcoma virus
- HTLV human T-cell leukemia virus type-1
- BLV
- the plasma membrane recruitment element can include a gag of HIV (e.g., HIV-1) or a biologically active mutant thereof.
- the plasma membrane recruitment element can include a gag of MLV or a biologically active mutant thereof.
- the plasma membrane recruitment element can include a gag of RSV or a biologically active mutant thereof.
- the plasma membrane recruitment element can include a gag of Friend murine leukemia virus (FMLV) or biologically active mutant thereof.
- FMLV Friend murine leukemia virus
- the envelope protein comprises one or more of the sequences set forth in Table 4D with at least one amino acid substitution, deletion, or insertion.
- N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein.
- the envelope protein comprises one or more of the sequences set forth in Table 4D and a heterologous peptide sequence fused to the N-terminus or C-terminus.
- the plasma membrane recruitment element comprises any one of the sequences described in Table 4D with a further truncation on the N-terminus.
- the N-terminal methionine can be absent.
- the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with a further truncation on the C-terminus.
- the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with one amino acid substitution.
- the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with two or more amino acid substitutions.
- the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D and a heterologous peptide sequence fused to the N-terminus or C- terminus.
- the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with a further truncation on the N-terminus. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with a further truncation on the C-terminus. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with one amino acid substitution. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with two or more amino acid substitutions. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth Table 4D and a heterologous peptide sequence fused to the N-terminus or C-terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4D.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4D fused to a heterologous peptide sequence on the N terminus.
- the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4D fused to a heterologous peptide sequence on the C terminus.
- the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with at least one amino acid substitution, deletion, or insertion.
- N- terminal methionine can be absent from the plasma membrane recruitment element of the lipid delivery particle provided herein relative to the wild-type viral envelope protein.
- the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D and a heterologous peptide sequence fused to the N-terminal or C-terminal.
- the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with a further truncation on the N-terminus.
- the N-terminal methionine can be absent.
- the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with a further truncation on the C-terminus.
- the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with one amino acid substitution.
- the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with two or more amino acid substitutions.
- the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D and a heterologous peptide sequence fused to the N-terminal or C-terminal. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with a further truncation on the N-terminus. For example, for those amino acid sequences start with a N-terminal methionine, the N-terminal methionine can be absent. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with a further truncation on the C-terminus. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with one amino acid substitution.
- the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with two or more amino acid substitutions. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D and a heterologous peptide sequence fused to the N-terminal or C-terminal.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4D. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 60% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48 In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 70% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 75% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 85% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 90% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 95% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 96% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 97% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48.
- the plasma membrane recruitment element comprises an amino acid sequence that has at least about 98% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 99% sequence identity to a sequence set forth in any one of
- HERV-K113 HERV-K101, HERV-K102, HERV-K104, HERV-K107, HERVK108, HERV-K109, HERV-K115, HERV- KI lp22, and HERV-K12ql3.
- the lipid delivery particle disclosed herein comprises a protein core that is composed of at least a structural protein of a viral origin, for instance, a retroviral gag protein.
- the lipid delivery particle comprises a retroviral gag-pro-pol polyprotein, e.g., a gag-pro-pol poly protein from HIV, MMLV, or FMLV, which can help assemble a protein core of the lipid delivery particle.
- some of the gag-pro-pol polyprotein is cleaved, e.g., by pro (protease) present freely or in the gag-pro-pol polyprotein.
- the cleavage by pro can be inefficient, and the resultant cleavage products can include gag polyprotein, gag-pro polyprotein, free pro, and free pol (polymerase).
- a retroviral gag polyprotein can be further cleaved into MA, CA, NC, and other small fragments, if any.
- the lipid delivery particle comprises a retroviral gag-pro polyprotein without the pol component, and the gag-pro polyprotein can help form a protein core of the lipid delivery particle.
- the gag -pro can also be cleaved by pro, in some cases, inefficiently, into separate gag and pro proteins.
- a gag -pro or gag-pro-pol polyprotein from one species of virus can help assemble form a protein core of the lipid delivery particle
- a chimeric protein in the lipid delivery particle, discussed infra can comprise a payload fused with a gag protein from a different species of virus (e.g., an MMLV), or from a HERV, or a PH domain or transmembrane domain of a huma protein (e.g. , a PH domain of human Aktl with E17K substitution).
- the present disclosure provides a chimeric protein comprising a plasma membrane recruitment element and a payload that is a protein or a fragment thereof.
- the lipid delivery particle comprises a chimeric protein comprising a plasma membrane recruitment element and a payload that is a protein or a fragment thereof.
- the plasma membrane recruitment element and the payload are fused directly in the chimeric protein.
- the plasma membrane recruitment element and the payload are fused indirectly via a linker.
- the linker between the plasma membrane recruitment element and the payload is a cleavable linker that is recognized by a protease.
- the chimeric protein (e.g., comprising a gag protein) can form at least part of a protein core of the lipid delivery particle.
- a lipid delivery particle can comprise two or more chimeric proteins.
- the chimeric protein can include a structural protein.
- the structural protein can comprise a plasma membrane recruitment element (e.g., retroviral gag protein).
- the plasma membrane recruitment element can be fused to a payload.
- the two or more chimeric proteins comprise the same structural protein.
- the two or more chimeric proteins comprise different structural proteins.
- the two or more chimeric proteins comprise different payloads.
- the chimeric protein comprises a payload that comprises a nucleic acid-binding moiety.
- the payload further comprises a guide nucleic acid molecule that forms a ribonucleoprotein complex with the nucleic acid-binding moiety.
- the chimeric protein is suitable for delivery by a lipid delivery particle disclosed herein.
- the lipid delivery particle of the present disclosure further comprises a protease that recognizes the cleavable linker in the chimeric protein and cuts the chimeric protein at the cleavable linker. As a result of the cleavage at the cleavable linker by the protease, the payload can be separated from the plasma membrane recruitment element. In some cases, the payload is present as a "free" entity separate from the plasma membrane recruitment element.
- the payload can be free and present within an inside of the protein core of the lipid delivery particle.
- the protease is part of a second chimeric protein comprising a second plasma membrane recruitment element and the protease, where the second plasma membrane recruitment element can be either different from or same as the plasma membrane recruitment element that is fused with the payload.
- the chimeric protein disclosed herein also comprises one or more non-cleavable linkers that operably link components together.
- the non-cleavable linker can be any suitable linker sequence that is used for chimeric protein construction, such as peptide linkers that consist of glycine (Gly) and serine (Ser) residues.
- the non-cleavable linker comprises an amino acid sequence selected from the group consisting of: (GS)x (SEQ ID NO: 564), (GGS)x (SEQ ID NO: 565), (GGGGS)x (SEQ ID NO: 566), (GGSG)x (SEQ ID NO: 567), and (SGGG)x (SEQ ID NO: 568), and wherein x is an integer from 1 to 50.
- the chimeric protein of the present disclosure comprises a nuclear export signal (NES) sequence that can direct transport of the chimeric protein out of the nucleus of a cell, e.g. , a producer cell.
- NES nuclear export signal
- the chimeric protein disclosed herein has one of the following configurations of components positioned in an order from N-terminus to C-terminus:
- n is an integer in the range of from 1 to 10, and denotes the number of repeats of the NES sequence.
- Non-cleavable linker sequence can be present or absent in any of the foregoing configurations between any two neighboring components.
- the payload sequence in the chimeric protein can have one or more NLS sequences, at its N-terminus, C-terminus, or both.
- n is an integer in the range of from 1 to 10, and denotes the number of repeats of the NES sequence.
- Non-cleavable linker sequence can be present or absent in any of the foregoing configurations between any two neighboring components.
- the payload sequence in the chimeric protein can have one or more NLS sequences, at its N-terminus, C-terminus, or both.
- nuclear export signal refers to a sequence of amino acids that targets a payload protein for export from the nucleus.
- NES nuclear export signal
- NES is a short target peptide sequence containing four hydrophobic residues. These residues target the protein for export from the nucleus to the cytoplasm through the nuclear pore complex.
- a chimeric protein provided herein can comprise 1 NES, 2 NESs, 3 NESs, 4 NESs, 5 NESs, 6 NESs, 7 NESs, 8 NESs, 9 NESs, or 10 NESs.
- the NES is located at the N-terminus, C-terminus, or in an internal region of the chimeric protein.
- a NES is coupled between the plasma membrane recruitment element and the payload in the chimeric protein.
- the NES sequence that is used in the chimeric protein comprises LQLPPLERLTL (SEQ ID NO: 403) derived from HIV-1 Rev protein, or any of the sequences having at least 80% identity thereto.
- the NES sequence comprises LALKLAGLDI (SEQ ID NO: 352) or NELALKLAGLDI (SEQ ID NO: 416), derived from PKIa, or any of the sequences having at least 80% identity thereto.
- the NES sequence that is used in the chimeric protein comprises an amino acid sequence as set forth in Table 5.
- the NES sequence comprises any one of the sequences set forth in Table 5.
- the NES sequence comprises one or more of the sequences set forth in Table 5.
- the NES sequence comprises more than one, more than two, more than three, more than four, more than five, more than six, more than seven, more than eight, more than nine, or more than ten of the sequences set forth in Table 5. In some cases, the NES sequences comprises multiple sequences set forth in Table 5.
- the NES sequence comprises an amino acid sequence having 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any sequence listed in Table 5.
- the NES sequence comprises an amino acid sequence having 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any sequence set forth in SEQ ID NOs: 353-453.
- the NES sequence described herein comprises a sequence with greater than 80% sequence identity to any sequence listed in Table 5. The transport of payload proteins within a cell is enabled through both NES and nuclear export receptors.
- the NES described herein is associated with a nuclear export receptor (e.g., CRM-1).
- the NES may be conditionally active or inactive.
- the NES sequence disclosed herein comprises a sequence such as those described in T la Cour, et al., Nucleic Acids Res. 2003;31 ( 1): 393-396; and Xu D, et al. Mol Biol Cell. 2012 Sep;23(18):3673-6, each of which is incorporated herein by reference in its entirety.
- any of the NES sequences described in the NES sequence database can be used in a chimeric protein disclosed herein, e.g., for the purpose of packaging a payload into the molecular assembly, e.g., the lipid delivery particle.
- a chimeric protein disclosed herein include a nuclear export sequence (NES).
- the NES facilitates localization of the chimeric protein in the cytosol of a target cell relative to the nucleus.
- a chimeric protein disclosed herein includes at least one NES sequences, such as, 2 or more, 3 or more, 4 or more, or 5 or more NES sequences.
- one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g., within 50 amino acids of) the N-terminus and/or the C- terminus of the chimeric protein.
- the chimeric protein disclosed herein comprises only one NES sequence.
- the chimeric protein disclosed herein comprises two NES sequences.
- the chimeric protein disclosed herein comprises three NES sequences.
- one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g. , within 50 amino acids of) the N- terminus of the chimeric protein. In some cases, one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g. , within 50 amino acids of) the C- terminus of the chimeric protein. In some cases, one or more NES sequences (3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g., within 50 amino acids of) both the N-terminus and the C-terminus of the chimeric protein. In some cases, an NES sequence is positioned at the N- terminus and an NES sequence is positioned at the C-terminus of the chimeric protein.
- a payload is a protein that is delivered as part of the chimeric protein disclosed herein, e.g., operably linked to a structural protein (e.g., human endogenous retroviral structural protein or a Plasma membrane recruitment element).
- the one or more NES sequences are positioned at or near the one or both ends of the payload protein sequence inside the chimeric protein.
- one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g., within 50 amino acids of) the N-terminus and/or the C- terminus of the payload protein sequence.
- one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g. , within 50 amino acids of) the N-terminus of the payload protein sequence. In some cases, one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g. , within 50 amino acids of) the C-terminus of the payload protein sequence. In some cases, one or more NES sequences (3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g. , within 50 amino acids of) both the N-terminus and the C-terminus of the payload protein sequence.
- an NES sequence is positioned at the N-terminus and an NES sequence is positioned at the C-terminus of the payload protein sequence.
- the chimeric protein disclosed herein comprises only one NES sequence.
- the chimeric protein comprises only one NES sequence, and the NES sequence is positioned at or near (e.g., within 50 amino acids of) the N-terminus of the payload protein.
- NESs nuclear export sequences
- NESs can direct export of proteins from the nucleus to the cytoplasm.
- NESs can bind directly to the export karyopherin CRM1 (also known as exportin 1), which can escort payload proteins through the nuclear pore complex.
- a payload described herein comprises one or more nuclear localization sequences (NLS).
- NLS nuclear localization sequences
- the term “nuclear localization signal” refers to a sequence of amino acids that targets a payload (e.g. , a protein or a short polypeptide), which the NLS is present within or coupled to, to localize to the nucleus.
- a payload e.g. , a protein or a short polypeptide
- an NLS facilitates the import of a polypeptide comprising an NLS into the cell nucleus.
- a polypeptide can comprise 1 NLS, 2 NLSs, 3 NLSs, 4 NLSs, 5 NLSs, 6 NLSs, 7 NLSs, 8 NLSs, 9 NLSs, or 10 NLSs.
- the NLS is located at the N- terminus, C-terminus, or in an internal region of the polypeptide. In some cases, a NLS is coupled to a nucleic acid binding domain described elsewhere herein. In some cases, a NLS is coupled to a nucleic acid modifying domain described elsewhere herein. In some cases, a NLS is coupled to a guidable polypeptide domain, a deaminase domain, or a reverse transcriptase domain. In some cases, a NLS is covalently linked to a nucleic acid binding domain described elsewhere herein. In some cases, a NLS is covalently linked to a nucleic acid modifying domain described elsewhere herein.
- a NLS is covalently linked to a guidable polypeptide domain, a deaminase domain, or a reverse transcriptase domain.
- a nucleic acid binding domain does not comprise an NLS.
- a nucleic acid binding domain does not comprise an NLS.
- a guidable polypeptide domain, a deaminase domain, or a reverse transcriptase domain does not comprise an NLS. Examples of NLS are provided in Table 6 below.
- the NLS comprises an amino acid sequence as set forth in Table 6. In some cases, the NLS comprises any one of the sequences set forth in Table 6. In some cases, the NLS comprises one or more of the sequences set forth in Table 6. In some cases, the NLS comprises more than one of the sequences set forth in Table 6. In some cases, the NLS comprises multiple sequences set forth in Table 6. In some cases, NLS sequence can comprise an amino acid sequence having 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any sequence listed in Table 6. In some cases, the NLS sequence described herein can comprise a sequence with greater than 80% sequence identity to any sequence listed in Table 6.
- NLS sequence can comprise an amino acid sequence having 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any sequence set forth in SEQ ID NOs: 454-477.
- a chimeric protein disclosed herein includes a nuclear localization sequence (NLS).
- NLS nuclear localization sequence
- the NLS facilitates delivery of the chimeric protein, or a payload released from the chimeric protein (for instance, released from the chimeric protein following cleavage of a cleavable linker), into the nucleus of a target cell.
- a payload is a protein and is delivered as part of the chimeric protein disclosed herein, e.g., operably linked to a structural protein (e.g., plasma membrane recruitment element).
- the one or more NLS sequences are positioned at or near the one or both ends of the payload protein sequence of the chimeric protein.
- a chimeric protein includes (e.g., is fused to) between 2 and 5 NLS sequences (e.g., 2-4, or 2-3 NLSs).
- NLS sequences include an NLS sequence derived from: the NLS of the SV40 virus large T-antigen, having the amino acid sequence PKKKRKV (SEQ ID NO: 468); the NLS from nucleoplasmin (e.g., the nucleoplasmin bipartite NLS with the sequence KRPAATKKAGQAKKKK (SEQ ID NO: 460); the c-myc NLS having the amino acid sequence PAAKRVKLD (SEQ ID NO: 467) or RQRRNELKRSP (SEQ ID NO: 541); the hRNPAl M9 NLS having the sequence NQSSNFGPMKGGNFGGRSSGPYGGGGQYFAKPRNQGGY (SEQ ID NO: 542); the sequence RMRIZFKNKGKDTAELRRRRVEVSVELRKAKKDEQILKRRNV (SEQ ID NO: 543) of the IBB domain from importin-alpha; the sequences VSRKRPRP (SEQ ID NO: 477) and PPKKA
- NLS sequence examples include KRTADGSEFESPKKKRKV (SEQ ID NO: 462), KKTELQTTNAENKTKKL (SEQ ID NO: 554), KRGINDRNFWRGENGRKTR (SEQ ID NO: 555), RKSGKIAAIVVKRPRK (SEQ ID NO: 556), and MDSLLMNRRKFLY QFKNVRWAKGRRETYLC (SEQ ID NO: 463), SPKKKRKVEAS (SEQ ID NO: 557), encoded by AGCCCCAAGAAgAAGAGaAAGGTGGAGGCCAGC (SEQ ID NO: 558), GPKKKRKVAAA (SEQ ID NO: 559), as well as any of those described in Cokol et al., EMBO Rep., 2000, 1(5): 411-415 and Freitas et al., Current Genomics, 2009, 10(8): 550-7; Lu, J., et la., Cell Commun Signal 19, 60 (20
- the chimeric protein comprises one NES sequence and two NLS sequences. In some cases of these embodiments, the NES sequence, NLS sequences, and the payload protein sequence are positioned in an order from N-terminus to C-terminus as follows: NES-NLS-payload protein-NLS. In some embodiments, the chimeric protein comprises two or more NES sequences and two NLS sequences. In some cases of these embodiments, the NES sequences, NLS sequences, and the payload protein sequence are positioned in an order from N-terminus to C-terminus as follows: n X NES
- the chimeric protein comprises a cleavable linker in between two or more components.
- the chimeric protein can comprise a cleavable linker between a payload protein sequence and a plasma membrane recruitment element sequence (e.g., retroviral gag protein sequence).
- the cleavable linker separates the plasma membrane recruitment element sequence from a NLS sequence, and/or a NES sequence at its N-terminus or C-terminus.
- the cleavable linker can separate the payload protein sequence from the plasma membrane recruitment element sequence, NLS sequence, and/or NES sequence at its N-terminus or C-terminus.
- cleavable linker sequences that can be used in the chimeric protein include TSTLLMENSS (SEQ ID NO: 560), PRSSLYPALTP (SEQ ID NO: 561), VQALVLTQ (SEQ ID NO: 562), and PLQVLTLNIERR (SEQ ID NO: 563), and sequences having at least 80% identity to any one of the foregoing.
- a payload in a lipid delivery particle of the present disclosure can comprise a protein, a polypeptide, a nucleic acid (e.g., DNA or RNA), or any combinations thereof.
- the payload can be a part of the chimeric protein disclosed herein or can comprise a part of the chimeric protein disclosed herein.
- the payload can include an entity in the lipid delivery particle separate from the chimeric protein disclosed herein.
- the payload is a protein or polypeptide coupled to a plasma membrane recruitment element.
- the payload comprises a first moiety (e.g, a nucleic acid-binding protein) that is fused to a plasma membrane recruitment element, and further comprises a second moiety that is coupled to the first moiety via covalent or non-covalent interaction.
- the first moiety can be a nucleic acid binding protein that is fused with the plasma membrane recruitment element
- the second moiety can be a nucleic acid molecule that binds to the nucleic acid binding protein.
- a payload is directly packaged within the lipid delivery particles and delivered into a target cell in its free form.
- a payload can be fused to a plasma membrane recruitment element (e.g., pleckstrin homology domain) and form a chimeric protein as part of the lipid delivery particles, and then delivered into the target cell.
- the plasma membrane recruitment element e.g., pleckstrin homology domain
- the payload in its free form or as part of a chimeric protein is within the inside cavity of the protein core of the lipid delivery particles disclosed herein.
- the payload in its free form derives from a cleavage of the chimeric protein comprising the payload.
- a lipid delivery particle can deliver more than one payload.
- Each of the payloads can independently comprise nucleic acid-binding moiety, a nucleic acid-modifying moiety, a fusion protein, or a nucleic acid, or any combinations thereof.
- the plasma membrane recruitment element and the payload are coupled via any suitable method.
- Covalent coupling between the plasma membrane recruitment element and a payload peptide can include inteins that can form peptide bonds, direct protein-protein chimeras generated from a single reading frame.
- nucleic acids base pairing to other nucleic acids via hydrogen bonding interactions (e.g., DNA/RNA, DNA/DNA, or RNA/RNA hybrids), protein-protein binding, or protein-nucleic acid molecule binding can be involved for the coupling between the plasma membrane recruitment element and the payload.
- protein-nucleic acid molecule binding examples include an RNA binding protein (RBP) and an RBP binding sequence (e.g., an RNA) that binds to the RBP.
- RBP RNA binding protein
- RBP binding sequence e.g., an RNA
- each of the plasma membrane recruitment element and the payload is fused to a heterologous sequence, and the two heterologous sequences dimerize or multimerize with or without the need for a chemical compound to induce the protein-protein binding, such as a single -stranded nucleic acid sequence or protein dimerization domains).
- each of the plasma membrane recruitment element and the payload is fused to one member of a pair of binding partners (e.g., antibody and its target antigen).
- the plasma membrane recruitment element is fused to an RBP, and the payload is fused to a RBP binding sequence.
- suitable protein domains or nucleic acid molecules for forming the non-covalent connections include single chain variable fragments, nanobodies, affibodies, DmrA/DmrB/DmrC, FKBP/FRB, dDZFs, Leucine zippers, proteins that bind to DNA and/or RNA, optogenetic protein domains that can dimerize or multimerize in the presence of certain light wavelengths, proteins with quaternary structural interactions, and/or naturally reconstituting split proteins.
- RBPs and their RBP binding sequences examples include a sequence having at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in Table 7.
- RBPs and their RBP binding sequences that can be used include a sequence having at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 478- 513.
- the RBP comprises an amino acid sequence as set forth in Table 7.
- the RBP comprises any one of the sequences set forth in Table 7.
- the RBP comprises one or more of the sequences set forth in Table 7.
- the RBP comprises more than one of the sequences set forth in Table 7.
- the RBP comprises multiple sequences set forth in Table 7.
- the RBP binding sequence comprises an amino acid sequence as set forth in Table 7.
- the RBP binding sequence comprises any one of the sequences set forth in Table 7.
- the RBP binding sequence comprises one or more of the sequences set forth in Table 7.
- the RBP binding sequence comprises more than one of the sequences set forth in Table 7.
- the RBP binding sequence comprises multiple sequences set forth in Table 7.
- RNA binding proteins RBP
- RBP RNA binding proteins
- nucleic acid binding domains and nucleic acid modifying domains are nucleic acid binding domains and nucleic acid modifying domains
- the payload comprises a nucleic acid-binding moiety, a nucleic acid-modifying moiety, a fusion protein, or a nucleic acid.
- the payload comprises a nucleic acid-binding domain, e.g., a DNA-binding protein domain or polypeptide or an RNA-binding domain or polypeptide e.g., an RNA-binding protein (RBP).
- a nucleic acid-binding moiety can be capable of binding a nucleic acid.
- a nucleic acid-binding domain can bind to a nucleic acid in a nonspecific or a site-specific manner.
- the nucleic acid-binding moiety binds to a nucleic acid in a site-specific manner.
- a nucleic acid-binding moiety can comprise an aptamer binding domain that selectively binds to a specific target.
- a nucleic acid-binding moiety recognizes a specific recognition sequence in the target nucleic acid.
- a nucleic acid-binding moiety comprises an aptamer binding domain.
- a nucleic acid binding moiety selectively binds to a sequence or a structural element in a nucleic acid molecule.
- an RNA-binding domain selectively binds to a specific sequence motif in an RNA molecule.
- a nucleic acid-binding moiety selectively binds to a structural element in a nucleic acid molecule.
- a nucleic acid-binding domain can bind to a stem -loop in a nucleic acid molecule.
- a nucleic acid-binding moiety is or comprises a guidable polypeptide domain, a transcriptional regulatory domain, or a nucleic acid-modifying domain.
- a guidable polypeptide domain can be capable of binding to a polynucleotide (e.g. an RNA guide) that can direct the guidable polypeptide domain a target site.
- the guidable polypeptide domain forms a complex with the RNA guide and recognizes the target sequence through DNA-RNA base pairing.
- a nucleic -acid binding moiety is or comprises a transcriptional regulatory domain.
- a nucleic- binding moiety can help recruit a transcriptional repressor or activator to a target site.
- a nucleic acid-binding moiety is or comprises a nucleic acid-modifying moiety.
- the present disclosure uses nucleic acid-binding moieties to recruit a nucleic acid-modifying moiety to a target site.
- a nucleic-acid binding moiety comprises catalytic activity.
- a nucleic acidbinding moiety is catalytically inactive.
- a nucleic-acid binding moiety comprising catalytic activity is modified to have a reduced level of activity compared to its wild-type counterpart.
- the payload in the present disclosure comprises a nucleic acid modifying domain.
- a nucleic acid-modifying domain can comprise a polypeptide domain, a nucleic acid or a combination thereof (e.g., a ribonucleoprotein complex).
- a nucleic acid-modifying domain can be capable of modifying nucleic acid, such as cleaving double-stranded nucleic acid; nicking a single -stranded nucleic acid; introducing a mutation, deletion, or insertion in a nucleic acid; methylating or demethylating a nucleic acid, or altering the structure of DNA (e.g., changing chromatin structure through modifying histones).
- a nucleic acid modifying domain can comprise a nuclease domain, a nickase domain, a deaminase domain, a polymerase, reverse transcriptase domain, a recombinase domain, a transposase domain, or an epigenetic modifying domain.
- a nuclease domain can be capable of cleaving phosphodiester bonds between nucleotides in nucleic acids.
- a nuclease domain can comprise an exonuclease (e.g., a nuclease capable of cleaving nucleic acids from the ends) or an endonuclease (e.g., a nuclease capable of cleaving nucleic acids in the middle).
- a nucleic acid modifying effector or nucleic acid binding domain is a nickase, which can be capable of cleaving a single-strand in a double -stranded DNA.
- Nucleic acid modifying domains can be useful for gene editing, or for regulating, activating, or inhibiting gene expression.
- the payload in the present disclosure comprises a guidable polypeptide domain (e.g., a CRISPR-Cas protein domain).
- a guidable polypeptide domain is capable of binding to a polynucleotide (e.g. , an RNA guide) that directs it to a target site .
- the guidable polypeptide domain forms a complex with the polynucleotide and recognizes the target sequence through DNA-RNA base pairing.
- a guidable polypeptide domain is a CRISPR/CRISPR-associated (Cas) domain.
- a CRISPR domain can be a natural or an engineered domain.
- a Cas protein or domain can be derived from a CRISPR system or share structural and/or functional similarities to a protein involved in a CRISPR system.
- the guidable polypeptide domain is any suitable nuclease, e.g., a CRISPR- associated (Cas) protein or a Cas nuclease which functions in a non-naturally occurring CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)/Cas (CRISPR-associated) system.
- a CRISPR-associated (Cas) protein or a Cas nuclease which functions in a non-naturally occurring CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)/Cas (CRISPR-associated) system.
- this system can provide adaptive immunity against foreign DNA (Barrangou, R., et al, “CRISPR provides acquired resistance against viruses in prokaryotes,” Science (2007) 315: 1709-1712; Makarova, K.S., et al, “Evolution and classification of the CRISPR-Cas systems,” Nat Rev Microbiol (2011) 9:467- 477; Gameau, J.
- Suitable nucleases include CRISPR-associated (Cas) proteins or Cas nucleases including type I CRISPR-associated (Cas) polypeptides, type II CRISPR-associated (Cas) polypeptides (e.g., Cas9 or Cas 14), type III CRISPR-associated (Cas) polypeptides, type IV CRISPR-associated (Cas) polypeptides, type V CRISPR-associated (Cas) polypeptides (e.g., Cpfl/Casl2a, C2cl, or c2c3), and type VI CRISPR- associated (Cas) polypeptides (e.g., C2c2/Casl3a, Cas 13b, Cas 13c, Cas 13d).
- type I CRISPR-associated (Cas) polypeptides e.g., Cas9 or Cas 14
- type III CRISPR-associated (Cas) polypeptides e.g.
- a CRISPR system is a system encoding DNA sequence arrays known as clustered regularly interspaced short palindromic repeats (CRISPRs), which can be found in microbial genomes or phage genomes.
- CRISPR systems comprise genes encoding CRISPR-associated (Cas) proteins and/or small RNA guide molecules (e.g., crRNA or tracrRNA) that assemble with the CRISPR domain.
- the CRISPR-Cas domain forms a complex with one or more RNA guide molecules to form an effector ribonucleoprotein complex.
- the effector ribonucleoprotein complex can recognize a target sequence through sequence specific DNA-RNA base pairing with a spacer sequence in the RNA guide.
- target recognition activates one or more nuclease domains (e.g., a RuvC domain or HNH domain) in the CRISPR domain to make a double -stranded cut at the target DNA.
- a CRISPR-Cas domain complexed with an RNA guide can be capable of inactivating target gene through a gene knockout.
- the CRISPR domain is used to enable gene insertion and/or deletion, which can inactivate, modify, or restore the gene’s function.
- One or more components of a CRISPR/Cas system (e.g., modified and/or unmodified) delivered by the lipid delivery particles disclosed herein can be utilized as a genome engineering tool in a wide variety of organisms including diverse mammals, animals, plants, and yeast.
- a CRISPR/Cas system can comprise a guide nucleic acid such as a guide RNA (gRNA) complexed with a Cas protein for targeted regulation of gene expression and/or activity or nucleic acid editing.
- gRNA guide RNA
- RNA-guided Cas protein e.g., a Cas nuclease such as a Cas9 nuclease
- a target polynucleotide e.g., DNA
- the Cas protein if possessing nuclease activity, can cleave the DNA (Gasiunas, G., etal, “Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria,” Proc Natl Acad Sci USA (2012) 109: E2579-E2 86; Jinek, M., et al, “A programmable dual -RNA-guided DNA endonuclease in adaptive bacterial immunity,” Science (2012) 337:816-821; Sternberg, S.
- the Cas protein is mutated and/or modified to yield a nuclease deficient protein or a protein with decreased nuclease activity relative to a wild-type Cas protein.
- a nuclease deficient protein can retain the ability to bind DNA but can lack or have reduced nucleic acid cleavage activity.
- a protein encoded by a donor sequence comprises a Cas nuclease (e.g., retaining wild-type nuclease activity, having reduced nuclease activity, and/or lacking nuclease activity) can function in a CRISPR/Cas system to regulate the level and/or activity of a target gene or protein (e.g., decrease, increase, or elimination).
- the Cas protein can bind to a target polynucleotide and prevent transcription by physical obstruction or edit a nucleic acid sequence to yield non -functional gene products.
- the Cas protein cleaves both strands of DNA.
- the Cas protein cleaves one strand of DNA.
- the nuclease is a Cas protein that forms a complex with a guide nucleic acid, such as a guide RNA (gRNA).
- the donor sequence disclosed herein encodes a Cas protein that forms a complex with a single guide nucleic acid, such as a single guide RNA (sgRNA).
- the donor sequence disclosed herein encodes a Cas protein that forms a complex with two separate RNA molecules of a dual guide nucleic acid (dgRNA).
- the donor sequence in the lipid delivery particles disclosed herein comprises or encodes an RNA-binding protein (RBP) optionally complexed with a guide nucleic acid, such as a guide RNA (e.g. , sgRNA, dgRNA), which is able to form a complex with a Cas protein.
- a guide RNA e.g. , sgRNA, dgRNA
- the gRNA comprises a scaffolding sequences that tethers the gRNA to the Cas protein.
- the gRNA comprises a scaffolding sequence and a spacer sequence that directs the Cas protein to a specific locus.
- the scaffolding sequence is configured to bind to the positively charged groves in the Cas9 protein.
- the scaffolding sequence is configured to bind to the Cas protein in the payload.
- Cas undergoes a conformational change when the gRNA binds to the target locus.
- the conformational change in Cas shifts the molecule from an inactive, non-DNA binding conformation into an active DNA-binding conformation.
- the Cas protein undergoes a confirmational change if the spacer sequence has sufficient homology to the sequence at the target locus.
- gRNAs can be modified.
- Exemplary modifications to the gRNA are provided in United States Patent Number 11,479,767 B2, United States Patent Application Publication Number US2020/0339980 Al, and United States Patent Application Publication Number US2021/0079389 Al, each of which is incorporated herein by reference in its entirety.
- One or more components of any suitable CRISPR/Cas system can be delivered by the lipid delivery particle described in the present disclosure.
- a CRISPR/Cas system can be referred to using a variety of naming systems. Exemplary naming systems are provided in Makarova, K.S. et al, “An updated evolutionary classification of CRISPR-Cas systems,” Nat Rev Microbiol (2015) 13:722-736 and Shmakov, S. et al, “Discovery and Functional Characterization of Diverse Class 2 CRISPR-Cas Systems,” Mol Cell (2015) 60: 1-13.
- a CRISPR/Cas system can be a type I, a type II, a type III, a type IV, a type V, a type VI system, or any other suitable CRISPR/Cas system.
- a CRISPR/Cas system as used herein can be a Class 1, Class 2, or any other suitably classified CRISPR/Cas system. Class 1 or Class 2 determination can be based upon the genes encoding the effector module. Class 1 systems generally have a multi-subunit crRNA-effector complex, whereas Class 2 systems generally have a single protein, such as Cas9, Cpfl, C2cl, C2c2, C2c3, or a crRNA-effector complex.
- a Class 1 CRISPR/Cas system can use a complex of multiple Cas proteins to effect regulation.
- a Class 1 CRISPR/Cas system can comprise, for example, type I (e.g, I, IA, IB, IC, ID, IE, IF, IU), type III (e.g., Ill, IIIA, IIIB, IIIC, IIID), and type IV (e.g., IV, IVA, IVB) CRISPR/Cas type.
- a Class 2 CRISPR/Cas system can use a single large Cas protein to effect regulation.
- a Class 2 CRISPR/Cas systems can comprise, for example, type II (e.g., II, IIA, IIB) and type V CRISPR/Cas type.
- CRISPR systems can be complementary to each other, and/or can lend functional units in trans to facilitate CRISPR locus targeting.
- Cas proteins that can be used as part of the CRISPR systems described herein include c2cl, Casl3a (formerly C2c2), Casl3b, Casl3c, Casl3d, c2c3, Casl, CaslB, Cas2, Cas3, Cas4, Cas5, Cas5e (CasD), Cas6, Cas6e, Cas6f, Cas7, Cas8a, Cas8al, Cas8a2, Cas8b, Cas8c, Cas9 (Csnl or Csxl2), CaslO, CaslOd, Casl4, CaslO, CaslOd, CasF, CasG, CasH, Casl2a (formerly Cpfl), Csyl, Csy2, Csy3, Csel (CasA), Cse2
- mutant Cas9 proteins or Cas9 variants include SpG, SpCas9-NG. Cas9-NRNH, SpG, SpRY, Cas9-VQR, Cas9-EQR, SaCas9-KKH, Nme2Cas9, eNme2-C, eNme2-C.NR, eNme2-T.l, eNme2-T.2, SpRY, eSpCas9(l.l), SpCas9-HFl, nSpCas9, eSpCas9, Sniper-Cas9, HypaCas9, evoCas9, Cas9TX, HscCas9-vl.2, superFi-Cas9, efSaCas9, SaCas9- HF, Cas9-HF, Cas mini, SaCas9, SpCas9(H840A), dSpCas9, SpCa
- a CRISPR system can comprise single subunit or multi-subunit effectors.
- a CRISPR system is a Class 1 CRISPR system.
- a Class 1 CRISPR system can be a type I, type III, or a type IV system.
- a Class 1 type I CRISPR system can comprise a multi-subunit effector.
- a Class 1 type I CRISPR system comprises a protein or domain in the Cascade-Cas3 protein complex.
- a Class 1 type I CRISPR system can comprise a Cas6, Cas7, Cas5, Casl 1, Cas8, or Cas3 domain.
- a Class 1 type III CRISPR system can comprise a multi-subunit effector.
- a Class 1 type III CRISPR system comprises a Csm complex or a Cmr complex.
- a Class 1 type III CRISPR system comprises a Cas6, a Cas7 (Csm3 or Cmr4), a Cas7-related (Csm5, Cmrl, or Cmr6), a Cas5 (e.g., Csm4 or Cmr5), a Casl 1 (e.g., Csm2 or Cmr3), or a CaslO (e.g., Csml or Cmr2) domain.
- a Class 1 type IV CRISPR system can comprise a Cas6, a Cas7, a Cas5, a Casl 1, a Cas8 (e.g., Csfl), or a DinG or CysH domain.
- a CRISPR system comprises Cmrl, Cmr3, Cmr4, Cmr5, or Cmr6.
- a CRISPR system comprises Csbl, Csb2, or Csb3.
- a CRISPR system can comprise Csfl, Csf2, Csf3, or Csf4.
- a CRISPR system can comprise Csn2, Csm2, Csm3, Csm4, Csm5, or Csm6.
- a CRISPR system can comprise Cscl or Csc2.
- a CRISPR system can comprise Casl, CaslB, Cas2, or Cas4.
- a CRISPR system can comprise Csyl, Csy2, or Csy3.
- a CRISPR system can comprise Csel or Cse2.
- a CRISPR system can comprise Csn2.
- a CRISPR system can comprise CsaX, Csxl, Csx3, CsxlO, Csxl 4, Csxl 5, Csxl 6, or Csxl7.
- a CRISPR system comprises a modified version of any one of the foregoing Cas proteins.
- a modified version of the foregoing Cas protein comprises a nickase mutation.
- the nickase mutation corresponds to the D10A mutation of the wild type Cas9 protein. In some cases, the nickase mutation corresponds to the H840A mutation of the wild type Cas9 protein. In some cases, the nickase mutation occurs in the RuvC domain of the wild type Cas9 protein. In some cases, the nickase mutation occurs in the HNH domain of the wild type Cas9 protein. In some cases the RuvC domain can be mutated to prevent cleavage of the non-target DNA strand. In some cases the HNH domain can be mutated to prevent cleavage of the target DNA strand. In some cases, a modified version of the foregoing Cas protein comprises one or more mutations that disrupt cleavage activity.
- a Cas protein with disrupted cleavage activity is catalytically inactive or catalytically dead.
- the catalytically dead mutations occur in the RuvC domain and the HNH domain of the wild type Cas9 protein.
- the catalytically inactive mutations correspond to the D10A mutation and the H840A mutation of the wild type Cas9 protein.
- a CRISPR system is a Class 2 CRISPR system.
- a Class 2 CRISPR system can be a Class 2 type II CRISPR system, a Class 2 type V CRISPR system, or a Class 2 type VI CRISPR system.
- a Class 2 type II CRISPR system can comprise a Cas9 domain (also known as Csnl and Csxl2).
- a Cas9 domain can be a SpyCas9, a GeoCas9, a SauCas9, a KhuCas9, a AinCas9, an FmaCas9, a SgaCas9, a ScCas9, a SauriCas9 domain.
- a Cas9 domain can be a hyperactive Cas9 domain.
- a Class 2 type V CRISPR system can comprise a Casl2 domain.
- a Casl2 domain can be a Casl2a, a Casl2b, a Casl2bl, a Casl2c, a Casl2d, a Casl2e, a Casl2f, a Casl2g, a Casl2h, a Casl2i, a Casl2j, a Casl2k, a Casl21, or a Cas 12m domain.
- a Class 2 type VI CRISPR system can comprise a Cas 13 domain.
- a CRISPR system comprises a circularly permuted Cas9.
- a CRISPR system comprises CjCas9, Casl3a, Casl3b, Casl3c, or Casl3d. In some cases, a CRISPR system comprises Cas 14, xCas9, or SpCas9-NG.
- a CRISPR-Cas domain comprises one or more subdomains.
- a Cas9 domain can comprise a Reel, a Rec2, a Rec3, a RuvC, an HNH, or a Wedge/PAM-interacting domain.
- a Casl2 domain can comprise a Reel, Rec2, a crRNA oligonucleotide binding domain (OBD), a Nuc domain, a PAM-interacting (PI) domain, or a RuvC domain.
- the RuvC domain comprises nuclease activity.
- the HNH domain comprises nuclease activity.
- the PAM-interacting domain can bind to a protospacer adjacent motif (PAM) sequence that is next to a target sequence in a target nucleic acid molecule.
- PAM recognition can help activate a nuclease domain to make a cut at the target sequence.
- a CRISPR protein or domain is an engineered or mutated variant of a protein involved in a CRISPR system.
- An engineered or mutated CRISPR domain can comprise a truncation, a deletion of a part of one or more domains or subdomains, or a mutation of an active site (e.g., a RuvC active site or HNH active site).
- a CRISPR domain with a mutation of one or more active sites is catalytically inactive (e.g., dCas9).
- a CRISPR domain with one or more mutated active sites comprises less than 90%, less than 80%, less than 70%, less than 60%, less than 50%, less than 40%, less than 30%, less than 20%, less than 10%, less than 5%, or less than 1% of the nuclease activity of its wildtype counterpart.
- a dCas9 can result from the point mutations D10A in the RuvC domain and the point mutation H840A in the HNH domain.
- a mutation can result in a CRISPR nickase.
- a nickase can generate nick or a single-stranded cut.
- a nickase can generate a nick in the strand complementary to the RNA guide (e.g., the targeting strand) or in the strand on the non-targeting strand.
- a RuvC mutation D10A in a Cas9 domain can produce a Cas9 nickase domain that nicks the targeting strand.
- An HNH mutation H840A in a Cas9 domain can produce a Cas9 nickase domain that nicks the non-targeting strand.
- a Cas protein can comprise one or more domains. Examples of domains include, guide nucleic acid recognition and/or binding domain, nuclease domains (e.g, DNase or RNase domains, RuvC, HNH), DNA binding domain, RNA binding domain, helicase domains, protein-protein interaction domains, and dimerization domains.
- a guide nucleic acid recognition and/or binding domain can interact with a guide nucleic acid.
- a nuclease domain can comprise catalytic activity for nucleic acid cleavage.
- a nuclease domain can lack catalytic activity to prevent nucleic acid cleavage.
- a Cas protein can be a chimeric Cas protein that is fused to other proteins or polypeptides.
- a Cas protein can be a chimera of various Cas proteins, for example, comprising domains from different Cas proteins.
- a CRISPR system comprises an Argonaute (Ago) domain.
- Cas 14 protein or polypeptide can bind and/or modify (e.g., cleave, nick, methylate, demethylate, etc.) a target nucleic acid and/or a polypeptide associated with target nucleic acid (e.g. , methylation or acetylation of a histone tail) (e.g. , in some cases the CasZ protein includes a chimeric partner with an activity, and in some cases the CasZ protein provides nuclease activity).
- the Casl4 protein or polypeptide is a naturally occurring protein (e.g., naturally occurs in prokaryotic cells) (e.g., a CasZ protein).
- the Cas 14 protein or polypeptide not a naturally occurring polypeptide (e.g, the Cas 14 protein is a variant Cas 14 protein, a chimeric protein, and the like).
- a Cas 14 protein includes 3 partial RuvC domains (RuvC-I, RuvC-II, and RuvC-III, also referred to herein as subdomains) that are not contiguous with respect to the primary amino acid sequence of the Cas 14 protein but form a RuvC domain once the protein is produced and folds.
- a naturally occurring Cas 14 protein functions as an endonuclease that catalyzes cleavage at a specific sequence in a targeted nucleic acid (e.g., a double stranded DNA (dsDNA)).
- the sequence specificity is provided by the associated guide RNA, which hybridizes to a target sequence within the target DNA.
- the naturally occurring Cas 14 guide RNA is a crRNA, where the crRNA includes (i) a guide sequence that hybridizes to a target sequence in the target DNA and (ii) a protein binding segment that binds to the Cas 14 protein.
- Examples of Casl4 proteins include those described U.S. Patent Publication Nos.
- the donor sequence disclosed herein encodes Casl4 polypeptide or a nucleic acid molecule encoding Cas 14 polypeptide. In some cases, the donor sequence disclosed herein encodes Cas 14a polypeptide. In some cases, the donor sequence disclosed herein encodes Casl4b polypeptide. In some cases, the donor sequence disclosed herein encodes Casl4c polypeptide.
- a Cas protein can be from any suitable organism. Examples include Streptococcus pyogenes, Streptococcus thermophilus, Streptococcus sp., Staphylococcus aureus, Nocardiopsis rougevillei, Streptomyces pristinae spiralis, Streptomyces viridochromo genes, Streptomyces viridochromogenes, Streptosporangium roseum, Streptosporangium roseum, AlicyclobacHlus acidocaldarius, Bacillus pseudomycoides, Bacillus selenitireducens, Exiguobacterium sihiricum, Lactobacillus delbrueckii, Lactobacillus salivarius, Microscilla marina, Burkholderiales bacterium, Polaromonas naphthalenivorans, Polaromonas sp., Crocosphaera watsonii, Cyanothece sp., Microcysti
- the organism is Streptococcus pyogenes (S. pyogenes). In some aspects, the organism is Staphylococcus aureus (S. aureus). In some aspects, the organism is Streptococcus thermophilus (.S'. thermophilus).
- a Cas protein can be derived from a variety of bacterial species including Veillonella atypical, Fusobacterium nucleatum, Filifactor alocis, Solobacterium moorei, Coprococcus catus, Treponema denticola, Peptoniphilus duerdenii, Catenibacterium mitsuokai, Streptococcus mutans, Listeria innocua, Listeria seeligeri.
- Listeria weihenstephanensis FSL R90317 Listeria weihenstephanensis FSL M60635, Staphylococcus pseudintermedius, Acidaminococcus intestine, Olsenella uli, Oenococcus kitaharae, Bifidobacterium bifidum, Lactobacillus rhamnosus, Lactobacillus gasseri, Finegoldia magna, Mycoplasma mobile, Mycoplasma gallisepticum, Mycoplasma ovipneumoniae, Mycoplasma canis, Mycoplasma synoviae, Eubacterium rectale, Streptococcus thermophilus, Eubacterium dolichum, Lactobacillus coryniformis subsp.
- Torquens Ilyobacter polytropus, Ruminococcus albus, Akkermansia muciniphila, Acidothermus cellulolyticus, Bifidobacterium longum, Bifidobacterium dentium, Corynebacterium diphtheria, Elusimicrobium minutum, Nitratifractor salsuginis, Sphaerochaeta globus, Fibrobacter succinogenes subsp.
- the nucleic acid encoding the Cas protein that is part of the prime editor can be codon optimized for efficient translation into protein in a particular cell or organism.
- a Cas protein is a dead Cas protein.
- a dead Cas protein can be a protein that lacks nucleic acid cleavage activity.
- a dCas9 polypeptide can associate with a guide nucleic acid molecule (e.g., PEgRNA) to activate or repress transcription of target DNA.
- Guide nucleic acid molecules can be introduced into cells expressing the engineered chimeric receptor polypeptide. In some cases, such cells contain one or more different guide nucleic acid molecules that target the same nucleic acid. In other cases, the guide nucleic acid molecules target different nucleic acids in the cell.
- the nucleic acids targeted by the guide nucleic acid molecule can be any that are expressed in a cell such as an immune cell.
- the nucleic acids targeted can be a gene involved in immune cell regulation. In some embodiments, the nucleic acid is associated with cancer.
- Enzymatically inactive can refer to an activity less than 1%, less than 2%, less than 3%, less than 4%, less than 5%, less than 6%, less than 7%, less than 8%, less than 9%, or less than 10% activity compared to a wild-type exemplary activity (e.g., nucleic acid cleaving activity, wild-type Cas9 activity).
- a wild-type exemplary activity e.g., nucleic acid cleaving activity, wild-type Cas9 activity.
- One or a plurality of the nuclease domains (e.g. , RuvC, HNH) of a Cas protein can be deleted or mutated so that they are no longer functional or comprise reduced nuclease activity (e.g., deactivated or dead Cas, i.e., “dCas”).
- dCas deactivated or dead Cas
- nickase can generate a single-strand break at a CRISPR RNA (crRNA) recognition sequence within a doublestranded DNA but not a double-strand break.
- crRNA CRISPR RNA
- Such a nickase can cleave the complementary strand or the non-complementary strand, but may not cleave both.
- the resulting Cas protein can have a reduced or no ability to cleave both strands of a double -stranded DNA.
- An example of a mutation that can convert a Cas9 protein into a nickase is a D10A (aspartate to alanine at position 10 of Cas9) mutation in the RuvC domain of Cas9 from S. pyogenes.
- H939A (histidine to alanine at amino acid position 839) or H840A (histidine to alanine at amino acid position 840) in the HNH domain of Cas9 from S. pyogenes can convert the Cas9 into a nickase.
- An example of a mutation that can convert a Cas9 protein into a dead Cas9 is a D10A (aspartate to alanine at position 10 of Cas9) mutation in the RuvC domain and H939A (histidine to alanine at amino acid position 839) or H840A (histidine to alanine at amino acid position 840) in the HNH domain of Cas9 from S. pyogenes.
- a dead Cas protein can comprise one or more mutations relative to a wild-type version of the protein.
- the mutation can result in less than 90%, less than 80%, less than 70%, less than 60%, less than 50%, less than 40%, less than 30%, less than 20%, less than 10%, less than 5%, or less than 1% of the nucleic acid-cleaving activity in one or more of the plurality of nucleic acid-cleaving domains of the wildtype Cas protein.
- the mutation can result in one or more of the plurality of nucleic acid-cleaving domains retaining the ability to cleave the complementary strand of the target nucleic acid but reducing its ability to cleave the non-complementary strand of the target nucleic acid.
- the mutation can result in one or more of the plurality of nucleic acid-cleaving domains retaining the ability to cleave the non-complementary strand of the target nucleic acid but reducing its ability to cleave the complementary strand of the target nucleic acid.
- the mutation can result in one or more of the plurality of nucleic acid-cleaving domains lacking the ability to cleave the complementary strand and the non-complementary strand of the target nucleic acid.
- the residues to be mutated in a nuclease domain can correspond to one or more catalytic residues of the nuclease.
- residues in the wild type exemplary .S', pyogenes Cas9 polypeptide such as Asp 10, His840, Asn854 and Asn856 can be mutated to inactivate one or more of the plurality of nucleic acid-cleaving domains (e.g., nuclease domains).
- the residues to be mutated in a nuclease domain of a Cas protein can correspond to residues AsplO, His840, Asn854 and Asn856 in the wild type .S'. pyogenes Cas9 polypeptide, for example, as determined by sequence and/or structural alignment.
- residues DIO, G12, G17, E762, H840, N854, N863, H982, H983, A984, D986, and/or A987 can be mutated.
- residues DIO, G12, G17, E762, H840, N854, N863, H982, H983, A984, D986, and/or A987 can be mutated.
- D10A, G12A, G17A, E762A, H840A, N854A, N863A, H982A, H983A, A984A, and/or D986A can be suitable.
- a D10A mutation can be combined with one or more of H840A, N854A, or N856A mutations to produce a Cas9 protein substantially lacking DNA cleavage activity (e.g., a dead Cas9 protein).
- a H840A mutation can be combined with one or more of D10A, N854A, or N856A mutations to produce a site- directed polypeptide substantially lacking DNA cleavage activity.
- a N854A mutation can be combined with one or more of H840A, D10A, or N856A mutations to produce a site-directed polypeptide substantially lacking DNA cleavage activity.
- a N856A mutation can be combined with one or more of H840A, N854A, or D10A mutations to produce a site-directed polypeptide substantially lacking DNA cleavage activity.
- a dCas9 can be fused to other proteins.
- dCas9 can be fused to SunTag, KRAB, VPS4, P3000, VPR, VP64, V64-p65-Rta, VP160, VP192, HDAC1, DNMT3A, TET1, SPH, KRAB-MeCP2, epigenetic regulators, or other proteins.
- a dCas9 fusion comprises a ZIM3 KRAB-Cas9 fusion.
- a Cas9 fusion can be a paired dCas9 system.
- dCas9 can be part of a SAM system or REDMAP system.
- Cas9 variants and fusion proteins can be found in Li, T. et al,, Sig Transduct Target Ther 8, 36 (2023), which is incorporated in its entirety, [0173]
- a Cas protein is a Class 2 Cas protein.
- a Cas protein is a type II Cas protein.
- the Cas protein is a Cas9 protein, a modified version of a Cas9 protein, or derived from a Cas9 protein. For example, a Cas9 protein lacking cleavage activity.
- the Cas9 protein is a Cas9 protein from .S', pyogenes (e.g., SwissProt accession number Q99ZW2). In some embodiments, the Cas9 protein is a Cas9 from S.aureus (e.g., SwissProt accession number J7RUA5). In some embodiments, the Cas9 protein is a modified version of a Cas9 protein from .S', pyogenes or .S'. Aureus. In some embodiments, the Cas9 protein is derived from a Cas9 protein from .S'. pyogenes or .S'. Aureus. For example, a .S' pyogenes or .S' Aureus Cas9 protein lacking cleavage activity.
- Cas9 can generally refer to a polypeptide with at least about 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100% sequence identity and/or sequence similarity to a wild type exemplary Cas9 polypeptide (e.g. , Cas9 from .S' pyogenes).
- Cas9 can refer to a polypeptide with at most about 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100% sequence identity and/or sequence similarity to a wild type exemplary Cas9 polypeptide (e.g. , from .S' pyogenes).
- Cas9 can refer to the wildtype or a modified form of the Cas9 protein that can comprise an amino acid change such as a deletion, insertion, substitution, variant, mutation, fusion, chimera, or any combination thereof.
- a guidable polypeptide domain is a Cas9 or variant thereof.
- the Cas9 or variant thereof is a nuclease active Cas9 domain, a nuclease inactive Cas9 domain, or a Cas9 nickase domain or a variant thereof.
- a guidable polypeptide domain is Cas9, Casl2e, Casl2d, Casl2a, Casl2bl, Casl3a, Casl2c, or Argonaute (Ago domain), any of which optionally has a nickase activity.
- a guidable polypeptide domain comprises an amino acid sequence at least 80%, 85%, 90%, 95%, or 99% identical to any one of sequences listed in Table 8 below. In some embodiments, a guidable polypeptide domain comprises an amino acid sequence at least 80%, 85%, 90%, 95%, or 99% identical to any one of sequences set forth in SEQ ID NOs: 1000- 1020. In some cases, a guidable polypeptide domain is a Cas9 H840A nickase. In some cases, a guidable polypeptide domain is Cas9 D10A nickase. Cas9-H840A. In some cases, a guidable polypeptide domain is a Casl2a/b nickase.
- the payload in the present disclosure comprises a polymerase (e.g., reverse transcriptase).
- a polymerase can comprise a natural or an engineered domain.
- a polymerase can be capable of synthesizing nucleic acids.
- a polymerase can be a DNA polymerase or an RNA polymerase.
- a polymerase is a reverse transcriptase.
- a reverse transcriptase can synthesize DNA from deoxyribonucleotides.
- a reverse transcriptase adds deoxyribonucleotides to the 3’ end of a nucleic acid primer to synthesize DNA.
- a reverse transcriptase uses an RNA template and uses base-pairing interactions to synthesize a DNA strand that is complementary to the RNA template.
- the reverse transcriptase domain can be a reverse transcriptase from any organism, phage, virus, or an engineered or mutated variant.
- the reverse transcriptase domain can be a reverse transcriptase derived from or sharing structural or sequencing similarity to a reverse transcriptase in a CRISPR system.
- the reverse transcriptase can be an M-MLV or HIV reverse transcriptase.
- the reverse transcriptase can be a human LINE-1 reverse transcriptase or a group II intron reverse transcriptase.
- the reverse transcriptase can be a human endogenous retrovirus reverse transcriptase.
- a nucleic-acid modifying effector or a nucleic acid-binding moiety comprises a transposase domain.
- a transposase domain can be a natural or an engineered domain.
- a transposase domain can be capable of aiding the translocation of a transposable element, a nucleic acid sequence that can change its position within a genome.
- a transposase domain comprises a TnsA, a TnsB, a TnsC, or a TnsD domain.
- a transposase domain comprises a TniQ domain.
- a transposase domain is derived from or shares sequence or structural similarity with a transposase in a CRISPR system (e.g., a CRISPR-associated transposase).
- a transposase domain is derived from or share sequence or structural similarity with a transposase domain from a type I CRISPR- associated transposon (CAST) system.
- transposase domain is derived from or share sequence or structural similarity with a transposase domain from a type V CRISPR-associated transposon (CAST) system.
- a transposase domain can be capable of binding to a guidable polypeptide domain.
- a transposase domain is coupled to a guidable polypeptide domain.
- a transposase domain is capable of binding to a type I CRISPR-Cas domain (e.g., a Cascade domain, a Cas8 domain, or a Cas5 domain).
- a transposase domain is capable of binding to a type V CRISPR-Cas domain (e.g., a Casl2 domain).
- a transposase domain is capable of mediating targeted insertion of a nucleic acid into a target nucleic acid.
- a transposase domain is capable of mediating targeted insertion of a nucleic acid that is at least 5 kb, at least 6 kb, at least 7 kb, at least 8kb, at least 9kb, at least lOkb, at least 1 Ikb, at least 12kb, at least 13kb, at least 14kb, or at least 15 kb into a target nucleic acid.
- a transcriptional activation domain can be or comprise a CAP domain, a VP64 domain, a p65 domain, an Rta domain, a synergistic activation mediator (SAM) domain, a SunTag domain, a VPR domain, a DNA demethylase domain, a histone methyltransferase domain, a histone acetyltransferase domain, or a histone demethylase domain.
- SAM synergistic activation mediator
- a two handed zinc finger domain can bind to two noncontiguous target DNA sequences.
- the spacing between the two noncontiguous target sequences comprises from 1 to 15, from 1 to 12, from 1 to 10, from 1 to 8, or from 1 to 5 nucleotides.
- a two handed type of zinc finger binding protein can be SIP1.
- a cluster of zinc finger domains in a two handed zinc finger domain can be capable of binding to a unique target nucleic acid sequence.
- the payload in the present disclosure comprises a fusion protein.
- a fusion protein can comprise two or more polypeptide domains of any of the polypeptide domains described elsewhere herein.
- a fusion protein can be a natural or an engineered fusion protein.
- the two or more polypeptide domains are coupled together.
- the two or more polypeptide domains can be coupled together directly or coupled together indirectly.
- a first polypeptide domain can be coupled directly to a second polypeptide domain.
- the first polypeptide domain can be coupled indirectly to the second polypeptide domain by coupling with a third polypeptide domain that is coupled directly to the second polypeptide domain.
- a first polypeptide domain is coupled to the N-terminus of a second polypeptide domain. In some cases, a first polypeptide domain is coupled to the C-terminus of a second polypeptide domain. In some cases, a first polypeptide domain is coupled to an internal component of a second polypeptide domain. In some cases, the two or more polypeptide domains are covalently linked. In some cases, the two or more polypeptide domains are noncovalently linked. In some cases, the two or more polypeptide domains are coupled together by a linker. For example, a linker may be a peptide linker. A linker can be a rigid linker, which helps maintain a fixed distance between the polypeptide domains that it links.
- a linker can be a flexible linker, which can allow some flexibility in movement of one polypeptide domain relative to the other polypeptide domain that it is linked to.
- a linker is a cleavable linker.
- a cleavable linker can comprise a disulfide bond.
- a cleavable linker can be an enzymatic cleavable linker, e.g., a linker comprising a protease cleavage site.
- the fusion protein comprises a guidable polypeptide domain (e.g., a CRISPR domain) coupled to a reverse transcriptase domain.
- a guidable polypeptide domain (e.g., a CRISPR domain) coupled to a reverse transcriptase can be capable of enabling prime editing.
- a prime editor can be capable of editing a nucleic acid sequence in a target nucleic acid molecule.
- a prime editor can be capable of mediating insertion or deletion of a nucleic acid sequence in a target nucleic acid molecule.
- the prime editor enables a sequence insertion or sequence deletion without introducing a double -stranded break.
- the prime editor introduces a nick at the target site.
- the prime editor can enable insertion of a template sequence in a target nucleic acid molecule.
- the template sequence can comprise the desired edit.
- a prime editor reverse transcribes a template sequence to synthesize a complementary strand.
- the synthesized complementary strand is inserted in the target nucleic acid molecule.
- the prime editor uses a primer to carry out reverse transcription.
- the prime editor can install nucleotides to the 3’ end of a primer strand.
- a primer strand is generated by nicking the target nucleic acid molecule.
- guidable polypeptide domain (e.g., a CRISPR domain) is coupled to a transcriptional regulatory domain.
- a guidable polypeptide domain (e.g., a CRISPR domain) coupled to a transcriptional regulatory domain can be capable of enabling CRISPR interference (CRISPRi) or CRISPR activation (CRISPRa).
- CRISPRi CRISPR interference
- CRISPRa CRISPR activation
- a guidable polypeptide domain (e.g., a CRISPR domain) can be coupled to a transcriptional regulatory domain such as P3000 or DNMT3.
- a guidable polypeptide domain (e.g., a CRISPR domain) is coupled to the C-terminus of a transcriptional regulatory domain.
- a guidable polypeptide domain (e.g., a CRISPR domain) is coupled to an internal component of a transcriptional regulatory domain.
- Any of the payloads described herein can further comprise a plasma membrane recruitment domain, transmembrane domain, a signaling domain, a receptor domain, a packaging domain, or a targeting domain.
- Any of payloads described herein can comprise or be engineered to comprise a protein tag, a peptide tag, or small molecule tag.
- a payload can comprise a small nuclear localization signal (NLS), a nuclear export signal (NES), a cell penetrating peptide (CPP), a mitochondria penetrating peptide (MPP), a solubility tag, or a fluorescent tag.
- the payload to be delivered by the lipid containing particles of the present disclosure comprises a nucleobase editor (also termed as “base editor”) or one or more components of a nucleobase editing (also termed as “base editing”) complex.
- base editor can refer to an agent comprising a polypeptide that is capable of making a modification to a base (e.g., A, T, C, G, or U) within a nucleic acid sequence (e.g., DNA or RNA).
- the base editor is capable of deaminating a base within a nucleic acid.
- the base editor is capable of deaminating a base within a DNA molecule.
- the base editor is capable of deaminating an adenosine (A) in DNA.
- the base editor is capable of deaminating a cytosine (C) in DNA.
- the base editor is capable of converting a guanine (G) in DNA through a glycoylase.
- the payload in the present disclosure comprises a deaminase domain.
- the deaminase domain can be a natural or an engineered domain.
- a deaminase domain can be capable of carrying out deamination reactions in DNA.
- a deaminase domain can be capable of enabling the generation of base conversions or point mutations in a target nucleic acid.
- a deaminase domain can be a cytidine deaminase domain or an adenosine deaminase domain.
- a cytidine deaminase domain can be capable of converting cytosine to uracil.
- a cytidine deaminase domain can be capable of enabling the conversion of a C-G base pair to a T-A base pair.
- a cytidine deaminase can be or comprise a APOBEC1 cytidine deaminase.
- An adenosine deaminase domain can be capable of converting an adenosine to hypoxanthine.
- An adenosine deaminase domain can be capable of converting an adenosine to an inosine.
- An adenosine deaminase can comprise TadA or a TadA mutant. In some embodiments, TadA comprises a monomer.
- TadA comprises a heterodimer comprising a wildtype TadA and a mutated Tad A. In some embodiments, TadA comprises a homodimer comprising two wildtype TadA domains or two mutated TadA domains.
- An adenosine deaminase domain can be capable of enabling the conversion of an A-T base pair to a G-C base pair.
- a deaminase domain can be a mutated variant. In some cases, a deaminase domain enables the conversion of C to G, A to I, or C to U.
- the payload in the present disclosure comprises a glycosylase domain.
- the glycosylase domain can be a natural or an engineered domain.
- a glycosylase-based guanine base editor can be designed to remove G, and the AP site generated is repaired by translesion synthesis and/or DNA replication, leading to G-to-C or G-to-T conversion.
- a glycosylase domain can be capable of enabling the generation of base conversions or point mutations in a target nucleic acid.
- a glycosylase domain can be a guanine glycosylase domain. Examples of glycosylase base edits can be found in Sun N, et al.. Mol Ther. 2022 Jul 6;30(7):2452-2463 and Huawei Tong, et al., National Science Review, Volume 10, Issue 8, August 2023, each of which is incorporated in its entirety herein.
- the base editor disclosed herein comprises a deaminase or a functional domain thereof (“deaminase domain”) that catalyzes deamination reaction.
- deaminase or “deaminase domain,” as used herein, refers to a protein or enzyme that catalyzes a deamination reaction.
- the deaminase or deaminase domain is an adenosine deaminase, catalyzing the deamination of adenosine, converting it to the nucleoside hypoxanthine.
- the deaminase or deaminase domain is a cytidine deaminase, catalyzing the hydrolytic deamination of cytidine or deoxycytidine to uridine or deoxyuridine, respectively.
- the deaminase or deaminase domain is a cytidine deaminase domain, catalyzing the hydrolytic deamination of cytosine to uracil.
- the deaminase or deaminase domain is a naturally-occurring deaminase from an organism, such as a human, chimpanzee, gorilla, monkey, cow, dog, rat, or mouse.
- the deaminase or deaminase domain is a variant of a naturally-occurring deaminase from an organism, that does not occur in nature.
- the deaminase or deaminase domain is at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75% at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or at least 99.5% identical to a naturally-occurring deaminase from an organism.
- an “adenosine deaminase” is an enzyme that catalyzes the deamination of adenosine, converting it to the nucleoside hypoxanthine. Under standard Watson-Crick hydrogen bond pairing, an adenosine base hydrogen bonds to a thymine base (or a uracil in case of RNA).
- the base editor is a chimeric protein comprising a nucleic acid programmable R/DNA binding protein (napR/DNAbp) fused to a deaminase (e.g., cytidine deaminase or adenosine deaminase) domain.
- a deaminase e.g., cytidine deaminase or adenosine deaminase
- nucleic acid programmable D/RNA binding protein refers to any protein that can associate (e.g., form a complex) with one or more nucleic acid molecules (i.e., which can broadly be referred to as a “napR/DNAbp-programming nucleic acid molecule” and includes, for example, guide RNA in the case of Cas systems) which direct or otherwise program the protein to localize to a specific target nucleotide sequence (e.g., a gene locus of a genome, or an RNA molecule) that is complementary to the one or more nucleic acid molecules (or a portion or region thereof) associated with the protein, thereby causing the protein to bind to the nucleotide sequence at the specific target site.
- a specific target nucleotide sequence e.g., a gene locus of a genome, or an RNA molecule
- napR/DNAbp embraces CRISPR Cas9 proteins, as well as Cas9 equivalents, homologs, orthologs, or paralogs, whether naturally occurring or non-naturally occurring (e.g., engineered or recombinant), and can include a Cas9 equivalent from any type of CRISPR system (e.g., type II, V, VI), including Cpfl (a type-V CRISPR-Cas systems), C2cl (a type V CRISPR-Cas system), C2c2 (a type VI CRISPR-Cas system) and C2c3 (a type V CRISPR-Cas system).
- Cpfl a type-V CRISPR-Cas systems
- C2cl a type V CRISPR-Cas system
- C2c2 a type VI CRISPR-Cas system
- C2c3 a type V CRISPR-Cas system
- C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector,” Science 2016; 353(6299), the contents of which are incorporated herein by reference.
- the nucleic acid programmable R/DNA binding protein (napR/DNAbp) that can be used in connection with this disclosure are not limited to CRISPR-Cas systems.
- the present disclosure embraces any such programmable protein, such as the Argonaute protein from Natronobacterium gregoryi (NgAgo) which can also be used for DNA-guided genome editing.
- NgAgo-guide DNA system does not require a PAM sequence or guide RNA molecules, which means genome editing can be performed simply by the expression of generic NgAgo protein and introduction of synthetic oligonucleotides on any genomic sequence. See Gao F, Shen X Z, Jiang F, Wu Y, Han C. DNA- guided genome editing using the Natronobacterium gregoryi Argonaute. Nat Biotechnol 2016; 34(7):768- 73, which is incorporated herein by reference.
- the napR/DNAbp is derived from a nuclease disclosed herein, such as, Cas9 (e.g., dCas9 and nCas9), CasX, CasY, Casl4, Cpfl, C2cl, C2c2, C2c3, Argonaute protein, or a variant thereof.
- Cas9 e.g., dCas9 and nCas9
- CasX e.g., CasX, CasY, Casl4, Cpfl, C2cl, C2c2, C2c3, Argonaute protein, or a variant thereof.
- the base editor comprises a Cas9 (e.g., dCas9 and nCas9), CasX, CasY, Cpfl, C2cl, C2c2, C2c3, or Argonaute protein fused to a deaminase (e.g, cytidine deaminase or adenosine deaminase).
- a deaminase e.g, cytidine deaminase or adenosine deaminase
- the base editor comprises a Cas9 nickase (nCas9) fused to an deaminase (e.g., cytidine deaminase or adenosine deaminase).
- the base editor comprises a CasX protein fused to a deaminase (e.g, cytidine deaminase or adenosine deaminase).
- the base editor comprises a nuclease-inactive Cas9 (dCas9) fused to a deaminase (e.g., cytidine deaminase or adenosine deaminase).
- the base editor comprises a CasY protein fused to a deaminase (e.g, cytidine deaminase or adenosine deaminase).
- the DNA-synthesis template can comprise a desired edit to be introduced in a target nucleic acid molecule.
- the DNA-synthesis template can be a template for a DNA polymerase or a reverse transcriptase to carry out DNA synthesis.
- a prime editor can use the DNA synthesis template sequence to synthesize a DNA strand that is complementary to the DNA synthesis template sequence.
- the DNA strand is inserted into the target nucleic acid.
- the nucleic acid comprising the DNA synthesis template sequence also comprises a primer-binding sequence.
- a primer-binding sequence can be complementary to a sequence in a primer strand to which a DNA polymerase or reverse transcriptase can add nucleotides.
- the primer strand is part of a target nucleic acid molecule.
- a primer-binding sequence can be complementary to a sequence in the target nucleic acid.
- the payload comprises a polynucleotide that is configured to bind to a guidable polypeptide domain.
- the polynucleotide directs a guidable polypeptide domain to a sequence in a target nucleic acid molecule.
- the polynucleotide comprises a scaffold segment configured to bind to a guidable polypeptide domain (e.g., Cas9 or Casl2).
- polynucleotide comprises a spacer sequence that is complementary to a target sequence in the target nucleic acid molecule and is capable of hybridizing to the target sequence.
- a guide RNA comprises (A) a primer binding site, (B) a clamp segment, (C) a sequence encoding new genetic information that replaces the targeted sequence, (D) an aptamer, (E) spacer, or (F) scaffold, or any combinations thereof.
- a guide RNA comprises a sequence encoding new genetic information that replaces the targeted sequence, a spacer, and scaffold.
- a guide RNA comprises a spacer and scaffold.
- the guide nucleic acid molecule is heterologous to the cell or host receiving the lipid delivery particle.
- the PEgRNA comprises an extended single guide RNA (sgRNA) containing a primer binding site (PBS) and a template sequence for nucleic acid polymerase (e.g., reverse transcriptase or DNA polymerase).
- a PEgRNA can comprise an architecture corresponding to 5'-[spacer]- [guide RNA core] -[extension arm]-3'.
- the spacer sequence can comprise about 20 nucleotides in length.
- the spacer sequence can bind to a protospacer in a target nucleic acid molecule.
- the spacer sequence can guide the nucleic acid-guided polypeptide (e.g., Cas9) to the target nucleic acid molecule.
- the PEgRNA comprises an aptamer and the prime editor further comprises an aptamer binding protein (e.g. , fused to Cas protein or reverse transcriptase).
- Guide RNAs including an aptamer include those described in International Publication No. W02023205708, which is hereby incorporated herein by reference in its entirety.
- Homology arm can encode a portion of a resulting reverse transcriptase-encoded single strand DNA flap to be integrated into the target DNA site by replacing the endogenous strand.
- the portion of the single strand DNA flap encoded by the homology arm is complementary to the non-edited strand of the target DNA sequence, which facilitates the displacement of the endogenous strand and annealing of the single strand DNA flap in its place, thereby installing the edit.
- the edit template can comprise a sequence corresponding to new genetic information that replaces the targeted sequence, i.e., a single strand RNA of the PEgRNA that codes for a complementary single strand DNA that is either the sense or the antisense strand of the new genetic information that replaces the targeted sequence and which is incorporated into the genomic DNA target locus through the prime editing process.
- the primer binding site allows the 3 ’ end of the nicked DNA strand to hybridize to the PEgRNA, while the reverse transcriptase template serves as a template for the synthesis of edited genetic information.
- a prime editing system can allow DNA synthesis based on the reverse transcriptase template at a nick site a single 3' flap, which becomes integrated into a target nucleic acid on the same strand.
- a prime editing system can be a multi-flap prime editing system that generate pairs or multiple pairs of 3' flaps on different strands, which form duplexes comprising desired edits and which become incorporated into target nucleic acid molecules, e.g., at specific loci or edit sites in a genome.
- the pairs or multiple pairs of 3' flaps form duplexes because they comprise reverse complementary sequences which anneal to one another once generated by the prime editors described herein.
- the duplexes can be incorporated into the target site by cell -driven mechanisms that naturally replace the endogenous duplex sequences located between adjacent nick sites.
- the new duplex sequences can be introduced at one or more locations (e.g., at adjacent genomic loci or on two different chromosomal locations), and can comprise one or more sequences of interest, e.g., protein-encoding sequence, peptide-encoding sequence, or RNA-encoding sequence.
- the payload comprises a polynucleotide comprising a scaffold segment, a spacer sequence, a DNA synthesis template, and a primer-binding sequence.
- the scaffold segment and a spacer sequence are on a first nucleic acid molecule and the DNA synthesis template and the primer-binding sequence are on a second nucleic acid molecule.
- the guide RNA further comprises a clamp segment.
- the guide RNA comprising, from 3’ to 5’, a primer binding site, a sequence encoding at least a portion of the first recombinase recognition sequence, a clamp segment, scaffold, and spacer.
- the clamp segment comprises a sequence that, after being reverse transcribed is at least partially complementary to a genomic site close to the primer binding site and where the spacer binds.
- the clamp segment can enhance integration efficiency of the new genetic material that replaces the target sequence at the double-stranded target DNA sequence relative to a guide RNA without the clamp segment.
- the clamp segment can allow for a reduced number of nucleotides in the primer binding site need to bind its genomic site and facilitate reverse transcription, which in turn enables design of a guide RNA that is shorter than conventional guide RNAs used for other gene editing methods.
- the clamp segment is described in International Publication No. WO2023215831, which is hereby incorporated herein by reference in its entirety.
- a guide RNA can complete the insertion of new genetic material that replaces the target sequence without another guide RNA when delivered to a cell together with a prime editor described herein.
- the guide RNA can complete the insertion of the new genetic material that replaces the target sequence with a second guide RNA that is a nicking guide RNA when delivered together with a prime editor described herein.
- a guide RNA comprises two or more guide RNAs.
- the two or more guide RNAs comprise a first guide RNA encoding at least a first portion of new genetic material that replaces the target sequence.
- the two or more guide RNAs comprise a second guide RNA encoding at least a second portion of the new genetic material that replaces the target sequence.
- the first guide RNA and the second guide RNA work in a pair and collectively encode the new genetic material that replaces the target sequence, thereby inserting the new genetic material that replaces the target sequence into the genome of a cell receiving the lipid delivery particles in a site-specific manner.
- the first and the second portion of the new genetic material that replaces the target sequence have at least 6bp overlap. In some cases, the first portion of the new genetic material that replaces the target sequence is 46 bp. In some cases, the first portion of the new genetic material that replaces the target sequence is 42 bp. In some cases, the first portion or the second portion of the new genetic material that replaces the target sequence is 36 bp, 38 bp, 40 bp, 42 bp, 44 bp, or 46 bp.
- the first guide RNA comprises a first spacer.
- the second guide RNA comprises a second spacer. The first spacer and the second spacer bind to two genomic target sites that are within 5-100 bp from each other.
- the double strand DNA between the two genomic target sites are deleted and the full sequence of the new genetic material that replaces the target sequence is inserted instead.
- the deletion can be mediated by the following steps: (a) reverse transcription of the sequence encoding the first portion of the new genetic material that replaces the target sequence in the first guide RNA and the sequence encoding the second portion of the new genetic material that replaces the target sequence in the second guide RNA, wherein the first and the second portion of the new genetic material that replaces the target sequence having at least 6bp overlap,
- the payload comprises a polypeptide domain described herein coupled to a polynucleotide domain described herein.
- the payload comprises a polypeptide domain described herein complexed to a polynucleotide domain described herein.
- the payload comprises a ribonucleoprotein.
- the payload may comprise a guidable polypeptide domain complexed to a polynucleotide configured to bind to the guidable polypeptide domain (e.g., Cas9 complexed with an RNA guide).
- Any of the payloads described herein can further comprise a plasma membrane recruitment element, a transmembrane domain, a signaling domain, a receptor domain, a packaging domain, or a targeting domain.
- Any of payloads described herein can comprise or be engineered to comprise a protein tag, a peptide tag, or small molecule tag.
- a payload can comprise a nuclear localization signal (NLS), a nuclear export signal (NES), a cell penetrating peptide (CPP), a mitochondria penetrating peptide (MPP), a solubility tag, a fluorescent tag, or any combinations thereof.
- the payload in the lipid delivery particle of the present disclosure comprises a recombinant protein.
- the payload can be a diagnostic imaging agent, such as a contrast agent.
- the payload comprises a therapeutic agent, including, but not limited to, a nuclease, a recombinase, a growth factor, an antibody, a chimeric antigen receptor, a T cell receptor, a cytokine, a cytokine inhibitor or agonist, a transcription factor, an organelle, a nucleic acid molecule, a therapeutic DNA, a therapeutic RNA, a retrotransposon, a reverse transcriptase, an oligonucleotide, an aptazyme, an aptamer, or a ribozyme, a generic or specific kinase inhibitor, a small molecule drug, an immunomodulator, a tumor suppressor, a developmental regulator, a cancer vaccine, an anesthetic, an enzyme, a
- the payload can be a prophylactic agent.
- the payload comprises a biomarker.
- the payload can also comprise an exogenous antigen or an enzyme.
- the payload comprises a metabolite molecule.
- the payload comprises a lipid molecule.
- the payload comprises a structural protein.
- the payload comprises a hormone or a hormonal protein.
- composition, methods of production, methods of purification related to the lipid delivery particles provided herein.
- the lipid delivery particles can be produced from producer cell lines that are either transiently transfected with at least one plasmid or stably expressing constructs that have been integrated into the producer cell line genomic DNA.
- Producer cell lines can be generated by stably integrating genetic material with a gene of interest into a host cell line.
- the genetic material is transiently expressed in a producer cell line.
- the genetic material is expressed via viral methods.
- the genetic material is expressed via non-viral methods.
- a producer cell line grows in a serum-free medium or in suspension.
- a producer cell line can be grown in serum-free medium and suspension simultaneously.
- producer cell lines can be generated with adherent cells (e.g., cells cultured in media and attached to a substrate).
- Producer cells can be used to produce the lipid delivery particles described herein.
- generating a producer cell line comprises transfecting cells (e.g., cells of a mammalian cell type) with genetic material of the present disclosure, culturing the cells to produce the lipid delivery particles, obtaining a media from the mammalian cell producing the lipid delivery particles, collecting and filtering the harvested media, and, optionally, purifying the lipid delivery particles to retain structural integrity.
- the method of producing the lipid delivery particle further comprises providing new media to promote transient production of the lipid delivery particles.
- the mammalian cell type includes a HT1080 cell, a COS cell, a HeLa cell, a Chinese Hamster Ovary (CHO) cell, or a HEK 293 cell.
- HEK293 cells are cells derived from human embryonic kidney cells grown in tissue culture.
- the HEK293 cell is a HEK293, 293E, 293T, 293F, 293FT, or 293T Gesicle cell.
- the producer cell line can be transformed with a viral vector or non-viral method in any number of means including calcium phosphate and the like.
- the cells can be cultured under conditions for production of lipid delivery particles.
- Exemplary culturing conditions can include refeeding cells in appropriate media, addition of CO2, and humidity.
- culturing conditions includes addition of antibiotics, anti-fungals, and/or growth factors.
- the medium can be harvested after 24, 48, 72, or 96 hours, or at any appropriate time point to allow sufficient production of the lipid delivery particles.
- the lipid delivery particles in the media can be isolated and collected using any number of techniques known in the art.
- the lipid delivery particles are purified, wherein the lipid delivery particles are washed or resuspended in an appropriate buffer or media or at particular concentration.
- disclosed herein are methods of manufacturing producer cell lines that comprise the lipid delivery particles of the present disclosure.
- Adherent cells can be first transfected to produce lipid delivery particles.
- transfection occurs by the addition or expression of exogenous nucleic acid sequences via non-viral methods (e.g., by electroporation, microinjection, or a chemical system such as DEAE-dextran or cationic polymers).
- transfection occurs by the addition or expression of exogenous nucleic acid sequences via viral methods (e.g. , by infecting the cells with a viral vector, such as an adenoviral vector, adeno-associate viral vector, a lentiviral vector, a herpes viral vector, or a HSV vector).
- a viral vector such as an adenoviral vector, adeno-associate viral vector, a lentiviral vector, a herpes viral vector, or a HSV vector.
- the cells are from a HEK293 cell line (e.g., HEK293, 293E, or 293T).
- the cells are cultured in a medium.
- cells can be cultured in the medium for 5, 10, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24 or 25 hours.
- cells can be cultured in the medium for between 10-20 hours.
- cells can be cultured in the medium for 18 hours.
- the new solution is new media.
- the new media promotes the production of the lipid delivery particles.
- the cells incorporate into the new media for between 10-50 hours.
- the cells incorporate into the new media for 10, 20, 30, 35, 40, 45, or 50 hours.
- the cells incorporate into the new media for 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39 or 40 hours.
- the media can then be harvested.
- the harvested media can be filtered, and the lipid delivery particles can be collected. Filtration can comprise microfiltration and/or depth filtration.
- the lipid delivery particles can undergo further purification and/or concentration methods that maintain the structural integrity of the particles.
- RNA and protein from a producer cell can get packaged and/or incorporated into lipid delivery vehicles of the present disclosure.
- the components of the lipid delivery particles, such as a payload is loaded via the packaging and assembly process of the lipid delivery particle.
- the payload can be a polypeptide or protein that is packaged into the lipid delivery particle as a part of a chimeric protein as disclosed herein.
- the payload is assembled into the lipid delivery particle as an independent entity, e.g., not as a part of a chimeric protein.
- the lipid delivery particle provided herein is loaded with a payload by utilizing any suitable method for delivering a biological or chemical payload through a lipid membrane, such as nucleofection, electroporation, lipid-based, polymer-based, or CaCF transfection, sonication, freeze thaw, incubation at various temperatures, or heat shock of lipid delivery particles mixed with payload.
- the nucleic acid molecules such as a template RNA described herein, are loaded into the lipid delivery particle by direct loading, such as electroporation of the lipid delivery particle in vitro.
- the nucleic acid molecules are loaded into the lipid delivery particle by binding to a nucleic acid binding protein (e.g., Cas protein) that is part of the lipid delivery particle or is already loaded into the lipid delivery particle.
- a nucleic acid binding protein e.g., Cas protein
- a first payload is a polypeptide that is assembled into the lipid delivery particle as a part of a chimeric protein
- a second payload is a separate protein or nucleic acid (RNA or DNA) that interacts with (e.g., binds) the first payload, and thus is loaded into the lipid delivery particle via the interaction between the first payload and the second payload.
- the second payload can be loaded into the lipid delivery particle via a transfection-like technique or any other suitable method.
- lipid delivery particle or pharmaceutical composition comprising contacting a cell with the lipid delivery particle described herein.
- the cell is a mammalian cell, such as a human cell.
- the cell is within a subject in need of treatment for a disease or a condition.
- contact comprising administering the lipid delivery particle described herein to the subject, such as via injections.
- the method comprises administering the lipid delivery particle, system, or pharmaceutical composition described herein to a subject in need thereof, such as via injections.
- the method comprises contacting a producer cell with compositions described herein.
- the lipid delivery particles are produced from producer cell lines that are either transiently transfected with at least one plasmid or stably expressing constructs that have been integrated into the producer cell line genomic DNA.
- the producer cell culture medium is harvested 24-, 48-, 72-, or 96-hours post-transfection.
- the producer cell culture medium is harvested between 40- and 48-hours post-transfection. The harvested medium can undergo centrifugation steps to remove producer cell debris while maintaining the structural integrity of the lipid delivery particle.
- the producer cell medium is centrifuged, e.g., at 500g for 5 minutes.
- the clarified lipid delivery particle containing supernatant can then be collected and filtered.
- the lipid delivery particles are further concentrated.
- the lipid delivery particles are further concentrated by ultracentrifugation.
- the lipid delivery particles are concentrated 50-fold, 100-fold, 200- fold, 500-fold, 1000-fold, 2000-fold, 3000-fold, or 5000-fold.
- the concentrated lipid delivery particles are resuspended, e.g., in cold PBS.
- the concentrated lipid delivery particles are frozen, e.g., frozen at a rate of -l°C/min and stored at -80°C.
- the purification methods can comprise chromatographic methods (e.g., anion exchange chromatography), ultrafiltration methods (e.g, tangential flow filtration), clarifying normal flow filtration, and/or sterilizing membrane filtration.
- chromatographic methods e.g., anion exchange chromatography
- ultrafiltration methods e.g, tangential flow filtration
- clarifying normal flow filtration e.g., tangential flow filtration
- sterilizing membrane filtration e.g., chromatographic methods
- Anion exchange chromatography can separate substances based on net-surface charge, using an ion-exchange resin.
- Tangential flow filtration can separate molecules using ultrafiltration membranes.
- the membrane pore size used for tangential flow filtration can retain a biological product of a size less than 1000 kDa, less than 750 kDa, less than 500 kDa, less than 250 kDa, less than 200 kDa, less than 150 kDa, less than 100 kDa, or less than 50 kDa.
- Normal flow filtration assists in the clarification of biofluid by convecting the substance directly toward a membrane under an applied pressure.
- compositions or systems can be used for producing a lipid delivery particle of the present disclosure, for instance, by transfecting or otherwise delivering the nucleic acid molecules in the compositions or systems into a producer cell.
- the nucleic acid molecules can be expressed in the producer cell, the result of which assemble, package, and subsequently cause the producer cell to release the lipid delivery particle.
- a lipid delivery particle of the present disclosure facilitates gene editing efficiency comprising 8-fold increase of prime editing efficiency when compared to a conventional VLP.
- a lipid delivery particle of the present disclosure exhibits reduced immunogenicity in transduced target cells.
- a lipid delivery particle of the present disclosure produces reduced off-target genome editing in target cells when delivering genome editing system into the target cells when compared to a conventional VLP.
- a lipid delivery particle of the present disclosure leads to more than 100-fold reduction in Cas-independent off-target editing when compared to a conventional VLP.
- a lipid delivery particle of the present disclosure leads to at least 10-fold, such as 12- to 900-fold, lower Cas-dependent off-target editing when compared to a conventional VLP.
- a pharmaceutical formulation comprising the lipid delivery particle disclosed herein and optionally further comprising a pharmaceutically acceptable carrier, excipient, or additive.
- pharmaceutical formulation refers to a composition formulated for pharmaceutical use.
- pharmaceutical formulations comprise an immunologically effective amount of one or more cells, vectors, lipid delivery particles, or compositions disclosed herein, and optionally one or more other components which are pharmaceutically acceptable.
- the pharmaceutical formulation comprises additional agents, e.g., for specific delivery, increasing half-life, or other therapeutic benefit.
- the pharmaceutical formulation may comprise one or more of dimethylsulfoxide (DMSO), dextrose, water, succinate, poly I: poly C, poly-L- lysine, carboxymethylcellulose, and/or chloride.
- DMSO dimethylsulfoxide
- a “pharmaceutically acceptable carrier” is an agent that is compatible with the other ingredients of the formulation and not injurious to the tissue of the subject (e.g., physiologically compatible, sterile, physiologic pH, etc.)
- a pharmaceutically acceptable carrier comprises any vehicle, such as a liquid or solid fdler, diluent, excipient, manufacturing aid (e.g., lubricant, talc magnesium, calcium or zinc stearate, or steric acid), or solvent encapsulating material, involved in carrying or transporting the compound from one site (e.g. , the delivery site) of the body, to another site (e.g., organ, tissue or portion of the body).
- Some exemplary materials which can serve as pharmaceutically-acceptable carriers include: (1) sugars, such as lactose, glucose and sucrose; (2) starches, such as com starch and potato starch; (3) cellulose, and its derivatives, such as sodium carboxymethyl cellulose, methylcellulose, ethyl cellulose, microcrystalline cellulose and cellulose acetate; (4) powdered tragacanthin; (5) malt; (6) gelatin; (7) lubricating agents, such as magnesium stearate, sodium lauryl sulfate and talc; (8) excipients, such as cocoa butter and suppository waxes; (9) oils, such as peanut oil, cottonseed oil, safflower oil, sesame oil, olive oil, com oil and soybean oil; (10) glycols, such as propylene glycol; (11) polyols, such as glycerin, sorbitol, mannitol and polyethylene glycol (PEG); (12) esters,
- Pharmaceutical formulation disclosed herein can comprise one or more pH buffering compounds to maintain the pH of the formulation at a predetermined level that reflects physiological pH, such as in the range of about 5.0 to about 8.0.
- the pH of the pharmaceutical formulation can be about 4, about 5, about 6, about 7, about 8 or about 9.
- the pH buffering compound used in the aqueous liquid formulation can be an amino acid or mixture of amino acids, such as histidine or a mixture of amino acids such as histidine and glycine.
- the pH buffering compound can be an agent which does not chelate calcium ions.
- Exemplary pH buffering compounds include imidazole and acetate ions.
- the pH buffering compound can be present in any amount suitable to maintain the pH of the formulation at a predetermined level.
- compositions described herein can be prepared by any method known or hereafter developed in the art of pharmacology.
- preparatory methods include the step of bringing the active ingredient(s) into association with an excipient and/or one or more other accessory ingredients, and then, optionally, shaping and/or packaging the product into a desired single- or multidose unit.
- compositions can additionally comprise a pharmaceutically acceptable excipient, which, as used herein, includes any and all solvents, dispersion media, diluents, or other liquid vehicles, dispersion or suspension aids, surface active agents, isotonic agents, thickening or emulsifying agents, preservatives, solid binders, lubricants, and the like, as suited to the particular dosage form desired.
- a pharmaceutically acceptable excipient includes any and all solvents, dispersion media, diluents, or other liquid vehicles, dispersion or suspension aids, surface active agents, isotonic agents, thickening or emulsifying agents, preservatives, solid binders, lubricants, and the like, as suited to the particular dosage form desired.
- a lipid delivery particle provided herein can find use in a variety of fields and methods.
- the lipid delivery particle of the present disclosure can be used to deliver one or more payloads, such as a ribonucleoprotein complex to a cell.
- the target cells to which the lipid delivery particles are delivered are in vitro cells, ex vivo cells, or in vivo cells.
- the lipid delivery particles of the present disclosure can be applicable for delivery of freights into a variety of cell types, such as, animal cells, plant cells, bacteria cells, algal cells, or fungal cells.
- lipid delivery particle described herein a system described herein, a composition described herein, or pharmaceutical composition according to some embodiments of the present disclosure.
- the present disclosure provides methods of treating, preventing, or diagnosing a condition, disease, or disorder.
- a composition, kit, or method described herein can be used to treat, prevent, or diagnose a condition, disease, or disorder.
- the condition, disease, or disorder can comprise a cancer, an immune disorder, an autoimmune disorder, a metabolic disorder, a hormonal disorder, an inflammatory disorder, a developmental disorder, a reproductive disorder, an imprinting disorder, a genetic disorder, a neurological disorder, or a neurodegenerative disorder.
- the condition, disease, or disorder comprises a liver disorder, an eye disorder, a heart disorder, a kidney disorder, a skin disorder, a blood disorder, a fibrotic disorder, a skeletal disorder, or a muscle order.
- the condition, disease, or disorder is caused by a genetic mutation (e.g, an insertion, deletion, or point mutation).
- the condition, disease, or disorder is hereditary.
- the condition, disease, or disorder is caused by a virus or bacteria or fungus.
- the condition, disease, or disorder is caused by aberrant gene expression.
- the condition, disease, or disorder is a result of age. In some embodiments, the condition, disease, or disorder is chronic.
- the subject in the method of present disclosure can be an animal.
- the subject is an animal cell.
- the subject is a mammal.
- the subject is a human.
- the subject is an aquaculture animal (fish, crabs, shrimp, oysters etc.), a mammal.
- the animals cell is from, for example, a pet or zoo animal (cats, dogs, lizards, birds (e.g., parrots), lions, tigers and bears etc.), from a farm or working animal (horses, cows (e.g., dairy and beef cattle) pigs, chickens, turkeys, hens or roosters, goats, sheep, etc.), or a human.
- the target cell as disclosed herein is in a subject to whom the method of the present disclosure is applicable.
- the methods described herein can be therapeutic or veterinary methods for treating a subject.
- the methods described herein are used to treat a disease resulting from a nonfunctional, poorly functional, or poorly expressed protein or gene product, for instance, resulting from a genetic mutation in one or more cells of the subject.
- the methods described herein are used to treat a genetic disease (e.g.
- a mutation, a substitution, a deletion, an expansion, or a recombination a monogenic disease, an inherited metabolic disease, a cancer, a neurodegenerative disease, a cardiovascular disease, a pulmonary disease, a renal disease, a liver disease, a genetic disease, a vascular disease, ophthalmic disease, musculoskeletal disease, lymphatic disease, auditory and inner ear disease, a metabolic disease, an inflammatory disease, an autoimmune disease, or an infectious disease.
- a retinal disease e.g., Leber congenital amaurosis
- kits comprising the unit doses containing the lipid delivery particles, systems, compositions or pharmaceutical compositions of the present disclosure.
- the kit comprises the lipid delivery particles, compositions, or pharmaceutical formulations of the present disclosure; and an informational medium containing instructions for administering the lipid delivery particle, composition, or pharmaceutical formulation to a subject.
- the kit can include a label indicating the intended use of lipid delivery particle, composition, or pharmaceutical formulation in the kit. Label can include any writing, or recorded material supplied on or with the kit, or which otherwise accompanies the kit.
- kits of the present disclosure can include, alternatively or additionally, diagnostic agents and/or other therapeutic agents.
- the kit includes cells or pharmaceutical formulations of the present disclosure and a diagnostic agent that can be used in a diagnostic method for diagnosing a condition, disease, or disorder in a subject.
- the composition or pharmaceutical formulation described herein is prepared for administration to a subject.
- the pharmaceutical formulation is prepared to induce a therapeutic or prophylactic effect in a subject.
- Suitable routes of administrating the pharmaceutical formulation described herein include transdermal, intravesical, intravenous, intravascular, intraosseus, topical, subcutaneous, intradermal, intralesional, intraarticular, intraperitoneal, transmucosal, gingival, intradental, intracochlear, transtympanic, intraorgan, epidural, intrathecal, intramuscular, periocular, intratumoral, intracerebral, intravitreal, and intracerebroventricular administration.
- the pharmaceutical formulation described herein is administered locally to a diseased site (e.g., site of infection or tumor site).
- the pharmaceutical composition described herein is delivered in a controlled release system.
- a pump is used.
- polymeric materials is used for controlled release.
- the pharmaceutical composition described herein is administered to a subject by injection, by means of a catheter, by means of a suppository, or by means of an implant, the implant being of a porous, non-porous, or gelatinous material, including a membrane, such as a sialastic membrane, or a fiber.
- the pharmaceutical formulation is formulated in accordance with routine procedures as a formulation adapted for intravenous or subcutaneous administration to a subject.
- pharmaceutical formulations for administration by injection are solutions in sterile isotonic use as solubilizing agent and a local anesthetic such as lignocaine to ease pain at the site of the injection.
- the ingredients can be supplied either separately or mixed together in unit dosage form, for example, as a dry lyophilized powder or water free concentrate in a hermetically sealed container such as an ampoule or sachets indicating the quantity of active agent.
- the pharmaceutical is to be administered by infusion, it is dispensed with an infusion bottle containing sterile pharmaceutical grade water or saline.
- a pharmaceutical formulation as described herein can be administered or packaged as a unit dose, for example, in reference to a pharmaceutical formulation to physically discrete units suitable as unitary dosage for the subject, each unit containing a predetermined quantity of active material calculated to produce the desired therapeutic effect in association with the required diluent, carrier, or vehicle.
- Example 1 Screens to identify Pleckstin Homology domains with specific properties
- a screen of Pleckstin Homology domains will be performed to identify mutations that confer a PH domain with enhanced or altered properties.
- mutations will be introduced through mutagenesis and/or gene editing-based methods.
- Unedited PH domains from alternative sources will also be screened.
- Non-limiting examples of enhanced or altered properties that will be measured include recruitment of a payload into the lipid delivery particle, the amount of a payload that can be encapsulated in the lipid delivery particle, release of a payload from the lipid delivery particle into a target cell, trafficking of the lipid delivery particle or payload protein to a subcellular locale in a target cell, or suitability of the PH domain for combination with other proteins, protein domains, or protein components.
- a single PH domain will be fused to a reporter protein, such as GFP, or another enzyme with measurable functional activity such as Cas9 or ABE8e.
- the PH domain-GFP and/or PH domain-Cas9 chimeric proteins will be packaged into lipid delivery particles.
- a range of cell lines, including HEK293T, HeLa, HepG2, ARPE-19, BEAS-2B, or SH-SYSY cells, will be treated with the packaged lipid delivery particles that were previously purified.
- a single PH domain will be fused to a reporter protein, such as GFP, or another enzyme with measurable functional activity such as Cas9 or ABE8e.
- the PH domain-GFP and/or PH domain-Cas9 chimeric proteins will be packaged into lipid delivery particles.
- a range of cell lines including HEK293T, He La, HepG2, ARPE-19, BEAS-2B, or SH-SYSY cells, will be treated with the packaged lipid delivery particles that were previously purified.
- the purification and concentration of the lipid delivery particle will be performed by filtration methods, such as PVDF filtration, and ultracentrifiigation.
- the amount of a payload that can be released into the cell via the lipid delivery particle using different PH domains will be measured and/or quantified by flow cytometry, microscopy, or sequencing.
- a single PH domain will be fused to a reporter protein, such as GFP, or another enzyme with measurable functional activity such as Cas9 or ABE8e.
- the PH domain- GFP and/or PH domain-Cas9 chimeric proteins will be packaged into lipid delivery particles.
- a range of cell lines, including HEK293T, HeLa, HepG2, ARPE-19, BEAS-2B, or SH-SYSY cells, will be treated with the packaged lipid delivery particles that were previously purified.
- the purification and concentration of the lipid delivery particle will be performed by filtration methods, such as PVDF filtration, and ultracentrifugation.
- the amount of a payload protein that can be trafficked into a subcellular locale using different PH domains will be measured and/or quantified by flow cytometry, microscopy, or sequencing.
- a single PH domain will be fused to a reporter protein, such as GFP, another enzyme with measurable functional activity such as Cas9 or ABE8e, or a range of other protein domains.
- the PH domain-GFP and/or PH domain-Cas9 chimeric proteins will be packaged into lipid delivery particles.
- a range of cell lines, including HEK293T, HeLa, HepG2, ARPE-19, BEAS- 2B, or SH-SYSY cells, will be treated with the packaged lipid delivery particles that were previously purified.
- the purification and concentration of the lipid delivery particle will be performed by filtration methods, such as PVDF filtration, and ultracentrifugation.
- the suitability of an PH domain for combination with other protein domains using different PH domains will be measured and/or quantified by flow cytometry, microscopy, sequencing, a combination thereof, or using another method.
- PH domains were tested for gene editing capacity as determined by the percentage of sequencing reads expressing edits (any derivation from the wildtype sequence at a specific locus was counted as an editing event).
- Lipid delivery particles were generated and delivered to two cell lines, HEK293 (FIGs. 2A and 2B) and K562 (FIGs. 2C and 2D) at a high and low dose.
- Different PH domain- Cas9 chimeric proteins were generated and encapsulated as payloads in the lipid delivery particle for delivery into HEK293 and K562 cells.
- Particles were concentrated lOOx by PEG precipitation prior to transduction of recipient cells.
- the lipid delivery vehicles were introduced to the cells at a high dose and a low dose.
- a Cas9 chimeric protein lacking a PH domain (No PH) and an untreated sample (NT) was included as a negative control.
- the lipid delivery particles were transduced.
- the cells were lysed, genomic DNA was extracted, amplicons inducing the target site were generated, and the efficiency of the Cas9 fused to different PH domains was assessed through quantification of edits installed at the target editing site. The results are summarized in FIGs. 2A-2D.
- the treated and control samples were harvested for genomic DNA using Qiagen Blood & Cell Culture DNA kit.
- the DNA was used as a template for PCR, wherein HEXA target specific primers flank the insertion locus.
- PCR was performed using a high fidelity, proof-reading Taq polymerase (Prime Star, Takara) to isolate the target region of interest.
- the PCR inserts was purified and prepared for a next generation sequencing run using a library preparation kit (Nextera XT, Illumina). The library was run on the Illumina MiSeq and editing efficiency of each guide RNA and the PH domains was determined (FIGs. 2A-2D).
- Example 3 Generation of a lipid delivery particle comprising a Pleckstin Homology domain, an envelope, and gene editing machinery
- a nucleic acid molecule will be generated that encodes a chimeric protein that comprises one of the PH domains disclosed herein and a payload protein.
- the nucleic acid construct will encode the PH domain AKT1 E17K/R25C (SEQ ID NO. 575).
- Another nucleic acid molecule that will be generated encodes an envelope and target specific guide RNA.
- the payload protein comprises a Cas domain.
- the nucleic acid molecules encoding the chimeric protein, the envelope, and the target specific guide RNA are introduced into HEK293T producer cells to generate the lipid delivery particles comprising protein payload.
- Example 4 Delivery of a lipid delivery particle comprising a Pleckstin Homology domain, an envelope, and gene editing machinery to a target cell
- the lipid delivery particle comprising the protein payload generated in Example 2 will be introduced into a target cell.
- the target cell will be an induced pluripotent stem cell derived neurons model (iPSC-N).
- the protein payload will comprise a guide RNA that targets alpha subunit of the enzyme [3-hexosaminidase (HEXA) located on human chromosome 15. Multiple guide RNAs will be designed to target HEXA in exon 11, the locus that encodes a pathogenic 4 base pair insertion.
- the guide RNAs will be analyzed for specificity, efficiency, secondary structure, and potential to induce off-target mutations.
- the PH domains SEQ ID NO: 569-596 will be modified for delivery to neurons and efficiency of the gene editing protein payload.
- nucleic acid molecules encoding the chimeric protein, the envelope, and the target specific guide RNA will be introduced into HEK293T producer cells.
- the lipid delivery particles will be isolated and purified using filtration methods and centrifugation.
- the iPSC-Ns expressing the 4 base pair insertion mutation will be treated with the lipid delivery particle and cultured for 48-72 hours.
- a control iPSC-Ns expressing the 4 bae pair insertion will be treated with the lipid delivery particle comprising a scramble guide RNA particle and cultured for 48-72 hours.
- the treated and control iPSC-Ns are harvested for genomic DNA using Qiagen Blood & Cell Culture DNA kit.
- the DNA will be used as a template for PCR, wherein HEXA target specific primers flank the insertion site.
- PCR will be performed using a high fidelity, proof-reading Taq polymerase (Prime Star, Takara) to isolate the target region of interest.
- the PCR inserts will be purified and prepared for a next generation sequencing run using a library preparation kit (Nextera XT, Illumina). The library will then be run on the Illumina MiSeq and editing efficiency of each guide RNA and the PH domains will be determined.
- Example 5 Delivery of a lipid delivery particle to a subject with genetic disease
- the guide RNA and PH domain will then be provided to a subject with familial hypertrophic cardiomyopathy caused by a pathogenic insertion in HEXA.
- the lipid delivery particle will be provided to the subject intravenously or through cerebral spinal fluid.
- the gene editing payload protein will be delivered to the target neurons, where the guide RNA delivers the gene editing machinery to exon 11 of HEXA, and removes the pathogenic insertion.
- a sample from the subject will be taken after treatment to assess if the genome alteration is sufficient to restore normal function of the gene or to block pathogenic function of the pathogenic gene.
- a control sample from the subject will be taken from the subject prior to treatment.
- the treated and control samples will be harvested for genomic DNA using Qiagen Blood & Cell Culture DNA kit.
- the DNA will be used as a template for PCR, wherein HEXA target specific primers flank the insertion locus.
- PCR will be performed using a high fidelity, proof-reading Taq polymerase (Prime Star, Takara) to isolate the target region of interest.
- the PCR inserts will be purified and prepared for a next generation sequencing run using a library preparation kit (Nextera XT, Illumina). The library will then be run on the Illumina MiSeq and editing efficiency of each guide RNA and the PH domains will be determined.
Landscapes
- Health & Medical Sciences (AREA)
- Life Sciences & Earth Sciences (AREA)
- Chemical & Material Sciences (AREA)
- Genetics & Genomics (AREA)
- Organic Chemistry (AREA)
- Engineering & Computer Science (AREA)
- Molecular Biology (AREA)
- Zoology (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Wood Science & Technology (AREA)
- Biomedical Technology (AREA)
- General Health & Medical Sciences (AREA)
- Biochemistry (AREA)
- General Engineering & Computer Science (AREA)
- Biotechnology (AREA)
- Microbiology (AREA)
- Medicinal Chemistry (AREA)
- Biophysics (AREA)
- Toxicology (AREA)
- Gastroenterology & Hepatology (AREA)
- Proteomics, Peptides & Aminoacids (AREA)
- Physics & Mathematics (AREA)
- Plant Pathology (AREA)
- Peptides Or Proteins (AREA)
Abstract
The present disclosure relates to the selection, engineering, development, and use of Plextrin Homology domains in lipid delivery particles. The Plextrin Homology can be coupled to protein payloads for the delivery of the protein payload to a target cell.
Description
PLECKSTRIN HOLOMOGY DOMAINS FOR DELIVERY OF PROTEIN PAYLOADS TO A
TARGET CELL
CROSS-REFERENCE
[0001] This application claims the benefit of U.S. Provisional Application No. 63/506,811, filed June 7, 2023, which application is incorporated herein by reference.
BACKGROUND
[0002] Delivery of therapeutic payloads into the cells has been a significant challenge in drug development because payloads, such as proteins, cannot freely diffuse across the cell membrane. Although viral based constructs have been developed to deliver therapeutic payloads, these constructs often have low efficacy and/or create considerable side effects. There is a need for improved delivery of payloads into cells.
SUMMARY
[0003] Disclosed herein, in some aspects, is a chimeric protein comprising a Pleckstrin Homology domain comprising: (i) at least 60% at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99.1%, at least 99.2%, at least 99.3%, at least 99.4%, at least 99.5%, at least 99.6%, at least 99.7%, at least 99.8%, or at least 99.9% sequence identity to the sequence of SEQ ID NO: 569; and (ii) at least two selected from the group of amino acid substitutions consisting of E17K, R25C, T81Y, T101C, and a combination of K142A, Hl 43 A, R144A (142-144A) relative to the sequence of SEQ ID NO: 569; and a protein payload.
[0004] In some cases, the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K and R25C; E17K and T81Y; E17K and T101C; E17K and 142-144A; R25C and T81Y; R25C and T101C; R25C and 142-144A; T81Y and T101C; T81Y and 142-144A; and T101C and 142-144A. In some cases, the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 572, 573, 574, 575, 576, 577, 578, 579, or 580. In some cases, the chimeric protein comprises at least three mutations selected from the group consisting of E17K, R25C, T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569. In some cases, the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K, R25C, and T81Y; E17K, R25C, and T101C; E17K, R25C, and 142-144A; E17K, T81Y, and T101C; E17K, T81Y, and 142-144A; E17K, T101C, and 142-144A; R25C, T81Y, and T101C; R25C, T81Y, and 142-144A; R25C, T101C, and 142-144A; and T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569.
[0005] In some cases, the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 581, 582, 583, 584, 585,586, 587, 588, 589, or 590. In some cases, the chimeric protein comprises at least four selected from the group consisting of E17K, R25C, T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569. In some cases, the chimeric protein comprises at least one mutation group
selected from the group consisting of: E17K, R25C, T81Y, and T101C; E17K, R25C, T81Y, and 142- 144A; R25C, T81Y, T101C, and 142-144A; R25C, T81Y, T101C, and 142-144A; and E17K, T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569. In some cases, the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 591, 592, 593, 594, or 595. In some cases, the chimeric protein comprises at least the mutations E17K, R25C, T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569. In some cases, the chimeric protein comprises the sequence of SEQ ID NO: 596.
[0006] In some cases, the chimeric protein further comprises one or more nuclear export sequences. In some cases, the one or more nuclear export sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 5. In some cases, the chimeric protein further comprises one or more nuclear localization sequences. In some cases, the one or more nuclear localization sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 6.
[0007] In some cases, the Pleckstrin Homology domain is coupled to the protein payload. In some cases, the Pleckstrin Homology domain is reversibly coupled to the protein payload. In some cases, the protein payload is reversibly coupled to the Pleckstrin Homology domain by a cleavable linker. In some cases, the cleavable linker is cleavable by a protease. In some cases, the protein payload is coupled to a C- terminal of the Pleckstrin Homology domain.
[0008] In some cases, the protein payload is coupled to an N-terminal of the Pleckstrin Homology domain.
[0009] In some cases, the protein payload comprises one or more viral proteins. In some cases, the one or more viral proteins comprise one or more retroviral proteins. In some cases, the one or more retroviral proteins comprises a retroviral Gag protein. In some cases, the protein payload comprises one or more non-viral proteins. In some cases, the one or more non-viral proteins comprises one or more mammalian proteins. In some cases, the one or more mammalian proteins comprises an Arc protein.
[0010] In some cases, the protein payload comprises a gene editing protein. In some cases, the gene editing protein comprises a prime editing protein. In some cases, the prime editing protein is coupled to a target-specific prime editing guide RNA. In some cases, the gene editing protein comprises a CRISPR system. In some cases, the CRISPR system comprises a Cas domain. In some cases, the gene editing protein is coupled to a target-specific guide RNA. In some cases, the protein payload comprises an epigenetic editing protein. In some cases, the epigenetic editing protein is coupled to a target-specific guide RNA. In some cases, the protein payload comprises a recombinase protein. In some cases, the protein payload comprises an integrase protein.
[0011] Disclosed herein, in some aspects, is a nucleic acid molecule encoding a chimeric protein.
[0012] Disclosed herein, in some aspects, is a lipid delivery particle comprising a chimeric protein. In some cases, the lipid delivery particle further comprises an envelope, wherein the envelope comprises a lipid bilayer encasing a lumen, wherein the chimeric protein is located in the lumen. In some cases, the lipid delivery particle further comprises a target ligand.
[0013] In some cases, the target ligand reversibly couples to a target receptor. In some cases, the target receptor comprises a cell surface protein on a target cell. In some cases, the envelope couples to the target cell at least partially through coupling of the target ligand and the target receptor on the target cell. In some cases, coupling of the target ligand and the target receptor on the target cell induces release of the protein payload in the target cell. In some cases, release of the protein payload in the target cell follows fusion of the envelope with cell membrane of the target cell.
[0014] Disclosed herein, in some aspects, is a method to deliver a payload to a target cell, the method comprising delivering the lipid delivery particle to a target cell.
[0015] Disclosed herein, in some aspects, is a chimeric protein comprising: a Pleckstrin Homology domain comprising at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99.1%, at least 99.2%, at least 99.3%, at least 99.4%, at least 99.5%, at least 99.6%, at least 99.7%, at least 99.8%, or at least 99.9% sequence identity to any one of the sequences set forth in Table 4B; and a protein payload. In some cases, the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 597, 598, 599, 600, 601, 602, 603, 604, 605, 606, 607, 608, 609, 610, 611, 612, 613, 614, 615, 616, 617, 618, 619, 620, 621, 622, 623, 624, 625, 626, 627, 628, 629,
630, 631, 632, 633, 634, 635, 636, 637, 638, 639, 640, 641, 642, 643, 644, 645, 646, 647, 648, 649, 650,
651, 652, 653, 654, 655, 656, 657, 658, 659, 660, 661, 662, 663, 664, 665, 666, 667, 668, 669, 670, 671,
672, 673, 674, 675, 676, 677, 678, 679, 680, 681, 682, 683, 684, 685, 686, 687, 688, 689, 690, 691, 692,
693, 694, 695, 696, 697, 698, 699, 700, 701, 702, 703, 704, 705, 706, 707,708, 709, 710, 711, 712, 713, 714, 715, 716, 717, 718, 719, 720, 721, or 722.
[0016] In some cases, the chimeric protein further comprises one or more nuclear export sequences. In some cases, the one or more nuclear export sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 5. In some cases, the chimeric protein further comprises one or more nuclear localization sequences. In some cases, the one or more nuclear localization sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 6.
[0017] In some cases, the Pleckstrin Homology domain is coupled to the protein payload. In some cases, the Pleckstrin Homology domain is reversibly coupled to the protein payload. In some cases, the protein payload is reversibly coupled to the Pleckstrin Homology domain by a cleavable linker. In some cases, the cleavable linker is cleavable by a protease. In some cases, the protein payload is coupled to C-terminal of the Pleckstrin Homology domain. In some cases, the protein payload is coupled to N-terminal of the Pleckstrin Homology domain.
[0018] In some cases, the protein payload comprises one or more viral proteins. In some cases, the one or more viral proteins comprise one or more retroviral proteins. In some cases, the one or more retroviral proteins comprises a retroviral Gag protein. In some cases, the protein payload comprises one or more non-viral proteins. In some cases, the one or more non-viral proteins comprises one or more mammalian proteins. In some cases, the one or more mammalian proteins comprises an Arc protein.
[0019] In some cases, the protein payload comprises a gene editing protein. In some cases, the gene editing protein comprises a prime editing protein. In some cases, the prime editing protein is coupled to a target-specific prime editing guide RNA. In some cases, the gene editing protein comprises a CRISPR system. In some cases, the CRISPR system comprises a Cas domain. In some cases, the gene editing protein is coupled to a target-specific guide RNA. In some cases, the protein payload comprises an epigenetic editing protein. In some cases, the epigenetic editing protein is coupled to a target-specific guide RNA. In some cases, the protein payload comprises a recombinase protein. In some cases, the protein payload comprises an integrase protein.
[0020] Disclosed herein, in some aspects, is a nucleic acid molecule encoding a chimeric protein.
[0021] Disclosed herein, in some aspects, is a lipid delivery particle comprising a chimeric protein. In some cases, the lipid delivery particle further comprises an envelope, wherein the envelope comprises a lipid bilayer encasing a lumen, wherein the chimeric protein is located in the lumen. In some cases, the lipid delivery particle further comprises a target ligand.
[0022] In some cases, the target ligand reversibly couples to a target receptor. In some cases, the target receptor comprises a cell surface protein on a target cell. In some cases, the envelope couples to the target cell through coupling of the target ligand and the target receptor on the target cell. In some cases, coupling of the target ligand and the target receptor on the target cell induces release of the protein payload in the target cell. In some cases, release of the protein payload in the target cell follows endocytosis of the lipid delivery particle.
[0023] Disclosed herein, in some aspects, is a method to deliver a payload to a target cell, the method comprising: delivering the lipid delivery particle to a target cell.
[0024] Disclosed herein, in some aspects, is a chimeric protein comprising: a Pleckstrin Homology domain comprising: (i) at least 60% at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99.1%, at least 99.2%, at least 99.3%, at least 99.4%, at least 99.5%, at least 99.6%, at least 99.7%, at least 99.8%, or at least 99.9% sequence identity to the sequence of SEQ ID NO: 613; and (ii) at least one amino acid substitution selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K relative to the sequence of SEQ ID NO: 613; and a protein payload.
[0025] In some cases, the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 603, 604, 605, 606, or 607. In some cases, the chimeric protein comprises at least two mutations selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613. In some cases, the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K and E58K; E17K and E52K; E17K and E53K; E17K and E55K; E17K and E65K; E58K and E52K; E58K and E53K; E58K and E55K; E58K and E65K; E52K and E53K; E52K and E55K; E52K and E65K; E53K and E55K; and E53K and E65K, relative to the sequence of SEQ ID NO: 613. In some cases, the chimeric protein comprises the sequence of SEQ ID NO: 608. In some cases, the chimeric protein comprises at least three mutations selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613.
[0026] In some cases, the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K, E58K, and E52K; E17K, E58K, and E53K; E17K, E58K, and E55K; E17K, E58K, and E65K; E17K, E52K, and E53K; E17K, E52K, and E55K; E17K, E52K, and E65K; E17K,
E53K, and E55K; E17K, E53K, and E65K; E58K, E52K, and E53K; E58K, E52K, and E55K; E58K,
E52K, and E65K; E58K, E53K, and E55K; E58K, E53K, and E65K; E52K, E53K, and E65K; and E52K,
E53K, and E55K, relative to the sequence of SEQ ID NO: 613. In some cases, the chimeric protein comprises the sequence of SEQ ID NO: 609. In some cases, the chimeric protein comprises at least four mutations selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613. In some cases, the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K, E58K, E52K, and E53K; E17K, E58K, E52K, and E55K; E17K, E58K, E52K, and E65K; E58K, E52K, E53K, and E55K; E58K, E52K, E53K, and E65K; E58K, E52K, E53K, and E65K, relative to the sequence of SEQ ID NO: 613. In some cases, the chimeric protein comprises the sequence of SEQ ID NO: 610.
[0027] In some cases, the chimeric protein comprises at least five selected from the group consisting of: E17K, E58K, E52K, E53K, and E55K; E17K, E58K, E52K, E53K, and E65K; E17K, E58K, E52K, E55K, and E65K; E17K, E58K, E53K, E55K, and E65K; E17K, E52K, E53K, E55K, and E65K; and E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613. In some cases, the chimeric protein comprises the sequence of SEQ ID NO: 611. In some cases, the chimeric protein comprises the mutations E17K, E58K, E52K, E53K, E55K, and E65K , relative to the sequence of SEQ ID NO: 569. In some cases, the chimeric protein comprises the sequence of SEQ ID NO: 612.
[0028] In some cases, the chimeric protein further comprises one or more nuclear export sequences. In some cases, the one or more nuclear export sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 5. In some cases, the chimeric protein further comprises one or more nuclear localization sequences. In some cases, the one or more nuclear localization sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 6. [0029] In some cases, the Pleckstrin Homology domain is coupled to the protein payload. In some cases, the Pleckstrin Homology domain is reversibly coupled to the protein payload. In some cases, the protein payload is reversibly coupled to the Pleckstrin Homology domain by a cleavable linker. In some cases, the cleavable linker is cleavable by a protease. In some cases, the protein payload is coupled to C-terminal of the Pleckstrin Homology domain. In some cases, the protein payload is coupled to N-terminal of the Pleckstrin Homology domain.
[0030] In some cases, the protein payload comprises one or more viral proteins. In some cases, the one or more viral proteins comprise one or more retroviral proteins. In some cases, the one or more retroviral proteins comprises a retroviral Gag protein. In some cases, the protein payload comprises one or more non-viral proteins. In some cases, the one or more non-viral proteins comprises one or more mammalian proteins. In some cases, the one or more mammalian proteins comprises an Arc protein.
[0031] In some cases, the protein payload comprises a gene editing protein. In some cases, the gene editing protein comprises a prime editing protein. In some cases, the prime editing protein is coupled to a target-specific prime editing guide RNA. In some cases, the gene editing protein comprises a CRISPR system. In some cases, the CRISPR system comprises a Cas domain. In some cases, the gene editing protein is coupled to a target-specific guide RNA. In some cases, the protein payload comprises an epigenetic editing protein. In some cases, the epigenetic editing protein is coupled to a target-specific guide RNA. In some cases, the protein payload comprises a recombinase protein. In some cases, the protein payload comprises an integrase protein.
[0032] Disclosed herein, in some aspects, is a nucleic acid molecule encoding a chimeric protein.
[0033] Disclosed herein, in some aspects, is a lipid delivery particle comprising a chimeric protein. In some cases, the lipid delivery particle further comprises an envelope, wherein the envelope comprises a lipid bilayer encasing a lumen, wherein the chimeric protein is located in the lumen. In some cases, the lipid delivery particle further comprises a target ligand. In some cases, the target ligand reversibly couples to a target receptor. In some cases, the target receptor comprises a cell surface protein on a target cell. In some cases, the envelope couples to the target cell through coupling of the target ligand and the target receptor on the target cell. In some cases, coupling of the target ligand and the target receptor on the target cell induces release of the protein payload in the target cell. In some cases, release of the protein payload in the target cell follows endocytosis of the lipid delivery particle.
[0034] Disclosed herein, in some aspects, is a method to deliver a payload to a target cell, the method comprising delivering the lipid delivery particle to a target cell.
INCORPORATION BY REFERENCE
[0035] All publications, patents, and patent applications mentioned in this specification are herein incorporated by reference to the same extent as if each individual publication, patent, or patent application was specifically and individually indicated to be incorporated by reference.
BRIEF DESCRIPTION OF THE DRAWINGS
[0036] The novel features of the disclosure are set forth with particularity in the appended claims. A better understanding of the features and advantages of the present disclosure will be obtained by reference to the following detailed description that sets forth illustrative embodiments, in which the principles of the disclosure are utilized, and the accompanying drawings of which:
[0037] FIG. 1 is a schematic showing nonlimiting examples of potential architecture for the chimeric protein disclosed herein. In the schematic, the black box designates the plasma membrane recruitment element, the checkered box designates the [n * nuclear export signal], the black line designates the cleavable linker, and the gray box designates a payload, which can further comprise a nuclear localization sequence.
[0038] FIGs. 2A-2D are graphs depicting the editing percentage in HEK293T and K5662 cells using Cas9 genetic editors fused to various PH domains and encapsulated in lipid delivery particles at high and low doses. FIG. 2A is a graph depicting editing percentages in HEK293T cells treated with a high dose of lipid delivery particle encapsulated with Cas9 genetic editors fused to various PH domains. FIG. 2B is a graph depicting editing percentages in HEK293T cells treated with a low dose of lipid delivery particle encapsulated with Cas9 genetic editors fused to various PH domains. FIG. 2C is a graph depicting editing percentages in K562 cells treated with a high dose of lipid delivery particle encapsulated with Cas9 genetic editors fused to various PH domains. FIG. 2D is a graph depicting editing percentages in K562 cells treated with a low dose of lipid delivery particle encapsulated with Cas9 genetic editors fused to various PH domains.
DETAILED DESCRIPTION
[0039] In some aspects, the present disclosure relates to the use of Pleckstrin Homology (PH) domains for delivery of protein payloads to target cells. In some cases, PH domains are used to facilitate recruitment and packaging of a payload in a lipid delivery particle. In some cases, PH domains have a range of attributes and functions that can be modified for particular applications. The PH domain can be selected or engineered to for one or more improved attributes and functions for a particular application. In some cases, the plasma membrane recruitment element is modified for more efficient recruitment of protein payloads into a lipid delivery particle. In some cases, the plasma membrane recruitment element is modified so that the total number of active protein payload that can be encapsulated in a lipid delivery particle is increased. The PH domain can be modified for more efficient release of the protein payload from a lipid delivery particle into a target cell. In some cases, the plasma membrane recruitment element is modified for more efficient trafficking of the protein payload to a particular target cell. In some cases, the plasma membrane recruitment element can be modified for more efficient trafficking of the protein payload to a particular subcellular locale in the target cell. In some cases, the plasma membrane recruitment element is modified for suitability for combination with other proteins, protein domains, protein components, or a combination thereof.
[0040] In some aspects, provided herein are chimeric proteins comprising engineered PH domains. In some aspects, provided herein are chimeric proteins comprising PH domains coupled to payloads. Also provided herein, in some aspects, are nucleic acid molecules encoding the chimeric proteins and protein payloads. Also provided herein, in some aspects, are lipid delivery particles comprising the chimeric proteins with PH domains. In some aspects, provided herein also includes cells comprising nucleic acid molecules encoding the chimeric proteins coupled to protein payloads. In some aspects, provided herein are cells comprising the chimeric proteins. Also provided herein, in some aspects, are methods of making lipid delivery vehicles comprising chimeric proteins comprising PH domains coupled to protein payloads. In some aspects, provided herein are methods of contacting target cells with lipid delivery particles comprising chimeric proteins comprising PH domains coupled to protein payloads. Also provided herein, in some aspects, are kits comprising nucleic acid molecules encoding chimeric proteins and protein payloads.
ENVELOPE PROTEIN
[0041] In some aspects, the lipid delivery particle provided herein comprises an envelope protein. The envelope protein can be associated with the outside boundary or the surface of the lipid delivery particle, for example, the membrane or envelope of the lipid delivery particle.
[0042] The membrane of the lipid delivery particle can comprise a lipid layer, such as a single layer or a lipid bilayer. In some cases, the membrane of the lipid delivery particle is from plasma membrane, endoplasmic reticulum, or a combination thereof. In some cases, the membrane of the lipid delivery particle is from Golgi complex, ER Golgi intermediate compartment, or nuclear envelope. In some cases, the membrane of the lipid delivery particle is from plasma membrane. In some cases, the membrane of the lipid delivery particle is a phospholipid bilayer.
[0043] The envelope protein can be associated with the membrane of the lipid delivery particle in various manners. For example, the envelope protein can be anchored or attached to the external membrane of the particle or anchored or attached to the internal membrane of the particle. The envelope protein can be embedded or inserted in the membrane, spanning through the membrane, with certain portions located at the outside of the membrane, or certain portions extending to the inside of the particle, or both. The envelope protein within the lipid delivery particle described herein can be overexpressed from an exogenous source, such as plasmids or stably integrated transgenes, in the production cells. [0044] The envelope protein can play a role in the delivery of the lipid delivery particle to a target cell and release of the components of the lipid delivery particle within the target cell. The envelope protein can contact with the surface of a target cell and participate in the fusion of the lipid delivery particle and the membrane of the target cell. The envelope protein can participate in the fusion of the lipid delivery particle with the membrane of the target cell via any appropriate mechanism, such as those described in White et al. Crit Rev Biochem Mol Biol. 2008; 43(3): 189-219. One example of the fusion mechanisms is unifying Trimer-of-Hairpins Fusion Mechanism. Membrane fusion can occur after allosteric priming by binding to a target receptor. In some cases, membrane fusion occurs after proteolysis. In some cases, membrane fusion occurs after isomerization of disulfide bridges. In some cases, membrane fusion occurs by internalization and then priming of fusion via (i) cathepsin-mediated proteolysis, or (ii) low pH/acidification. The cathepsin-mediated proteolysis can be pH dependent or pH independent. Other fusion triggering mechanisms can include low PH, binding to target cell receptors, and a receptor followed by low pH. The envelope protein can also play a role in the formation of the lipid delivery particle. The envelope protein can interact with another component within the lipid delivery particle and participate in the assembly of the lipid delivery particle, for example, in a producer cell. The envelope protein can make contact with another envelope protein and form an oligomer embedded within the membrane. The envelope protein can be a glycoprotein, for example, a transmembrane glycoprotein. In some cases, envelope protein comprises multiple membrane -spanning regions. These multiple membranespanning regions can oligomerize and form channels in the membrane.
[0045] In some cases, the envelope protein is fused with a targeting moiety. In some cases, the targeting moiety recognizes a specific molecule (e.g, antigen, receptor, or other membrane protein) on the surface of a target cell to allow targeted cell entry with more specificity. In some cases, the targeting moiety is specific for a certain cell type or is specific for a certain target cell. The targeting moiety can be fused to the envelope protein at a position that is located at an outside of the lipid delivery particle. For example, the targeting moiety includes scFvs, antibody variable regions, nanobodies, T-cell receptor variable regions, other antigen-binding fragments or their mimetics, such as DARPins. In some cases, the targeting moiety is a protein ligand from the human ligandome. The targeting moiety can be a natural peptide or a synthetic peptide. In some cases, the targeting moiety is not fused with the envelope protein and is attached to the membrane of the lipid delivery particle from the outside, for example, via a transmembrane domain.
[0046] A targeting moiety can include, e.g., an antibody or an antigen-binding fragment thereof (e.g., Fab, Fab', F(ab')2, Fv fragments, scFv antibody fragments, disulfide-linked Fvs (sdFv), a Fd fragment consisting of the VH and CHI domains, linear antibodies, single domain antibodies such as sdAb (either VL or VH), nanobodies, or camelid VHH domains), an antigen-binding fibronectin type III (Fn3) scaffold such as a fibronectin polypeptide minibody, a ligand, a cytokine, a chemokine, or a T cell receptor (TCRs). Membrane-fusion proteins can be re-targeted by non-covalently conjugating a targeting moiety to the membrane-fusion protein or targeting protein (e.g. the hemagglutinin protein). For example, the membrane -fusion protein can be engineered to bind the Fc region of an antibody that targets an antigen on a target cell, redirecting the membrane fusion activity towards cells that display the antibody’s target. [0047] In some cases, the targeting moiety linked to the membrane -fusion protein binds a cell surface marker on the target cell, e.g, a protein, glycoprotein, receptor, cell surface ligand, agonist, lipid, sugar, class I transmembrane protein, class II transmembrane protein, or class III transmembrane protein.
[0048] In some cases, the lipid delivery particles disclosed herein display targeting moieties that are not conjugated to the membrane -fusion protein or other proteins in order to redirect the fusion activity of the lipid delivery particle towards a cell that is bound by the targeting moiety, or to affect tropism of the lipid delivery particle toward the target cell.
Envelope protein of viral origin
[0049] In some cases, an envelope protein has a viral origin. For example, a suitable envelope protein is from a DNA virus, an RNA virus, or a retrovirus. The envelope protein can be envelope protein from Herpesviruses, Avian sarcoma leukosis virus, Poxviruses, Hepadnaviruses, Asfarviridae, Flaviviruses, Alphaviruses, Togaviruses, Coronaviruses, Hepatitis D, Orthomyxoviruses, Rhabdovirus, Bunyaviruses, Filoviruses, Oncoretroviruses, lentiviruses, Spumaviruses. In some cases, envelope protein can be envelope protein from lentiviruses, for example, human immunodeficiency virus (HIV), simian immunodeficiency virus (SIV), feline immunodeficiency virus (FIV) and equine infectious anemia virus (EIAV). In some cases, an envelope protein is a fusion of two different envelope proteins, wherein each comes from a different virus. Additional suitable envelope proteins that are from viral origins and their functions are described in White JM et al., Crit Rev Biochem Mol Biol. 2008 May-Jun;43(3): 189-219.
[0050] In some cases, the envelope protein is a vesicular stomatitis virus glycoprotein (VSVG) or a biologically active mutant thereof. A “biologically active mutant” disclosed herein in connection with a reference protein can refer to a mutant of the reference protein that remains displaying one or more biological activities that are of same nature as the reference protein, which are relevant to the context in which the reference protein is used in the lipid delivery particle disclosed herein, while the level of the one or more biological activities of the biologically active mutant can be either similar as or different than the reference protein. For instance, the biologically active mutant of a VSVG in the context of an envelope protein remains displaying the biological activities of an envelope protein, e.g., mediating membrane fusion, tropism of the lipid delivery particle toward a target cell, or both. Unless otherwise noted, a mutant as described in the present disclosure is equivalent to a biologically active mutant. In some cases, the envelope protein is a Human immunodeficiency virus GP160 or a biologically active mutant thereof. In some cases, the envelope protein is a Baboon Endogenous Retrovirus (BaEVTR) glycoprotein or a biologically active mutant thereof. In some cases, the envelope protein is a modified Baboon Endogenous Retrovirus (BaEVTRless) glycoprotein or a biologically active mutant thereof. In some cases, the envelope protein is the fusion protein of Vesicular stomatitis Indiana virus and Rabies virus Glycoproteins (FuG-E) or a biologically active mutant thereof. In some cases, the envelope protein pantropic murine leukemia virus envelope protein (MLV) or a biologically active mutant thereof. In some cases, the envelope protein is a modified Fusion protein of Vesicular stomatitis Indiana virus and Rabies virus Glycoproteins (FuG-E P440E) or a biologically active mutant thereof. In some cases, the envelope protein is an ecotropic Murine Leukemia Virus envelope protein (MLV ENV ecotropic) or a biologically active mutant thereof. In some cases, the envelope protein is an amphotrophic Murine Leukemia Virus envelope protein (MLV ENV amphotropic) or a biologically active mutant thereof. In some cases, the envelope protein is a Moloney murine leukemia virus envelope protein (MMLV) or a biologically active mutant thereof. In some cases, the envelope protein is a Moloney murine sarcoma virus envelope protein (MoMSVg) or a biologically active mutant thereof. In some cases, the envelope protein is a moloney murine leukemia virus 10A1 strain Glycoprotein (MLV 10A1) or a biologically active mutant thereof. In some cases, the envelope protein is a xenotropic murine leukemia virus envelope protein (MLV ENV xenotropic) or a biologically active mutant thereof. In some cases, the envelope protein is a xenotropic murine leukemia virus-related envelope protein (XMRV) or a biologically active mutant thereof. In some cases, the envelope protein is a Baculovirus envelope glycoprotein (GP64) or a biologically active mutant thereof. In some cases, the envelope protein is an endogenous feline virus envelope protein (RD114 ENV) or a biologically active mutant thereof. In some cases, the envelope protein is a mammalian endogenous retrovirus protein, or a biologically active mutant thereof. The mammalian endogenous retrovirus protein can be a koala retrovirus protein (KoRV) or a Jaagsiekte sheep retrovirus protein (enJSRV), or a biologically active mutant thereof.
[0051] In some cases, the envelope protein is a simian endogenous type D retrovirus protein (RD-114) or a biologically active mutant thereof. In some cases, the envelope protein is a gibbon ape leukemia virus envelope protein (GALV) or a biologically active mutant thereof. In some cases, the envelope protein is a
feline leukemia virus envelope protein (FLV) or a biologically active mutant thereof. In some cases, the envelope protein is a mouse mammary tumor virus envelope protein (MMTV) or a biologically active mutant thereof. In some cases, the envelope protein is an avian leukosis virus envelope protein or a biologically active mutant thereof. In some cases, the envelope protein is a rous sarcoma virus envelope protein or a biologically active mutant thereof.
[0052] In some cases, the envelope protein can direct the lipid delivery particles to fuse with a certain type of target cells rather than other cells. For example, based on the specific type of envelope protein associated with the membrane of the lipid delivery particle, the lipid delivery particle can preferentially target different cell types (z.e., tropisms of the lipid delivery particles), such as liver cells, ocular cells, nerve cells, lung cells, immune cells, muscle cells, and any other cell types of interest. For example, to fuse with a target liver cells, the envelope protein can be a glycoprotein from human hepatitis viruses or a biologically active mutant thereof, e.g., Hepatitis B virus (HBV) or hepatitis C virus (HCV), VSV-G glycoprotein or a biologically active mutant thereof, a Marburg virus glycoprotein or a biologically active mutant thereof, an Ebola virus glycoprotein or a biologically active mutant thereof. To fuse with a target muscle cell, for example, a skeletal muscle cell, the envelope protein can be a Ross River virus glycoprotein or a biologically active mutant thereof, or a VSV-G or a biologically active mutant thereof. To fuse with a target ocular cell, for example, a photoreceptor cell or a retinal cell, the envelope protein can be an Ebola virus glycoprotein or a biologically active mutant thereof, a Marburg virus glycoprotein or a biologically active mutant thereof, or a VSV-G or a biologically active mutant thereof. To fuse with a target immune cell, for example, CD8+ T cell, an HTLV-1 glycoprotein or a biologically active mutant thereof, or a VSV- G glycoprotein or a biologically active mutant thereof. To fuse with a target immune cell, for example, CD4+ T cell, the envelope protein can be a HIV-1 envelope or a biologically active mutant thereof, a HTLV-1 glycoprotein or a biologically active mutant thereof, or a VSV-G glycoprotein or a biologically active mutant thereof. To fuse with a target lung cells, the envelope protein can be a respiratory syncytial virus glycoprotein or a biologically active mutant thereof, or a SARS-CoV glycoprotein or a biologically active mutant thereof. To fuse with a target nerve cell, such as a cell from the central nervous system cell (e.g., neurons, glial cells including oligodendrocytes, astrocytes and microglia), the envelope protein can be a rabies glycoprotein or a biologically active mutant thereof, a Mokola virus glycoprotein or a biologically active mutant thereof, a Semliki Forest virus glycoprotein or a biologically active mutant thereof, a Venezuelan equine encephalitis virus glycoprotein or a biologically active mutant thereof, or a VSV-G or a biologically active mutant thereof. To fuse with a target sensory cell, such as an auditory cell, including hair cells, cochlear cells, etc., the envelope protein can be an Ebola virus glycoprotein or a biologically active mutant thereof, a Marburg virus glycoprotein or a biologically active mutant thereof, or a VSV-G or a biologically active mutant thereof.
[0053] In some cases, the envelope protein comprises the sequences set forth in Table 1. In some cases, the envelope protein comprises the sequences set forth in Table 1 with at least one amino acid substitution, deletion, or insertion. For instance, N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein. In
some cases, the envelope protein comprises the sequences set forth in Table 1 and a heterologous peptide sequence fused to the N-terminal or C-terminal.
[0054] In some cases, the envelope protein comprises one or more of the sequences set forth in Table 1 with at least one amino acid substitution, deletion, or insertion. For instance, N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein. In some cases, the envelope protein comprises one or more of the sequences set forth in Table 1 and a heterologous peptide sequence fused to the N-terminal or C-terminal.
[0055] In some cases, the envelope protein comprises any one of the sequences set forth in Table 1 with at least one amino acid substitution, deletion, or insertion. For instance, N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein. In some cases, the envelope protein comprises any one of the sequences set forth in Table 1 and a heterologous peptide sequence fused to the N-terminal or C-terminal.
[0056] In some cases, the envelope protein comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in Table 1. In some cases, the envelope protein comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 50% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 60% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104 In some cases, the envelope protein comprises an amino acid sequence that has at least about 70% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 75% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 80% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104 In some cases, the envelope protein comprises an amino acid sequence that has at least about 85% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 90% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 95% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104 In some cases, the envelope protein comprises an amino acid sequence that has at least about 96% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 97% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 98% sequence identity to a sequence set
forth in any one of SEQ ID NOs: 83-104. In some cases, the envelope protein comprises an amino acid sequence that has at least about 99% sequence identity to a sequence set forth in any one of SEQ ID NOs: 83-104.
Envelope protein of human origin
[0057] In some aspects, the envelope protein in the lipid delivery particle described herein has a human origin, e.g., has significant sequence similarity to a human wild-type protein, such as at least 90%, at least 95%, at least 98%, or at least 99%. Using an envelope protein of a human origin can have benefits such as providing a minimized immunogenicity and better tolerance in a human subject receiving the lipid delivery particles. The lipid delivery particle comprising an envelope protein of a human origin can comprise another component that is from human origin or from non-human origin (e.g. , a payload or a plasma membrane recruitment element). An envelope protein that is from human origin can include, example, envelope proteins or glycoproteins of human endogenous retroviruses (HERVs), other human endogenous envelope proteins, or other human endogenous proteins that serve a similar function of recognizing and/or fusing with membrane of a target cell (e.g., clathrin adaptor protein complex- 1, CHMP4C, Proteolipid protein 1, TSAP6, immunoglobulin variable domains, or a biologically active mutant thereof).
[0058] In some cases, the envelope protein is a HERV envelope protein such as any one of those listed in Table 2. In some cases, the envelope protein is a hENVHl or a biologically active mutant thereof. In some cases, the envelope protein is a hENVH2 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVH3 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVKl or a biologically active mutant thereof. In some cases, the envelope protein is a hENVK2 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVK3 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVK4 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVK5 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVK6 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVT or a biologically active mutant thereof. In some cases, the envelope protein is a hENVW or a biologically active mutant thereof. In some cases, the envelope protein is a hENVFRD or a biologically active mutant thereof. In some cases, the envelope protein is a hENVR or a biologically active mutant thereof. In some cases, the envelope protein is a hENVR(b) or a biologically active mutant thereof. In some cases, the envelope protein is a hENVR(c)2 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVR(c) 1 or a biologically active mutant thereof. In some cases, the envelope protein is a hENVKcon or a biologically active mutant thereof. In some cases, the envelope protein is a truncated HERV protein.
[0059] In some cases, the envelope protein comprises the sequences set forth in Table 3. In some cases, the envelope protein comprises the sequences set forth in Table 3 with at least one amino acid substitution, deletion, or insertion. For example, for those amino acid sequences start with aN-terminal
methionine, the N-terminal methionine can be absent. In some cases, the envelope protein comprises the sequences set forth in Table 3 and a heterologous peptide sequence fused to the N-terminal or C-terminal. [0060] In some cases, the envelope protein comprises one or more of the sequences set forth in Table 3 with at least one amino acid substitution, deletion, or insertion. For instance, N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein. In some cases, the envelope protein comprises one or more of the sequences set forth in Table 3 and a heterologous peptide sequence fused to the N-terminal or C-terminal.
[0061] In some cases, the envelope protein comprises any one of the sequences set forth in Table 3 with at least one amino acid substitution, deletion, or insertion. For instance, N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein. In some cases, the envelope protein comprises any one of the sequences set forth in Table 3 and a heterologous peptide sequence fused to the N-terminal or C-terminal.
[0062] In some cases, the envelope protein comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 50% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 60% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 70% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 75% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 80% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 85% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 90% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 95% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 96% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 97% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82.In some cases, the envelope protein comprises an amino acid sequence that has at least about 98% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82. In some cases, the envelope protein comprises an amino acid sequence that has at least about 99% sequence identity to a sequence set forth in any one of SEQ ID NOs: 49-82.
Table 3. Exemplary sequences for human HERV envelope proteins
PLASMA MEMBRANE RECRUITMENT ELEMENT
[0063] In some aspects, the lipid delivery particle provided herein comprises a plasma membrane recruitment element. The lipid delivery particle disclosed herein can comprise a membrane. In some cases, the membrane encapsulates a payload. In some cases, the lipid delivery particle comprises a plasma membrane recruitment element, for example, inside the cavity of the lipid delivery particle. The plasma membrane recruitment element can localize itself to the membrane of the lipid delivery particles. The plasma membrane recruitment element can be utilized to recruit a component (e.g., a payload) to the membrane of the lipid delivery particles via forming a chimeric protein of the plasma membrane recruitment element and a component to be localized to the membrane or other mechanisms of attachment. In some cases, the membrane encapsulates a protein core. In some cases, at least a portion of the plasma membrane recruitment element forms the basic structure of the lipid delivery particle, such as a portion of the protein core inside the lipid delivery particle. In some cases, at least a portion of the plasma membrane recruitment element binds to the membrane of the lipid delivery particle from the inside.
[0064] The plasma membrane recruitment element can play a role in the assembly of the lipid delivery particle, such as packing various components (e.g., a payload) into the lipid delivery particles. The plasma membrane recruitment element can direct budding of the lipid delivery particles from a producer cell. In some cases, expressing plasma membrane recruitment element alone or together with an envelope protein disclosed herein in a producer cell can lead to formation of the lipid delivery particle.
[0065] In some cases, the plasma membrane recruitment element has a viral origin. For instance, the plasma membrane recruitment element comprises a retroviral gag protein, e.g., a retroviral polyprotein that comprises one or more of a matrix (MA) polypeptide, an RNA-binding phosphoprotein polypeptide, a capsid (CA) polypeptide, or a nucleocapsid (NC) polypeptide. The plasma membrane recruitment element can comprise HIV gag or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a gag from murine leukemia virus (MLV) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a gag from Moloney murine leukemia virus (MMLV) or a biologically active mutant thereof. In some cases, the plasma membrane recruitment element forms structural protein that forms the protein core of the lipid delivery particles described herein. The plasma membrane recruitment element can comprise Respiratory syncytial virus (RSV) M or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Human Papillomavirus (HPV) LI protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise HPV L2 protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Hepatitis B virus (HBV) core protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Hepatitis C virus (HCV) core protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise hepatitis E virus (HeV) M protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Chikungunya virus (CHIKV) C-E3-E2-6k-El or a biologically active mutant thereof. The plasma membrane recruitment element can comprise RSV NP or a
biologically active mutant thereof. The plasma membrane recruitment element can comprise Human metapneumovirus (HMPV) M or a biologically active mutant thereof. The plasma membrane can comprise a glycoprotein from a flavivirus. The flavivirus can comprise Chikungunya virus, Zika virus, Dengue virus, or West Niles virus. The plasma membrane recruitment element can comprise Zika virus (ZIKV) C or a biologically active mutant thereof. The plasma membrane recruitment element can comprise ZIKV prM/M or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Dengaue virus (DENV) C-prM or a biologically active mutant thereof. The plasma membrane recruitment element can comprise West Nile Virus (WNV) prME protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise WNV CprME protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Filovirus VP40 or Z protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Baculovirus Pl protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Rotavirus VP7 or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Rotavirus VP2 protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Rotavirus VP6 protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Porcine Circovirus Type 2 (PCV2) capsid or a biologically active mutant thereof. The plasma membrane recruitment element can comprise baculovirus VP2 protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise baculovirus VP5 protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise baculovirus VP3 protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise or baculovirus VP7 protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Ebola nucleocapsid or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Parovirus VP1 protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Parovirus VP2 protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Newcastle disease virus (NDV) M protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Human polyomavirus 2 (JCPyV) VP1 protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise Human parainfluenza virus type 3 (HPIV3) M protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise HPIV3N protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise or Mumps virus (MuV) M proteins or a biologically active mutant thereof. The plasma membrane recruitment element can comprise SARS M protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise SARS E protein or a biologically active mutant thereof. The plasma membrane recruitment element can comprise SARS N protein or a biologically active mutant thereof. [0066] In some cases, the plasma membrane recruitment element is a mammalian protein or part thereof. For example, the plasma membrane recruitment element can include a pleckstrin homology (PH) domain or a transmembrane domain of a mammalian protein, such as a mouse protein or a human protein. In
some cases, the plasma membrane recruitment element has a human origin. Utilizing the plasma membrane recruitment element of a human origin in the lipid delivery particle can give rise to reduced immunogenicity for administration to a human subject. The plasma membrane recruitment element can include a gag from human endogenous retrovirus, such as Human Endogenous Retrovirus K (e.g., HERV- K113, HERV-K101, HERV-K102, HERV-K104, HERV-K107, HERV-K108, HERV-K109, HERV- K115, HERV- KI lp22, and HERV-K12ql3) and Human Endogenous Retrovirus-W (HERV-W) or a biologically active mutant thereof. The plasma membrane recruitment element can include a hGAGKcon or a biologically active mutant thereof. The plasma membrane recruitment element can include an endogenous gag of a mammal (e.g, human) from retrotransposons (e.g., Arc from vertebrate lineage of Ty3/gypsy retrotransposon), which are also ancestral to retroviruses. In some cases, the plasma membrane recruitment element comprises a portion from human Arc.
[0067] The plasma membrane recruitment element can include a pleckstrin homology (PH) domain from a mammalian protein or a biologically active mutant thereof. The plasma membrane recruitment element can include a pleckstrin homology (PH) domain from a human protein or a biologically active mutant thereof. The PH domains can play a role in protein-membrane interactions via binding to phosphatidylinositol phosphate (PIP), for example PIP2 or PIP3, or other lipids or proteins within the membrane of the lipid delivery particles. PH domains with different sequences can have varied affinities and selectivity when binding different PIPs. The plasma membrane recruitment element can include a pleckstrin homology (PH) domain from a human protein or a mutant thereof. The PH domains can play a role in protein-membrane interactions via binding to phosphatidylinositol phosphate (PIP), for example PIP2 or PIP3, or other lipids or proteins within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more anionic lipids within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more monophosphorylated derivatives of phosphatidylinositol. In some cases, the plasma membrane recruitment element interacts with one or more bisphosphorylated derivatives of phosphatidylinositol. In some cases, the plasma membrane recruitment element interacts with one or more trisphosphorylated derivatives of phosphatidylinositol. In some cases, the plasma membrane recruitment element interacts with one or more phosphatidylinositol 4,5-bisphosphate (PI(4,5)P2) molecules or phosphatidylinositol 3,5-bisphosphate (PI(3,5)P2) molecules within a lipid layer. In some cases, the plasma membrane recruitment element interacts with one or more phosphatidylinositol(3,4,5)- trisphosphate (PIP3) molecules within a lipid layer. PH domains with different sequences can have varied affinities and selectivity when binding different PIPs.
[0068] In some cases, the plasma membrane recruitment element interacts with one or more proteins within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more G-protein coupled receptors (GPRC) within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more bg-subunits of GPRC within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more kinase proteins within the membrane of the
lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more protein kinase C (PKC) proteins within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more members of an Arf family small G- proteins within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with one or more GDP-bound form of ADP-ribosylation factor (ARF) within the membrane of the lipid delivery particles. In some cases, the plasma membrane recruitment element interacts with other proteins, protein domains, protein components, or a combination thereof within or tethered to the membrane of the lipid delivery particles.
[0069] In some embodiments, the PH domain is selected or engineered for enhanced or altered properties. In some cases, the plasma membrane recruitment element comprises an enhancement or alteration of the recruitment of a payload into a lipid delivery particle, the amount of a payload that can be encapsulated in a lipid delivery particle, release of a payload from the lipid delivery particle into a target cell, trafficking of a lipid delivery particle to a subcellular locale in a target cell, suitability of the PH for combination with other protein domains, or a combination thereof. In some cases, the plasma membrane recruitment element is engineered to enhance or alter the delivery of a payload to a target cell or target cellular compartment. In some cases, the plasma membrane recruitment element is engineered to enhance or alter the delivery of protein payload to a target cell or target cellular compartment. In some cases, the plasma membrane recruitment element is engineered to enhance or alter the delivery of a homogenous protein payload to a target cell or target cellular compartment. In some cases, the plasma membrane recruitment element is engineered to enhance or alter the delivery of a heterogenous protein payload to a target cell or target cellular compartment. In some cases, the plasma membrane recruitment element is engineered to enhance, alter, or decrease other properties and/or functions of the PH domain.
[0070] In some cases, the plasma membrane recruitment element is engineered to comprise mutations in particular structurally and functionally important regions of the PH domain. In some cases, PH domain comprises one or more mutations that affect interaction and/or binding between the PH domains and a lipid. In some cases, the plasma membrane recruitment element comprises one or more mutations that affect the lipid binding surface. In some cases, the plasma membrane recruitment element comprises one or more mutations that affect interaction and/or binding between the PH domains and a phosphatidylinositol or derivative thereof. In some cases, the plasma membrane recruitment element comprises one or more mutations that affect the binding surface for phosphatidylinositol or derivative thereof. In some cases, the plasma membrane recruitment element comprises one or more mutations that affect interaction between the PH domains and a protein. In some cases, the plasma membrane recruitment element comprises one or more mutations that affect the protein-interacting surfaces. In some cases, the plasma membrane recruitment element comprises one or more mutations that affect the other structural and functional features of the PH domains, nonlimiting examples of which can include tertiary structure, hydrophobicity, a helix architecture and configuration, [3-barrel architecture and configuration, electrostatic interactions, hydrogen-bonding, dimerization, oligomerization, any other structural and/or functional features, of a combination thereof. In some cases, the plasma membrane recruitment elements
can be screened for particular functional properties. In some cases, the plasma membrane recruitment elements can be screened for particular functional properties in a high throughput manner.
[0071] In some cases, the PH is selected from the genera: Homo, Mus, Rattus, Bos, Sus, Gallus, Danio, Canis, Xenopus, Macaca, Pan, Felis, Pongo, Dasypus, Anser, Cavia, Capra, Ovis, Meriones, Mustela, Columba, Mauremys, Callorhinchus, Pteropus, Cricetulus, Scophthalmus, Protopterus, Lates, Thunnus, Callithrix, Equus, Mesocricetus, Oryzias, Mya, Polypterus, Gallus, Drosophila, and other genera belonging to the Chordata phylum. In some cases, the plasma membrane recruitment element is selected from birds, turtles, alligators, lizards, snakes, mammals, amphibians, lungfishes, bony fishes, cartilaginous fishes, othered jawed vertebrates, or a combination thereof.
[0072] In some cases, the plasma membrane recruitment element is selected from human proteins. The plasma membrane recruitment element can include a PH domain of phospholipase C51 (e.g., human phospholipase C51) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of Aktl (e.g, human Aktl) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a mutant PH domain of human Aktl with E17K substitution or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of 3 -phosphoinositide-dependent protein kinase 1 (e.g., human 3- phosphoinositide-dependent protein kinase 1) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of Dappl (e.g., human Dappl) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of Grp 1 (e.g. , mouse Grp 1) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of human Grpl or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of OSBP (e.g., human OSBP) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of Btkl (e.g. , human Btkl) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of FAPP1 (e.g., human FAPP1) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of CERT (e.g. , human CERT) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of PKD (e.g. , human PKD) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of PHLPP1 (e.g., human PHLPP1) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of SWAP70 (e.g., human SWAP70) or a biologically active mutant thereof. The plasma membrane recruitment element can comprise a PH domain of MAPKAP1 (e.g., human MAPKAP1) or a biologically active mutant thereof.
[0073] In some cases, the plasma membrane recruitment element comprises a PH domain of SH3 domain binding protein 2 (3BP2), Rho guanine nucleotide exchange factor 28-like (AKP13), Oxysterol binding protein like 7 (OSBPL7), FYVE, RhoGEF And PH Domain Containing 6 (FGD6), Bruton agammaglobulinemia tyrosine kinase (BTK), connector enhancer of kinase suppressor of Ras 2 (CNKSR2), Rho associated coiled-coil containing protein kinase 1 (Rock 1), Rho associated coiled-coil containing protein kinase 2 (Rock 2), ceramide kinase like (CERKL), ceramide kinase (CERK), SAPK-
interacting protein 1 (SIN1), pleckstrin homology like domain family A member 2 (PHLDA2), tubby bipartite transcription factor (TUB), PH domain and leucine rich repeat protein phosphatase 1 (PHLPP1), ArfGAP with coiled-coil, ankyrin repeat and PH domains 1 (ACAP1), ArfGAP with dual PH domains 1 (AD API PHI), ArfGAP with dual PH domains 2 (AD API PH2), actin filament associated protein 1 like
2 (AFAP1L2), ArfGAP with GTPase domain, ankyrin repeat and PH domain 2 (AGAP2), A-kinase anchoring protein 13 (akapl3), AKT serine/threonine kinase 2 (AKT2), AKT serine/threonine kinase
3 (AKT3), anillin, actin binding protein (anln), amyloid beta precursor protein binding family B member
1 interacting protein (Apbblip), adaptor protein, phosphotyrosine interacting with PH domain and leucine zipper 1 (APPL1), adaptor protein, phosphotyrosine interacting with PH domain and leucine zipper 2 (APPL2), ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 2 (ARAP2), Rho GTPase activating protein 21 (ARHGAP21), Rho GTPase activating protein 25 (ARHGAP25), Rho GTPase activating protein 27 (ARHGAP27), Rho GTPase activating protein 9 (Arhgap9), Rho guanine nucleotide exchange factor 1 (ARHGEF1), Rho guanine nucleotide exchange factor 16 (Arhgefl6), Rho/Rac guanine nucleotide exchange factor 18 (Arhgefl8), Rho/Rac guanine nucleotide exchange factor 2 (ARHGEF2), Rho guanine nucleotide exchange factor 28 (ARHGEF28), Rho guanine nucleotide exchange factor 3 (Arhgef3), Rho guanine nucleotide exchange factor 4 (ARHGEF4), Rac/Cdc42 guanine nucleotide exchange factor 6 (Arhgef6), Cdc42 guanine nucleotide exchange factor 9 (Arhgef9), ArfGAP with SH3 domain, ankyrin repeat and PH domain 1 (asapl), Bruton agammaglobulinemia tyrosine kinase (btk), alkylglycerone phosphate synthase (ADPS), ceramide transfer protein (CERT), cytohesin 3 (cyth3), dual adaptor for phosphotyrosine and 3 -phosphoinositides 1 (dappl), dynamin 1 (dnml), dynamin 2 (DNM2), dedicator of cytokinesis 9 (dock9), dedicator of cytokinesis 2 (dok2), dedicator of cytokinesis 7 (Dok7), engulfment and cell motility 1 (ELM01), exocyst complex component 8 (Exoc8), FERM, ARH/RhoGEF and pleckstrin domain protein 2 (Farp2), FERM domain containing kindlin 1 (Fermtl), FERM domain containing kindlin 2 (fermt2), FERM domain containing kindlin 3 (FERMT3), FYVE, RhoGEF and PH domain containing 3 (FGD3), FYVE, RhoGEF and PH domain containing 6 (Fgd6), growth factor receptor bound protein 10 (GRB10), growth factor receptor bound protein 14 (GRB14), G protein-coupled receptor kinase 2 (GRK2), inositol polyphosphate-5 -phosphatase B (inpp5b), interaction protein for cytohesin exchange factors 1 (ipcefl), IQ motif and Sec7 domain ArfGEF 1 (IQSEC1), insulin receptor substrate 1 (IRS1), MCF.2 cell line derived transforming sequence like (Mcf21), OCRL inositol polyphosphate-5-phosphatase (OCRL), oxysterol binding protein like 11 (OSBPL11), oxysterol binding protein like 8 (OSBPL8), 3 -phosphoinositide dependent protein kinase 1 (pdpkl), phospholipase C delta 1 (plcdl), phospholipase C gamma 1 (Plcgl), pleckstrin 2 (PLEK2), pleckstrin homology domain containing Al (PLEKHA1), pleckstrin homology domain containing A2 (Plekha2), pleckstrin homology domain containing A3 (PLEKHA3), pleckstrin homology domain containing A4 (plekha4), pleckstrin homology domain containing, family A member 5 (PLEKHA5), pleckstrin homology domain containing A6 (PLEKHA6), pleckstrin homology domain containing A7 (PLEKHA7), pleckstrin homology domain containing, family B (evectins) member 1 (Plekhbl), pleckstrin homology domain containing B2 (Plekhb2), pleckstrin homology and RUN domain containing M2 (PLEKHM2), pleckstrin (PHI) (PLEK
PHI), pleckstrin (PH2) (PLEK PH2), phosphatidylinositol-3,4,5-trisphosphate dependent Rae exchange factor 1 (p-rexl), phosphatidylinositol-3,4,5-trisphosphate dependent Rac exchange factor 2 (PREX2), protein kinase D2 (PRKD2), protein kinase D3 (PRKD3), Rai GEF with PH domain and SH3 binding motif 1 (Ralgpsl), Ras association (RalGDS/AF-6) and pleckstrin homology domains 1 (RAPH1), Rho associated coiled-coil containing protein kinase 2 (Rock2), SET binding factor 1 (Sbfl), SH2B adaptor protein 2 (Sh2b2), src kinase associated phosphoprotein 1 (SKAP1), src family associated phosphoprotein 2 (Skap2), SOS Ras/Rac guanine nucleotide exchange factor 1 (Sos 1 spectrin beta, non-erythrocytic 1 (sptbnl), spectrin beta, non-erythrocytic 2 (SPTBN2), signal transducing adaptor family member 1 (stapl), SWA-70 protein (SWAP70), TBC1 domain family member 2 (TBC1D2), tec protein tyrosine kinase (TEC), TIAM Rael associated GEF 1 (Tiaml), T cell lymphoma invasion and metastasis 2 (Tiam2), trio Rho guanine nucleotide exchange factor (PHI) (TRIO), trio Rho guanine nucleotide exchange factor (PH2) (TRIO ), vav guanine nucleotide exchange factor 1 (VAV1), or mutants thereof. [0074] In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to the sequence of SEQ ID NO: 569. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to the sequence of SEQ ID NO: 569. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to the sequence set forth in any one of SEQ ID NOs: 570-596. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to the sequence set forth in any one of SEQ ID NOs: 570-596. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to any one of the sequences listed in Table 4A. In some cases, the plasma membrane recruitment element comprises any one of the sequences listed in Table 4A without the N- terminal Methionine. In some cases, the plasma membrane recruitment element comprises the sequence set forth in any one of SEQ ID NOs: 569-596 without the N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises any one of the sequences listed in Table 4A with the
N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises the sequence set forth in any one of SEQ ID NOs: 569-596 with the N-terminal Methionine.
[0075] In some cases, the plasma membrane recruitment element comprises any one of the sequences described in Table 4A with a further truncation on the N-terminus. For example, for those amino acid sequences that start with a N-terminal methionine, the N-terminal methionine can be absent. In some cases, the plasma membrane recruitment element comprises any one of the sequences described in Table 4A with atruncation (e.g., truncation of about 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 12, 14, 16, 18, 20, or more amino acids) on the C-terminus. In some cases, the plasma membrane recruitment element includes those described in Table 4A with one amino acid substitution. In some cases, the plasma membrane recruitment element comprises any one of the sequences described in Table 4A with two or more amino acid substitutions. In some cases, the plasma membrane recruitment element comprises one or more of the sequences described in Table 4A and further comprises a heterologous peptide sequence fused to the N- terminus or C-terminus.
[0076] In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to one or more of the sequences listed in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4A fused to a heterologous peptide sequence on the N terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to one or more of the sequences listed in Table 4A fused to a heterologous peptide sequence on the N terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4A fused to a heterologous peptide sequence on the C terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to one or more of the sequences listed in Table 4A fused to a heterologous peptide sequence on the C terminus.
[0077] In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A fused to a heterologous peptide sequence on the N terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A fused to a heterologous peptide sequence on the N terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A fused to a heterologous peptide sequence on the C terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4A fused to a heterologous peptide sequence on the C terminus.
[0078] In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 60% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 70% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 75% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least
about 85% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 90% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 95% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 96% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 97% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 98% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 99% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has about 100% sequence identity to any one of the sequences set forth in Table 4A. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4A.
Table 4A. RAC-alpha serine/threonine kinase 1 (AKT1) Pleckstrin Homology domains and corresponding mutations (mut.)
[0079] In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences set forth in SEQ ID NOs: 597-722. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to any one of the sequences set forth in SEQ ID NOs: 597-722. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, or about 99% or more sequence identity to any one of the sequences listed in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4B. In some cases, the plasma membrane recruitment element comprises any one of the sequences listed in Table 4B without the N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in SEQ ID NOs: 597-722 without the N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises any one of the sequences listed in Table 4B with an N- terminal Methionine. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in SEQ ID NOs: 597-722 with an N-terminal Methionine.
[0080] In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences set forth in SEQ ID NOs: 597-722. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to a one or more of the sequences set forth in SEQ ID NOs: 597-722. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, or about 99% or more
sequence identity to one or more of the sequences listed in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4B. In some cases, the plasma membrane recruitment element comprises one or more of the sequences listed in Table 4B without the N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in SEQ ID NOs: 597-722 without the N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises one or more of the sequences listed in Table 4B with an N-terminal Methionine. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in SEQ ID NOs: 597-722 with an N-terminal Methionine.
[0081] In some cases, the plasma membrane recruitment element comprises any one of the sequences described in Table 4B with a further truncation on the N-terminus. For example, for those amino acid sequences start with a N-terminal methionine, the N-terminal methionine can be absent. In some cases, the plasma membrane recruitment element comprises any one of the sequences described in Table 4B with a truncation on the C-terminus. In some cases, the plasma membrane recruitment element comprises any one of the sequences described in Table 4B with one amino acid substitution. In some cases, the plasma membrane recruitment element comprises any one of the sequences described in Table 4B with two or more amino acid substitutions. In some cases, the plasma membrane recruitment element comprises any one of the sequences described in Table 4B and a heterologous peptide sequence fused to the N-terminus or C-terminus.
[0082] In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more sequences listed in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more sequences listed in Table 4B fused to a heterologous peptide sequence on the N terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4B fused to a heterologous peptide sequence on the C terminus.
[0083] In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more,
about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4B fused to a heterologous peptide sequence on the N terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4B fused to a heterologous peptide sequence on the C terminus.
[0084] In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 60% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 70% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 75% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 85% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 90% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 95% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 96% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 97% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 98% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 99% sequence identity to any one of the sequences set forth in Table 4B. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has about 100% sequence identity to any one of the sequences set forth in Table 4B.
[0085] In some cases, the plasma membrane recruitment element is engineered to comprise one or more mutations to yield PH domain variants. In some cases, the plasma membrane recruitment element variants comprises about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to the sequence set forth in any one of SEQ ID NOs: 569-722. In some cases, the plasma membrane recruitment element is engineered to comprise one or more mutations to yield PH domain variants. In some cases, the plasma membrane recruitment element variants comprises about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to the sequence set forth in one or more of SEQ ID NOs: 569-722.
[0086] In some cases, the plasma membrane recruitment element comprises the PH domain of any one of the proteins or protein fragments set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises the PH domain of one or more of the proteins or protein fragments set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a PH domain that comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or
more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to any one of the sequences set forth in any one of SEQ ID NOs: 161-290. In some cases, the plasma membrane recruitment element comprises a PH domain that comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to one or more of the sequences set forth in any one of SEQ ID NOs: 161-290. In some cases, the plasma membrane recruitment element comprises a PH domain that comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to any one of the sequences listed in Table 4C. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that is about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more sequence identity to the PH domain of one or more of the sequences listed in Table 4C.
[0087] In some cases, the plasma membrane recruitment element comprises the protein or protein fragment of any one of the proteins or protein fragments set forth in Table 4C with a further truncation on the N-terminus. For example, for those amino acid sequences start with a N-terminal methionine, the N- terminal methionine can be absent. In some cases, the plasma membrane recruitment element comprises the protein or protein fragment of any one of the proteins or protein fragments set forth in Table 4C with a further truncation on the C-terminus. In some cases, the plasma membrane recruitment element comprises the protein or protein fragment of any one of the proteins or protein fragments set forth in Table 4C with one amino acid substitution. In some cases, the plasma membrane recruitment element comprises the protein or protein fragment of any one of the proteins or protein fragments set forth in Table 4C with two or more amino acid substitutions. In some cases, the plasma membrane recruitment element comprises the protein or protein fragment of any one of the proteins or protein fragments set forth in Table 4C and a heterologous peptide sequence fused to the N-terminus or C-terminus.
[0088] In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more sequences listed in Table 4C. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more sequences listed in Table 4C fused to a heterologous peptide sequence on the N terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about
96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4C fused to a heterologous peptide sequence on the C terminus.
[0089] In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C fused to a heterologous peptide sequence on the N terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C fused to a heterologous peptide sequence on the C terminus.
[0090] In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C fused to a heterologous peptide sequence on the N terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4C fused to a heterologous peptide sequence on the C terminus.
[0091] In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4C. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4C fused to a heterologous peptide sequence on the N terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about
85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4C fused to a heterologous peptide sequence on the C terminus.
[0092] In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 50% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 60% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 70% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 75% sequence identity to any one of the sequences set forth in Table 4C.
[0093] In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragments that comprises an amino acid sequence that has at least about 80% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 85% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 90% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 95% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 96% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 97% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 98% sequence identity to any one of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 99% sequence identity to any one of the sequences set forth in Table 4C.
[0094] In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 50% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 60% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 70% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 75% sequence identity to one or more of the sequences set forth in Table 4C.
[0095] In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragments that comprises an amino acid sequence that has at least about 80% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 85% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 90% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 95% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 96% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 97% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 98% sequence identity to one or more of the sequences set forth in Table 4C. In some cases, the plasma membrane recruitment element comprises a protein or protein fragment that comprises an amino acid sequence that has at least about 99% sequence identity to one or more of the sequences set forth in Table 4C.
[0096] In some cases, the plasma membrane recruitment element is identified and/or isolated. In some cases, the plasma membrane recruitment element comprises at least about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or
more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to the sequence set forth in any one of SEQ ID NOs: 161-290. In some cases, the plasma membrane recruitment element is engineered to comprise one or more mutations to yield PH domain variants. In some cases, the plasma membrane recruitment element variants comprises at least about 60% or more, about 65% or more, about 70% or more, about 75% or more, about 80% or more, about 85% or more, about 90% or more, about 95% or more, about 96% or more, about 97% or more, about 98% or more, about 99% or more, or about 100% sequence identity to the sequence set forth in any one of SEQ ID NOs: 161-290.
[0097] The plasma membrane recruitment element can also include a membrane protein (e.g. , a human membrane protein), a transmembrane domain thereof, or a biologically active mutant thereof. For example, the transmembrane domain of a human protein can be a tetraspanin or a biologically active mutant thereof. In some cases, the plasma membrane recruitment element comprises a transmembrane domain of human CD9 or a biologically active mutant thereof. In some cases, the plasma membrane recruitment element comprises a transmembrane domain of human CD47 or a biologically active mutant thereof. In some cases, the plasma membrane recruitment element comprises a transmembrane domain of human CD63 or a biologically active mutant thereof. In some cases, the plasma membrane recruitment element comprises a transmembrane domain of human CD81, or a biologically active mutant thereof. [0098] The plasma membrane recruitment element can comprise a retroviral gag or a biologically active mutant thereof. The mutant of a retroviral gag can include only a portion of the retroviral gag. The plasma membrane recruitment element can include a gag of an alpha retrovirus or a biologically active mutant thereof. The plasma membrane recruitment element can a beta retrovirus or biologically active mutant thereof. The plasma membrane recruitment element can include a gamma retrovirus or biologically active mutant thereof. The plasma membrane recruitment element can include a delta retrovirus or biologically active mutant thereof. The plasma membrane recruitment element can include or biologically active mutant thereof. The plasma membrane recruitment element can include an epsilon retrovirus or biologically active mutant thereof. The plasma membrane recruitment element can include a spumavirus or biologically active mutant thereof. The retroviral gag can include a gag of HIV (e.g., HIV-1), a gag of murine leukemia virus (MLV), a gag of Moloney murine leukemia virus (MMLV), a gag of Simian immunodeficiency virus (SIV), a gag of Rous sarcoma virus (RSV), a gag of human T-cell leukemia virus type-1 (HTLV), or a gag of bovine leukemia virus (BLV), or a biologically active mutant thereof. The plasma membrane recruitment element can include a gag of HIV (e.g., HIV-1) or a biologically active mutant thereof. The plasma membrane recruitment element can include a gag of MLV or a biologically active mutant thereof. The plasma membrane recruitment element can include a gag of RSV or a biologically active mutant thereof. The plasma membrane recruitment element can include a gag of Friend murine leukemia virus (FMLV) or biologically active mutant thereof.
[0099] In some cases, the envelope protein comprises one or more of the sequences set forth in Table 4D with at least one amino acid substitution, deletion, or insertion. For instance, N-terminal methionine can be absent from the envelope protein of the lipid delivery particle provided herein relative to the wild-type viral envelope protein. In some cases, the envelope protein comprises one or more of the sequences set forth in Table 4D and a heterologous peptide sequence fused to the N-terminus or C-terminus. In some cases, the plasma membrane recruitment element comprises any one of the sequences described in Table
4D with a further truncation on the N-terminus. For example, forthose amino acid sequences start with a N-terminal methionine, the N-terminal methionine can be absent. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with a further truncation on the C-terminus. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with one amino acid substitution. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with two or more amino acid substitutions. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D and a heterologous peptide sequence fused to the N-terminus or C- terminus.
[0100] In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with a further truncation on the N-terminus. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with a further truncation on the C-terminus. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with one amino acid substitution. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with two or more amino acid substitutions. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth Table 4D and a heterologous peptide sequence fused to the N-terminus or C-terminus.
[0101] In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4D. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to one or more of the sequences listed in Table 4D fused to a heterologous peptide sequence on the N terminus. In some cases, the plasma membrane recruitment element comprises an amino acid sequence comprising about 60%, about 65%, about 70%, about 75%, about 80%, about 85%, about 90%, about 95%, about 96%, about 97%, about 98%, about 99%, or about 100% sequence identity to any one of the sequences listed in Table 4D fused to a heterologous peptide sequence on the C terminus.
[0102] In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with at least one amino acid substitution, deletion, or insertion. For instance, N- terminal methionine can be absent from the plasma membrane recruitment element of the lipid delivery particle provided herein relative to the wild-type viral envelope protein. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D and a heterologous peptide sequence fused to the N-terminal or C-terminal.
[0103] In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with a further truncation on the N-terminus. For example, for those amino acid sequences start with a N-terminal methionine, the N-terminal methionine can be absent. In some cases,
the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with a further truncation on the C-terminus. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with one amino acid substitution. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D with two or more amino acid substitutions. In some cases, the plasma membrane recruitment element comprises any one of the sequences set forth in Table 4D and a heterologous peptide sequence fused to the N-terminal or C-terminal. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with a further truncation on the N-terminus. For example, for those amino acid sequences start with a N-terminal methionine, the N-terminal methionine can be absent. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with a further truncation on the C-terminus. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with one amino acid substitution. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D with two or more amino acid substitutions. In some cases, the plasma membrane recruitment element comprises one or more of the sequences set forth in Table 4D and a heterologous peptide sequence fused to the N-terminal or C-terminal.
[0104] In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to any one of the sequences set forth in Table 4D. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 50% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 60% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48 In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 70% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 75% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. [0105] In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 80% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 85% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 90% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma
membrane recruitment element comprises an amino acid sequence that has at least about 95% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 96% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 97% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 98% sequence identity to a sequence set forth in any one of SEQ ID NOs: 1-48. In some cases, the plasma membrane recruitment element comprises an amino acid sequence that has at least about 99% sequence identity to a sequence set forth in any one of
SEQ ID NOs: 1-48
Table 4D. Exemplary plasma membrane recruitment elements and their sequences
*hGAGKcon is a consensus sequence derived from ten proviral GAG sequences encoded by human genomic sequences. The GAG sequences used to derive this consensus GAG sequence are from the following HERVs: HERV-K113, HERV-K101, HERV-K102, HERV-K104, HERV-K107, HERVK108, HERV-K109, HERV-K115, HERV- KI lp22, and HERV-K12ql3.
[0106] In some cases, the lipid delivery particle disclosed herein comprises a protein core that is composed of at least a structural protein of a viral origin, for instance, a retroviral gag protein. In some of these cases, the lipid delivery particle comprises a retroviral gag-pro-pol polyprotein, e.g., a gag-pro-pol poly protein from HIV, MMLV, or FMLV, which can help assemble a protein core of the lipid delivery particle. In some of these cases, some of the gag-pro-pol polyprotein is cleaved, e.g., by pro (protease) present freely or in the gag-pro-pol polyprotein. Without wishing to be bound by any particular theory, the cleavage by pro can be inefficient, and the resultant cleavage products can include gag polyprotein, gag-pro polyprotein, free pro, and free pol (polymerase). In some cases, a retroviral gag polyprotein can be further cleaved into MA, CA, NC, and other small fragments, if any. In some other cases, the lipid delivery particle comprises a retroviral gag-pro polyprotein without the pol component, and the gag-pro polyprotein can help form a protein core of the lipid delivery particle. The gag -pro can also be cleaved by pro, in some cases, inefficiently, into separate gag and pro proteins. In some cases, there can be different plasma membrane recruitment elements in a lipid delivery particle. For instance, a gag -pro or gag-pro-pol polyprotein from one species of virus (e.g., a retrovirus, e.g., a HIV) can help assemble form a protein core of the lipid delivery particle, while a chimeric protein in the lipid delivery particle, discussed infra, can comprise a payload fused with a gag protein from a different species of virus (e.g., an MMLV), or from a HERV, or a PH domain or transmembrane domain of a huma protein (e.g. , a PH domain of human Aktl with E17K substitution).
CHIMERIC PROTEIN
[0107] In aspects, the present disclosure provides a chimeric protein comprising a plasma membrane recruitment element and a payload that is a protein or a fragment thereof. In some aspects, the lipid delivery particle comprises a chimeric protein comprising a plasma membrane recruitment element and a payload that is a protein or a fragment thereof. In some cases, the plasma membrane recruitment element and the payload are fused directly in the chimeric protein. In other cases, the plasma membrane recruitment element and the payload are fused indirectly via a linker. In some cases, the linker between the plasma membrane recruitment element and the payload is a cleavable linker that is recognized by a protease.
[0108] The chimeric protein (e.g., comprising a gag protein) can form at least part of a protein core of the lipid delivery particle. A lipid delivery particle can comprise two or more chimeric proteins. The chimeric protein can include a structural protein. The structural protein can comprise a plasma membrane recruitment element (e.g., retroviral gag protein). The plasma membrane recruitment element can be fused to a payload. In some cases, the two or more chimeric proteins comprise the same structural protein. In some cases, the two or more chimeric proteins comprise different structural proteins. In some cases, the
two or more chimeric proteins comprise different payloads. In some cases, the chimeric protein comprises a payload that comprises a nucleic acid-binding moiety. In some cases, the payload further comprises a guide nucleic acid molecule that forms a ribonucleoprotein complex with the nucleic acid-binding moiety. In some cases, the chimeric protein is suitable for delivery by a lipid delivery particle disclosed herein. [0109] In some cases, the lipid delivery particle of the present disclosure further comprises a protease that recognizes the cleavable linker in the chimeric protein and cuts the chimeric protein at the cleavable linker. As a result of the cleavage at the cleavable linker by the protease, the payload can be separated from the plasma membrane recruitment element. In some cases, the payload is present as a "free" entity separate from the plasma membrane recruitment element. For instance, the payload can be free and present within an inside of the protein core of the lipid delivery particle. In some cases, the protease is part of a second chimeric protein comprising a second plasma membrane recruitment element and the protease, where the second plasma membrane recruitment element can be either different from or same as the plasma membrane recruitment element that is fused with the payload.
[0110] In some cases, the chimeric protein disclosed herein also comprises one or more non-cleavable linkers that operably link components together. The non-cleavable linker can be any suitable linker sequence that is used for chimeric protein construction, such as peptide linkers that consist of glycine (Gly) and serine (Ser) residues. In some embodiments, the non-cleavable linker comprises an amino acid sequence selected from the group consisting of: (GS)x (SEQ ID NO: 564), (GGS)x (SEQ ID NO: 565), (GGGGS)x (SEQ ID NO: 566), (GGSG)x (SEQ ID NO: 567), and (SGGG)x (SEQ ID NO: 568), and wherein x is an integer from 1 to 50.
[oni] In some cases, the chimeric protein of the present disclosure comprises a nuclear export signal (NES) sequence that can direct transport of the chimeric protein out of the nucleus of a cell, e.g. , a producer cell.
[0112] In some cases, the chimeric protein disclosed herein has one of the following configurations of components positioned in an order from N-terminus to C-terminus:
[plasma membrane recruitment element]-[cleavable linker] -[payload];
[plasma membrane recruitment element]-[n * NES] -[cleavable linker]-[payload]; [plasma membrane recruitment element]-[cleavable linker] -[payload] -[n * NES]; [plasma membrane recruitment element]-[cleavable linker]-[n * NES]-[payload];
[plasma membrane recruitment element]-[cleavable linker l]-[payload]- [cleavable linker 2]-[n * NES]; [payload] -[cleavable linker]-[n * NES]-[plasma membrane recruitment element];
[payload]-[n * NES]-[cleavable linker] -[plasma membrane recruitment element]; [n * NES] -[payload] -[cleavable linker] -[plasma membrane recruitment element];
[n * NES] -[cleavable linker l]-[payload]-[cleavable linker 2] -[plasma membrane recruitment element]; and
[payload] -[cleavable linker] -[plasma membrane recruitment element]; wherein n is an integer in the range of from 1 to 10, and denotes the number of repeats of the NES sequence. Non-cleavable linker sequence can be present or absent in any of the foregoing configurations
between any two neighboring components. As provided herein, the payload sequence in the chimeric protein can have one or more NLS sequences, at its N-terminus, C-terminus, or both.
[n * NES] -[cleavable linker l]-[payload]-[cleavable linker 2] -[plasma membrane recruitment element]; and
[payload] -[cleavable linker] -[plasma membrane recruitment element]; wherein n is an integer in the range of from 1 to 10, and denotes the number of repeats of the NES sequence. Non-cleavable linker sequence can be present or absent in any of the foregoing configurations between any two neighboring components. As provided herein, the payload sequence in the chimeric protein can have one or more NLS sequences, at its N-terminus, C-terminus, or both.
Nuclear Export Signal
[0113] Direction of nuclear transport within the cell can be governed by nuclear targeting signals within payload proteins or coupled to (e.g., fused with) the payload proteins. As used herein, the term “nuclear export signal” refers to a sequence of amino acids that targets a payload protein for export from the nucleus. In some cases, a nuclear export signal (NES) is a short target peptide sequence containing four hydrophobic residues. These residues target the protein for export from the nucleus to the cytoplasm through the nuclear pore complex. A chimeric protein provided herein can comprise 1 NES, 2 NESs, 3 NESs, 4 NESs, 5 NESs, 6 NESs, 7 NESs, 8 NESs, 9 NESs, or 10 NESs. In some cases, the NES is located at the N-terminus, C-terminus, or in an internal region of the chimeric protein. In some cases, a NES is coupled between the plasma membrane recruitment element and the payload in the chimeric protein. In some cases, there is a cleavable linker between the plasma membrane recruitment element and the payload in the chimeric protein, and one or more NESs present on the same of the cleavable linker as the plasma membrane recruitment element.
[0114] In some cases, the NES sequence that is used in the chimeric protein comprises LQLPPLERLTL (SEQ ID NO: 403) derived from HIV-1 Rev protein, or any of the sequences having at least 80% identity thereto. In some cases, the NES sequence comprises LALKLAGLDI (SEQ ID NO: 352) or NELALKLAGLDI (SEQ ID NO: 416), derived from PKIa, or any of the sequences having at least 80% identity thereto. In some cases, the NES sequence that is used in the chimeric protein comprises an amino acid sequence as set forth in Table 5. In some cases, the NES sequence comprises any one of the sequences set forth in Table 5. In some cases, the NES sequence comprises one or more of the sequences set forth in Table 5. In some cases, the NES sequence comprises more than one, more than two, more than three, more than four, more than five, more than six, more than seven, more than eight, more than nine, or more than ten of the sequences set forth in Table 5. In some cases, the NES sequences comprises multiple sequences set forth in Table 5.
[0115] In some cases, the NES sequence comprises an amino acid sequence having 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any sequence listed in Table 5. In some cases, the NES sequence comprises an amino acid sequence having 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any sequence set forth in SEQ ID NOs: 353-453. In some cases, the NES sequence described herein comprises a sequence with greater than
80% sequence identity to any sequence listed in Table 5. The transport of payload proteins within a cell is enabled through both NES and nuclear export receptors. In some cases, the NES described herein is associated with a nuclear export receptor (e.g., CRM-1). In some cases, the NES may be conditionally active or inactive. In some cases, the NES sequence disclosed herein comprises a sequence such as those described in T la Cour, et al., Nucleic Acids Res. 2003;31 ( 1): 393-396; and Xu D, et al. Mol Biol Cell. 2012 Sep;23(18):3673-6, each of which is incorporated herein by reference in its entirety. Any of the NES sequences described in the NES sequence database (NESdb°; prodata.swmed.edu/LRNes) or (NESbase; services.healthtech.dtu.dk/datasets/NESbase-1.0) can be used in a chimeric protein disclosed herein, e.g., for the purpose of packaging a payload into the molecular assembly, e.g., the lipid delivery particle.
[0116] In some cases, a chimeric protein disclosed herein include a nuclear export sequence (NES). In some cases, the NES facilitates localization of the chimeric protein in the cytosol of a target cell relative to the nucleus.
[0117] In some cases, a chimeric protein disclosed herein includes at least one NES sequences, such as, 2 or more, 3 or more, 4 or more, or 5 or more NES sequences. In some cases, one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g., within 50 amino acids of) the N-terminus and/or the C- terminus of the chimeric protein. In some cases, the chimeric protein disclosed herein comprises only one NES sequence. In some cases, the chimeric protein disclosed herein comprises two NES sequences. In some cases, the chimeric protein disclosed herein comprises three NES sequences. In some cases, one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g. , within 50 amino acids of) the N- terminus of the chimeric protein. In some cases, one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g. , within 50 amino acids of) the C- terminus of the chimeric protein. In some cases, one or more NES sequences (3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g., within 50 amino acids of) both the N-terminus and the C-terminus of the chimeric protein. In some cases, an NES sequence is positioned at the N- terminus and an NES sequence is positioned at the C-terminus of the chimeric protein.
[0118] In some cases, a payload is a protein that is delivered as part of the chimeric protein disclosed herein, e.g., operably linked to a structural protein (e.g., human endogenous retroviral structural protein or a Plasma membrane recruitment element). In some embodiments, the one or more NES sequences are positioned at or near the one or both ends of the payload protein sequence inside the chimeric protein. For example, in some cases, one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g., within 50 amino acids of) the N-terminus and/or the C- terminus of the payload protein sequence. In some cases, one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g. , within 50 amino acids of) the N-terminus of the payload protein sequence. In some cases, one or more NES sequences (2 or more, 3 or more, 4 or more, or 5 or more NES sequences) are positioned at or near (e.g. , within 50 amino acids of) the C-terminus of the payload protein sequence. In some cases, one or more NES sequences (3 or more, 4
or more, or 5 or more NES sequences) are positioned at or near (e.g. , within 50 amino acids of) both the N-terminus and the C-terminus of the payload protein sequence. In some cases, an NES sequence is positioned at the N-terminus and an NES sequence is positioned at the C-terminus of the payload protein sequence. In some cases, the chimeric protein disclosed herein comprises only one NES sequence. In some cases, the chimeric protein comprises only one NES sequence, and the NES sequence is positioned at or near (e.g., within 50 amino acids of) the N-terminus of the payload protein.
[0119] In eukaryotic cells, transport of proteins between the nucleus and the cytoplasm can be mediated by transport factors in the karyopherin-[3 family, which are also known as importins and exportins. The direction of nuclear-cytoplasmic transport can be dictated by nuclear targeting signals within the payload proteins. Nuclear export sequences (NESs) can direct export of proteins from the nucleus to the cytoplasm. NESs can bind directly to the export karyopherin CRM1 (also known as exportin 1), which can escort payload proteins through the nuclear pore complex.
Nuclear Localization Signal
[0120] In some instances, a payload described herein comprises one or more nuclear localization sequences (NLS). As used herein, the term “nuclear localization signal” refers to a sequence of amino acids that targets a payload (e.g. , a protein or a short polypeptide), which the NLS is present within or coupled to, to localize to the nucleus. In some cases, an NLS facilitates the import of a polypeptide comprising an NLS into the cell nucleus. A polypeptide can comprise 1 NLS, 2 NLSs, 3 NLSs, 4 NLSs, 5 NLSs, 6 NLSs, 7 NLSs, 8 NLSs, 9 NLSs, or 10 NLSs. In some cases, the NLS is located at the N- terminus, C-terminus, or in an internal region of the polypeptide. In some cases, a NLS is coupled to a nucleic acid binding domain described elsewhere herein. In some cases, a NLS is coupled to a nucleic acid modifying domain described elsewhere herein. In some cases, a NLS is coupled to a guidable polypeptide domain, a deaminase domain, or a reverse transcriptase domain. In some cases, a NLS is covalently linked to a nucleic acid binding domain described elsewhere herein. In some cases, a NLS is covalently linked to a nucleic acid modifying domain described elsewhere herein. In some cases, a NLS is covalently linked to a guidable polypeptide domain, a deaminase domain, or a reverse transcriptase domain. In some cases, a nucleic acid binding domain does not comprise an NLS. In some cases, a nucleic acid binding domain does not comprise an NLS. In some cases, a guidable polypeptide domain, a deaminase domain, or a reverse transcriptase domain does not comprise an NLS. Examples of NLS are provided in Table 6 below.
[0121] In some cases, the NLS comprises an amino acid sequence as set forth in Table 6. In some cases, the NLS comprises any one of the sequences set forth in Table 6. In some cases, the NLS comprises one or more of the sequences set forth in Table 6. In some cases, the NLS comprises more than one of the
sequences set forth in Table 6. In some cases, the NLS comprises multiple sequences set forth in Table 6. In some cases, NLS sequence can comprise an amino acid sequence having 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any sequence listed in Table 6. In some cases, the NLS sequence described herein can comprise a sequence with greater than 80% sequence identity to any sequence listed in Table 6. In some cases, NLS sequence can comprise an amino acid sequence having 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any sequence set forth in SEQ ID NOs: 454-477.
[0122] In some cases, a chimeric protein disclosed herein includes a nuclear localization sequence (NLS). In some cases, the NLS facilitates delivery of the chimeric protein, or a payload released from the chimeric protein (for instance, released from the chimeric protein following cleavage of a cleavable linker), into the nucleus of a target cell.
[0123] In some cases, a payload is a protein and is delivered as part of the chimeric protein disclosed herein, e.g., operably linked to a structural protein (e.g., plasma membrane recruitment element). In some embodiments, the one or more NLS sequences are positioned at or near the one or both ends of the payload protein sequence of the chimeric protein. In some cases, a chimeric protein includes (e.g., is fused to) between 2 and 5 NLS sequences (e.g., 2-4, or 2-3 NLSs). Examples of NLS sequences include an NLS sequence derived from: the NLS of the SV40 virus large T-antigen, having the amino acid sequence PKKKRKV (SEQ ID NO: 468); the NLS from nucleoplasmin (e.g., the nucleoplasmin bipartite NLS with the sequence KRPAATKKAGQAKKKK (SEQ ID NO: 460); the c-myc NLS having the amino acid sequence PAAKRVKLD (SEQ ID NO: 467) or RQRRNELKRSP (SEQ ID NO: 541); the hRNPAl M9 NLS having the sequence NQSSNFGPMKGGNFGGRSSGPYGGGGQYFAKPRNQGGY (SEQ ID NO: 542); the sequence RMRIZFKNKGKDTAELRRRRVEVSVELRKAKKDEQILKRRNV (SEQ ID NO: 543) of the IBB domain from importin-alpha; the sequences VSRKRPRP (SEQ ID NO: 477) and PPKKARED (SEQ ID NO: 544) of the myoma T protein; the sequence PQPKKKPL (SEQ ID NO: 545) of human p53; the sequence SALIKKKKKMAP (SEQ ID NO: 546) of mouse c-abl IV; the sequences DRLRR (SEQ ID NO: 547) and PKQKKRK (SEQ ID NO: 548) of the influenza virus NS1; the sequence RKLKKKIKKL (SEQ ID NO: 549) of the Hepatitis virus delta antigen; the sequence REKKKFLKRR (SEQ ID NO: 550) of the mouse Mxl protein; the sequence KRKGDE VDGVDEV AKKKS KK (SEQ ID NO: 551) of the human poly(ADP-ribose) polymerase; and the sequence RKCLQAGMNLEARKTKK (SEQ ID NO: 552) of the steroid hormone receptors (human) glucocorticoid, and sequences having at least 80% identity to the foregoing. In some cases, an NLS comprises the amino acid sequence MDSLLMNRRKFLY QFKNVRWAKGRRETYLC (SEQ ID NO: 553).
[0124] Other examples of an NLS sequence include KRTADGSEFESPKKKRKV (SEQ ID NO: 462), KKTELQTTNAENKTKKL (SEQ ID NO: 554), KRGINDRNFWRGENGRKTR (SEQ ID NO: 555), RKSGKIAAIVVKRPRK (SEQ ID NO: 556), and MDSLLMNRRKFLY QFKNVRWAKGRRETYLC (SEQ ID NO: 463), SPKKKRKVEAS (SEQ ID NO: 557), encoded by AGCCCCAAGAAgAAGAGaAAGGTGGAGGCCAGC (SEQ ID NO: 558), GPKKKRKVAAA (SEQ ID NO: 559), as well as any of those described in Cokol et al., EMBO Rep., 2000, 1(5): 411-415 and
Freitas et al., Current Genomics, 2009, 10(8): 550-7; Lu, J., et la., Cell Commun Signal 19, 60 (2021); international publication no. WO/2001/038547, each of which is incorporated herein by reference in its entirety, and sequences having at least 80% identity to the foregoing.
[0125] In some embodiments, the chimeric protein comprises one NES sequence and two NLS sequences. In some cases of these embodiments, the NES sequence, NLS sequences, and the payload protein sequence are positioned in an order from N-terminus to C-terminus as follows: NES-NLS-payload protein-NLS. In some embodiments, the chimeric protein comprises two or more NES sequences and two NLS sequences. In some cases of these embodiments, the NES sequences, NLS sequences, and the payload protein sequence are positioned in an order from N-terminus to C-terminus as follows: n X NES
(n >=2)-NLS-payload protein-NLS.
Cleavable Linker
[0126] In some cases, the chimeric protein comprises a cleavable linker in between two or more components. For instance, the chimeric protein can comprise a cleavable linker between a payload protein sequence and a plasma membrane recruitment element sequence (e.g., retroviral gag protein sequence). In some cases, the cleavable linker separates the plasma membrane recruitment element sequence from a NLS sequence, and/or a NES sequence at its N-terminus or C-terminus. The cleavable linker can separate the payload protein sequence from the plasma membrane recruitment element sequence, NLS sequence, and/or NES sequence at its N-terminus or C-terminus. Examples of cleavable linker sequences that can be used in the chimeric protein include TSTLLMENSS (SEQ ID NO: 560), PRSSLYPALTP (SEQ ID NO: 561), VQALVLTQ (SEQ ID NO: 562), and PLQVLTLNIERR (SEQ ID NO: 563), and sequences having at least 80% identity to any one of the foregoing.
PAYLOAD
[0127] A payload in a lipid delivery particle of the present disclosure can comprise a protein, a polypeptide, a nucleic acid (e.g., DNA or RNA), or any combinations thereof.
[0128] The payload can be a part of the chimeric protein disclosed herein or can comprise a part of the chimeric protein disclosed herein. Alternatively or additionally, the payload can include an entity in the lipid delivery particle separate from the chimeric protein disclosed herein. For instance, in some cases, the payload is a protein or polypeptide coupled to a plasma membrane recruitment element. In some cases, the payload comprises a first moiety (e.g, a nucleic acid-binding protein) that is fused to a plasma membrane recruitment element, and further comprises a second moiety that is coupled to the first moiety via covalent or non-covalent interaction. For instance, the first moiety can be a nucleic acid binding protein that is fused with the plasma membrane recruitment element, and the second moiety can be a nucleic acid molecule that binds to the nucleic acid binding protein.
[0129] In some cases, a payload is directly packaged within the lipid delivery particles and delivered into a target cell in its free form. In some cases, a payload can be fused to a plasma membrane recruitment element (e.g., pleckstrin homology domain) and form a chimeric protein as part of the lipid delivery particles, and then delivered into the target cell. In some cases, the plasma membrane recruitment element (e.g., pleckstrin homology domain) forms at least part of a protein core of the lipid delivery particle. In some embodiments, the payload in its free form or as part of a chimeric protein is within the inside cavity
of the protein core of the lipid delivery particles disclosed herein. In some cases, the payload in its free form derives from a cleavage of the chimeric protein comprising the payload.
[0130] In some cases, a lipid delivery particle can deliver more than one payload. Each of the payloads can independently comprise nucleic acid-binding moiety, a nucleic acid-modifying moiety, a fusion protein, or a nucleic acid, or any combinations thereof.
[0131] In some embodiments, the plasma membrane recruitment element and the payload are coupled via any suitable method. Covalent coupling between the plasma membrane recruitment element and a payload peptide can include inteins that can form peptide bonds, direct protein-protein chimeras generated from a single reading frame. In some cases, nucleic acids base pairing to other nucleic acids via hydrogen bonding interactions (e.g., DNA/RNA, DNA/DNA, or RNA/RNA hybrids), protein-protein binding, or protein-nucleic acid molecule binding can be involved for the coupling between the plasma membrane recruitment element and the payload. Examples of protein-nucleic acid molecule binding include an RNA binding protein (RBP) and an RBP binding sequence (e.g., an RNA) that binds to the RBP. In some embodiments, each of the plasma membrane recruitment element and the payload is fused to a heterologous sequence, and the two heterologous sequences dimerize or multimerize with or without the need for a chemical compound to induce the protein-protein binding, such as a single -stranded nucleic acid sequence or protein dimerization domains). In some embodiments, each of the plasma membrane recruitment element and the payload is fused to one member of a pair of binding partners (e.g., antibody and its target antigen). In some embodiments, the plasma membrane recruitment element is fused to an RBP, and the payload is fused to a RBP binding sequence. Examples of suitable protein domains or nucleic acid molecules for forming the non-covalent connections include single chain variable fragments, nanobodies, affibodies, DmrA/DmrB/DmrC, FKBP/FRB, dDZFs, Leucine zippers, proteins that bind to DNA and/or RNA, optogenetic protein domains that can dimerize or multimerize in the presence of certain light wavelengths, proteins with quaternary structural interactions, and/or naturally reconstituting split proteins. Examples of RBPs and their RBP binding sequences that can be used include a sequence having at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in Table 7. Examples of RBPs and their RBP binding sequences that can be used include a sequence having at least about 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity to a sequence set forth in any one of SEQ ID NOs: 478- 513. In some cases, the RBP comprises an amino acid sequence as set forth in Table 7. In some cases, the RBP comprises any one of the sequences set forth in Table 7. In some cases, the RBP comprises one or more of the sequences set forth in Table 7. In some cases, the RBP comprises more than one of the sequences set forth in Table 7. In some cases, the RBP comprises multiple sequences set forth in Table 7. In some cases, the RBP binding sequence comprises an amino acid sequence as set forth in Table 7. In some cases, the RBP binding sequence comprises any one of the sequences set forth in Table 7. In some cases, the RBP binding sequence comprises one or more of the sequences set forth in Table 7. In some
cases, the RBP binding sequence comprises more than one of the sequences set forth in Table 7. In some cases, the RBP binding sequence comprises multiple sequences set forth in Table 7.
Nucleic acid binding domains and nucleic acid modifying domains
[0132] In some cases, the payload comprises a nucleic acid-binding moiety, a nucleic acid-modifying moiety, a fusion protein, or a nucleic acid. In some cases, the payload comprises a nucleic acid-binding domain, e.g., a DNA-binding protein domain or polypeptide or an RNA-binding domain or polypeptide e.g., an RNA-binding protein (RBP). A nucleic acid-binding moiety can be capable of binding a nucleic acid. A nucleic acid-binding domain can bind to a nucleic acid in a nonspecific or a site-specific manner. [0133] In some cases, the nucleic acid-binding moiety binds to a nucleic acid in a site-specific manner. For example, a nucleic acid-binding moiety can comprise an aptamer binding domain that selectively binds to a specific target. In some cases, a nucleic acid-binding moiety recognizes a specific recognition sequence in the target nucleic acid. In some cases, a nucleic acid-binding moiety comprises an aptamer binding domain. In some cases, a nucleic acid binding moiety selectively binds to a sequence or a structural element in a nucleic acid molecule. In some cases, an RNA-binding domain selectively binds to a specific sequence motif in an RNA molecule. In some cases, a nucleic acid-binding moiety selectively binds to a structural element in a nucleic acid molecule. For example, a nucleic acid-binding domain can bind to a stem -loop in a nucleic acid molecule.
[0134] In some cases, a nucleic acid-binding moiety is or comprises a guidable polypeptide domain, a transcriptional regulatory domain, or a nucleic acid-modifying domain. A guidable polypeptide domain can be capable of binding to a polynucleotide (e.g. an RNA guide) that can direct the guidable polypeptide domain a target site. In some cases, the guidable polypeptide domain forms a complex with the RNA guide and recognizes the target sequence through DNA-RNA base pairing. In some cases, a nucleic -acid binding moiety is or comprises a transcriptional regulatory domain. In other cases, a nucleic- binding moiety can help recruit a transcriptional repressor or activator to a target site. In some cases, a
nucleic acid-binding moiety is or comprises a nucleic acid-modifying moiety. In some cases, the present disclosure uses nucleic acid-binding moieties to recruit a nucleic acid-modifying moiety to a target site. In some cases, a nucleic-acid binding moiety comprises catalytic activity. In other cases, a nucleic acidbinding moiety is catalytically inactive. In some cases, a nucleic-acid binding moiety comprising catalytic activity is modified to have a reduced level of activity compared to its wild-type counterpart.
[0135] In some cases, the payload in the present disclosure comprises a nucleic acid modifying domain. A nucleic acid-modifying domain can comprise a polypeptide domain, a nucleic acid or a combination thereof (e.g., a ribonucleoprotein complex). A nucleic acid-modifying domain can be capable of modifying nucleic acid, such as cleaving double-stranded nucleic acid; nicking a single -stranded nucleic acid; introducing a mutation, deletion, or insertion in a nucleic acid; methylating or demethylating a nucleic acid, or altering the structure of DNA (e.g., changing chromatin structure through modifying histones). For example, a nucleic acid modifying domain can comprise a nuclease domain, a nickase domain, a deaminase domain, a polymerase, reverse transcriptase domain, a recombinase domain, a transposase domain, or an epigenetic modifying domain. A nuclease domain can be capable of cleaving phosphodiester bonds between nucleotides in nucleic acids. A nuclease domain can comprise an exonuclease (e.g., a nuclease capable of cleaving nucleic acids from the ends) or an endonuclease (e.g., a nuclease capable of cleaving nucleic acids in the middle). In some cases, a nucleic acid modifying effector or nucleic acid binding domain is a nickase, which can be capable of cleaving a single-strand in a double -stranded DNA. Nucleic acid modifying domains can be useful for gene editing, or for regulating, activating, or inhibiting gene expression.
Guidable polypeptide domain
[0136] In some cases, the payload in the present disclosure comprises a guidable polypeptide domain (e.g., a CRISPR-Cas protein domain). In some cases, a guidable polypeptide domain is capable of binding to a polynucleotide (e.g. , an RNA guide) that directs it to a target site . In some cases, the guidable polypeptide domain forms a complex with the polynucleotide and recognizes the target sequence through DNA-RNA base pairing.
[0137] In some cases, a guidable polypeptide domain is a CRISPR/CRISPR-associated (Cas) domain. A CRISPR domain can be a natural or an engineered domain. A Cas protein or domain can be derived from a CRISPR system or share structural and/or functional similarities to a protein involved in a CRISPR system.
[0138] In some cases, the guidable polypeptide domain is any suitable nuclease, e.g., a CRISPR- associated (Cas) protein or a Cas nuclease which functions in a non-naturally occurring CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)/Cas (CRISPR-associated) system. In bacteria, this system can provide adaptive immunity against foreign DNA (Barrangou, R., et al, “CRISPR provides acquired resistance against viruses in prokaryotes,” Science (2007) 315: 1709-1712; Makarova, K.S., et al, “Evolution and classification of the CRISPR-Cas systems,” Nat Rev Microbiol (2011) 9:467- 477; Gameau, J. E., et al, “The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid
DNA,” Nature (2010) 468:67-71 ; Sapranauskas, R., et al, “The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coh,” Nucleic Acids Res (2011) 39: 9275-9282).
[0139] Suitable nucleases include CRISPR-associated (Cas) proteins or Cas nucleases including type I CRISPR-associated (Cas) polypeptides, type II CRISPR-associated (Cas) polypeptides (e.g., Cas9 or Cas 14), type III CRISPR-associated (Cas) polypeptides, type IV CRISPR-associated (Cas) polypeptides, type V CRISPR-associated (Cas) polypeptides (e.g., Cpfl/Casl2a, C2cl, or c2c3), and type VI CRISPR- associated (Cas) polypeptides (e.g., C2c2/Casl3a, Cas 13b, Cas 13c, Cas 13d).
[0140] A CRISPR system is a system encoding DNA sequence arrays known as clustered regularly interspaced short palindromic repeats (CRISPRs), which can be found in microbial genomes or phage genomes. In some cases, CRISPR systems comprise genes encoding CRISPR-associated (Cas) proteins and/or small RNA guide molecules (e.g., crRNA or tracrRNA) that assemble with the CRISPR domain. In some cases, the CRISPR-Cas domain forms a complex with one or more RNA guide molecules to form an effector ribonucleoprotein complex. The effector ribonucleoprotein complex can recognize a target sequence through sequence specific DNA-RNA base pairing with a spacer sequence in the RNA guide. In some cases, target recognition activates one or more nuclease domains (e.g., a RuvC domain or HNH domain) in the CRISPR domain to make a double -stranded cut at the target DNA. A CRISPR-Cas domain complexed with an RNA guide can be capable of inactivating target gene through a gene knockout. In some cases, the CRISPR domain is used to enable gene insertion and/or deletion, which can inactivate, modify, or restore the gene’s function.
[0141] One or more components of a CRISPR/Cas system (e.g., modified and/or unmodified) delivered by the lipid delivery particles disclosed herein can be utilized as a genome engineering tool in a wide variety of organisms including diverse mammals, animals, plants, and yeast. A CRISPR/Cas system can comprise a guide nucleic acid such as a guide RNA (gRNA) complexed with a Cas protein for targeted regulation of gene expression and/or activity or nucleic acid editing. An RNA-guided Cas protein (e.g., a Cas nuclease such as a Cas9 nuclease) can specifically bind a target polynucleotide (e.g., DNA) in a sequence-dependent manner. The Cas protein, if possessing nuclease activity, can cleave the DNA (Gasiunas, G., etal, “Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria,” Proc Natl Acad Sci USA (2012) 109: E2579-E2 86; Jinek, M., et al, “A programmable dual -RNA-guided DNA endonuclease in adaptive bacterial immunity,” Science (2012) 337:816-821; Sternberg, S. H., et al, “DNA interrogation by the CRISPR RNA-guided endonuclease Cas9,” Nature (2014) 507:62; Deltcheva, E., et al, “CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III,” Nature (201 1) 471 : 602-607), and has been widely used for programmable genome editing in a variety of organisms and model systems (Cong, L., et al, “Multiplex genome engineering using CRISPR Cas systems,” Science (2013) 339:819-823; Jiang, W., et al, “RNA- guided editing of bacterial genomes using CRISPR-Cas systems,” Nat. Biotechnol. (2013) 31: 233-239; Sander, J. D. & Joung, J. K, “CRISPR-Cas systems for editing, regulating and targeting genomes,” Nature Biotechnol. (2014) 32:347-355).
[0142] In some cases, the Cas protein is mutated and/or modified to yield a nuclease deficient protein or a protein with decreased nuclease activity relative to a wild-type Cas protein. A nuclease deficient protein can retain the ability to bind DNA but can lack or have reduced nucleic acid cleavage activity. A protein encoded by a donor sequence comprises a Cas nuclease (e.g., retaining wild-type nuclease activity, having reduced nuclease activity, and/or lacking nuclease activity) can function in a CRISPR/Cas system to regulate the level and/or activity of a target gene or protein (e.g., decrease, increase, or elimination). The Cas protein can bind to a target polynucleotide and prevent transcription by physical obstruction or edit a nucleic acid sequence to yield non -functional gene products. In some cases, the Cas protein cleaves both strands of DNA. In some cases, the Cas protein cleaves one strand of DNA.
[0143] In some embodiments, the nuclease is a Cas protein that forms a complex with a guide nucleic acid, such as a guide RNA (gRNA). In some embodiments, the donor sequence disclosed herein encodes a Cas protein that forms a complex with a single guide nucleic acid, such as a single guide RNA (sgRNA). In some embodiments, the donor sequence disclosed herein encodes a Cas protein that forms a complex with two separate RNA molecules of a dual guide nucleic acid (dgRNA). In some embodiments, the donor sequence in the lipid delivery particles disclosed herein comprises or encodes an RNA-binding protein (RBP) optionally complexed with a guide nucleic acid, such as a guide RNA (e.g. , sgRNA, dgRNA), which is able to form a complex with a Cas protein. In some embodiments, the gRNA comprises a scaffolding sequences that tethers the gRNA to the Cas protein. In some embodiments, the gRNA comprises a scaffolding sequence and a spacer sequence that directs the Cas protein to a specific locus. In some embodiments, the scaffolding sequence is configured to bind to the positively charged groves in the Cas9 protein. In some embodiments, the scaffolding sequence is configured to bind to the Cas protein in the payload. In some cases, Cas undergoes a conformational change when the gRNA binds to the target locus. In some cases, the conformational change in Cas shifts the molecule from an inactive, non-DNA binding conformation into an active DNA-binding conformation. In some cases, the Cas protein undergoes a confirmational change if the spacer sequence has sufficient homology to the sequence at the target locus. In some embodiments, gRNAs can be modified. Exemplary modifications to the gRNA are provided in United States Patent Number 11,479,767 B2, United States Patent Application Publication Number US2020/0339980 Al, and United States Patent Application Publication Number US2021/0079389 Al, each of which is incorporated herein by reference in its entirety.
[0144] One or more components of any suitable CRISPR/Cas system can be delivered by the lipid delivery particle described in the present disclosure. A CRISPR/Cas system can be referred to using a variety of naming systems. Exemplary naming systems are provided in Makarova, K.S. et al, “An updated evolutionary classification of CRISPR-Cas systems,” Nat Rev Microbiol (2015) 13:722-736 and Shmakov, S. et al, “Discovery and Functional Characterization of Diverse Class 2 CRISPR-Cas Systems,” Mol Cell (2015) 60: 1-13. A CRISPR/Cas system can be a type I, a type II, a type III, a type IV, a type V, a type VI system, or any other suitable CRISPR/Cas system. A CRISPR/Cas system as used herein can be a Class 1, Class 2, or any other suitably classified CRISPR/Cas system. Class 1 or Class 2 determination can be based upon the genes encoding the effector module. Class 1 systems generally have
a multi-subunit crRNA-effector complex, whereas Class 2 systems generally have a single protein, such as Cas9, Cpfl, C2cl, C2c2, C2c3, or a crRNA-effector complex. A Class 1 CRISPR/Cas system can use a complex of multiple Cas proteins to effect regulation. A Class 1 CRISPR/Cas system can comprise, for example, type I (e.g, I, IA, IB, IC, ID, IE, IF, IU), type III (e.g., Ill, IIIA, IIIB, IIIC, IIID), and type IV (e.g., IV, IVA, IVB) CRISPR/Cas type. A Class 2 CRISPR/Cas system can use a single large Cas protein to effect regulation. A Class 2 CRISPR/Cas systems can comprise, for example, type II (e.g., II, IIA, IIB) and type V CRISPR/Cas type. CRISPR systems can be complementary to each other, and/or can lend functional units in trans to facilitate CRISPR locus targeting. Examples of Cas proteins that can be used as part of the CRISPR systems described herein include c2cl, Casl3a (formerly C2c2), Casl3b, Casl3c, Casl3d, c2c3, Casl, CaslB, Cas2, Cas3, Cas4, Cas5, Cas5e (CasD), Cas6, Cas6e, Cas6f, Cas7, Cas8a, Cas8al, Cas8a2, Cas8b, Cas8c, Cas9 (Csnl or Csxl2), CaslO, CaslOd, Casl4, CaslO, CaslOd, CasF, CasG, CasH, Casl2a (formerly Cpfl), Csyl, Csy2, Csy3, Csel (CasA), Cse2 (CasB), Cse3 (CasE), Cse4 (CasC), Cscl, Csc2, Csa5, Csn2, Csm2, Csm3, Csm4, Csm5, Csm6, Cmrl, Cmr3, Cmr4, Cmr5, Cmr6, Csbl, Csb2, Csb3, Csxl7, Csxl4, CsxlO, Csxl6, CasX, Csx3, Csxl, Csxl5, Csfl, Csf2, Csf3, Csf4, and Cul966, and homologs or modified versions thereof. Examples of mutant Cas9 proteins or Cas9 variants include SpG, SpCas9-NG. Cas9-NRNH, SpG, SpRY, Cas9-VQR, Cas9-EQR, SaCas9-KKH, Nme2Cas9, eNme2-C, eNme2-C.NR, eNme2-T.l, eNme2-T.2, SpRY, eSpCas9(l.l), SpCas9-HFl, nSpCas9, eSpCas9, Sniper-Cas9, HypaCas9, evoCas9, Cas9TX, HscCas9-vl.2, superFi-Cas9, efSaCas9, SaCas9- HF, Cas9-HF, Cas mini, SaCas9, SpCas9(H840A), dSpCas9, SpCas9(N863A), SpCas9(D839A), SpCas9(H983A), as well as others described in Chuang CK et al., Int J Mol Sci. 2021 Sep 13;22(18):9872, and Li, T. et al.. Sig Transduct Target Ther 8, 36 (2023), each of which is incorporated herein by reference in its entirety.
[0145] A CRISPR system can comprise single subunit or multi-subunit effectors. In some cases, a CRISPR system is a Class 1 CRISPR system. A Class 1 CRISPR system can be a type I, type III, or a type IV system. A Class 1 type I CRISPR system can comprise a multi-subunit effector. In some cases, a Class 1 type I CRISPR system comprises a protein or domain in the Cascade-Cas3 protein complex. A Class 1 type I CRISPR system can comprise a Cas6, Cas7, Cas5, Casl 1, Cas8, or Cas3 domain. A Class 1 type III CRISPR system can comprise a multi-subunit effector. In some cases, a Class 1 type III CRISPR system comprises a Csm complex or a Cmr complex. In some cases, a Class 1 type III CRISPR system comprises a Cas6, a Cas7 (Csm3 or Cmr4), a Cas7-related (Csm5, Cmrl, or Cmr6), a Cas5 (e.g., Csm4 or Cmr5), a Casl 1 (e.g., Csm2 or Cmr3), or a CaslO (e.g., Csml or Cmr2) domain. A Class 1 type IV CRISPR system can comprise a Cas6, a Cas7, a Cas5, a Casl 1, a Cas8 (e.g., Csfl), or a DinG or CysH domain. In some cases, a CRISPR system comprises Cmrl, Cmr3, Cmr4, Cmr5, or Cmr6. In some cases, a CRISPR system comprises Csbl, Csb2, or Csb3. A CRISPR system can comprise Csfl, Csf2, Csf3, or Csf4. A CRISPR system can comprise Csn2, Csm2, Csm3, Csm4, Csm5, or Csm6. A CRISPR system can comprise Cscl or Csc2. A CRISPR system can comprise Casl, CaslB, Cas2, or Cas4. A CRISPR system can comprise Csyl, Csy2, or Csy3. A CRISPR system can comprise Csel or Cse2. A CRISPR system can comprise Csn2. A CRISPR system can comprise CsaX, Csxl, Csx3, CsxlO, Csxl 4, Csxl 5, Csxl 6, or
Csxl7. In some cases, a CRISPR system comprises a modified version of any one of the foregoing Cas proteins. In some cases, a modified version of the foregoing Cas protein comprises a nickase mutation. In some cases, the nickase mutation corresponds to the D10A mutation of the wild type Cas9 protein. In some cases, the nickase mutation corresponds to the H840A mutation of the wild type Cas9 protein. In some cases, the nickase mutation occurs in the RuvC domain of the wild type Cas9 protein. In some cases, the nickase mutation occurs in the HNH domain of the wild type Cas9 protein. In some cases the RuvC domain can be mutated to prevent cleavage of the non-target DNA strand. In some cases the HNH domain can be mutated to prevent cleavage of the target DNA strand. In some cases, a modified version of the foregoing Cas protein comprises one or more mutations that disrupt cleavage activity. In some cases, a Cas protein with disrupted cleavage activity is catalytically inactive or catalytically dead. In some cases, the catalytically dead mutations occur in the RuvC domain and the HNH domain of the wild type Cas9 protein. In some cases, the catalytically inactive mutations correspond to the D10A mutation and the H840A mutation of the wild type Cas9 protein.
[0146] In some cases, a CRISPR system is a Class 2 CRISPR system. A Class 2 CRISPR system can be a Class 2 type II CRISPR system, a Class 2 type V CRISPR system, or a Class 2 type VI CRISPR system. A Class 2 type II CRISPR system can comprise a Cas9 domain (also known as Csnl and Csxl2). A Cas9 domain can be a SpyCas9, a GeoCas9, a SauCas9, a KhuCas9, a AinCas9, an FmaCas9, a SgaCas9, a ScCas9, a SauriCas9 domain. A Cas9 domain can be a hyperactive Cas9 domain. A Class 2 type V CRISPR system can comprise a Casl2 domain. A Casl2 domain can be a Casl2a, a Casl2b, a Casl2bl, a Casl2c, a Casl2d, a Casl2e, a Casl2f, a Casl2g, a Casl2h, a Casl2i, a Casl2j, a Casl2k, a Casl21, or a Cas 12m domain. A Class 2 type VI CRISPR system can comprise a Cas 13 domain.
[0147] In some cases, a CRISPR system comprises a circularly permuted Cas9.
[0148] In some cases, a CRISPR system comprises CjCas9, Casl3a, Casl3b, Casl3c, or Casl3d. In some cases, a CRISPR system comprises Cas 14, xCas9, or SpCas9-NG.
[0149] In some cases, a CRISPR-Cas domain comprises one or more subdomains. For example, a Cas9 domain can comprise a Reel, a Rec2, a Rec3, a RuvC, an HNH, or a Wedge/PAM-interacting domain. A Casl2 domain can comprise a Reel, Rec2, a crRNA oligonucleotide binding domain (OBD), a Nuc domain, a PAM-interacting (PI) domain, or a RuvC domain. In some cases, the RuvC domain comprises nuclease activity. In some cases, the HNH domain comprises nuclease activity. The PAM-interacting domain can bind to a protospacer adjacent motif (PAM) sequence that is next to a target sequence in a target nucleic acid molecule. PAM recognition can help activate a nuclease domain to make a cut at the target sequence. In some cases, a CRISPR protein or domain is an engineered or mutated variant of a protein involved in a CRISPR system. An engineered or mutated CRISPR domain can comprise a truncation, a deletion of a part of one or more domains or subdomains, or a mutation of an active site (e.g., a RuvC active site or HNH active site). In some cases, a CRISPR domain with a mutation of one or more active sites is catalytically inactive (e.g., dCas9). In some cases, a CRISPR domain with one or more mutated active sites comprises less than 90%, less than 80%, less than 70%, less than 60%, less than 50%, less than 40%, less than 30%, less than 20%, less than 10%, less than 5%, or less than 1% of the
nuclease activity of its wildtype counterpart. For example, a dCas9 can result from the point mutations D10A in the RuvC domain and the point mutation H840A in the HNH domain. In other cases, a mutation can result in a CRISPR nickase. A nickase can generate nick or a single-stranded cut. A nickase can generate a nick in the strand complementary to the RNA guide (e.g., the targeting strand) or in the strand on the non-targeting strand. For example, a RuvC mutation D10A in a Cas9 domain can produce a Cas9 nickase domain that nicks the targeting strand. An HNH mutation H840A in a Cas9 domain can produce a Cas9 nickase domain that nicks the non-targeting strand.
[0150] A Cas protein can comprise one or more domains. Examples of domains include, guide nucleic acid recognition and/or binding domain, nuclease domains (e.g, DNase or RNase domains, RuvC, HNH), DNA binding domain, RNA binding domain, helicase domains, protein-protein interaction domains, and dimerization domains. A guide nucleic acid recognition and/or binding domain can interact with a guide nucleic acid. A nuclease domain can comprise catalytic activity for nucleic acid cleavage. A nuclease domain can lack catalytic activity to prevent nucleic acid cleavage. A Cas protein can be a chimeric Cas protein that is fused to other proteins or polypeptides. A Cas protein can be a chimera of various Cas proteins, for example, comprising domains from different Cas proteins.
[0151] In some cases, a CRISPR system comprises an Argonaute (Ago) domain.
[0152] Another example of a Cas protein that can be used as part of the prime editor includes Cas 14. A Cas 14 protein or polypeptide (also termed as “CasZ” protein or polypeptide) can bind and/or modify (e.g., cleave, nick, methylate, demethylate, etc.) a target nucleic acid and/or a polypeptide associated with target nucleic acid (e.g. , methylation or acetylation of a histone tail) (e.g. , in some cases the CasZ protein includes a chimeric partner with an activity, and in some cases the CasZ protein provides nuclease activity). In some cases, the Casl4 protein or polypeptide is a naturally occurring protein (e.g., naturally occurs in prokaryotic cells) (e.g., a CasZ protein). In other cases, the Cas 14 protein or polypeptide not a naturally occurring polypeptide (e.g, the Cas 14 protein is a variant Cas 14 protein, a chimeric protein, and the like). A Cas 14 protein includes 3 partial RuvC domains (RuvC-I, RuvC-II, and RuvC-III, also referred to herein as subdomains) that are not contiguous with respect to the primary amino acid sequence of the Cas 14 protein but form a RuvC domain once the protein is produced and folds. A naturally occurring Cas 14 protein functions as an endonuclease that catalyzes cleavage at a specific sequence in a targeted nucleic acid (e.g., a double stranded DNA (dsDNA)). The sequence specificity is provided by the associated guide RNA, which hybridizes to a target sequence within the target DNA. The naturally occurring Cas 14 guide RNA is a crRNA, where the crRNA includes (i) a guide sequence that hybridizes to a target sequence in the target DNA and (ii) a protein binding segment that binds to the Cas 14 protein. Examples of Casl4 proteins include those described U.S. Patent Publication Nos. US20200172886 and US20210214697, Harrington LB et al., Science. 2018 Nov 16;362(6416):839-842; Aquino-Jarquin G. Nanomedicine. 2019 Jun;18:428-431; each of which is incorporated herein by reference in its entirety. In some cases, the donor sequence disclosed herein encodes Casl4 polypeptide or a nucleic acid molecule encoding Cas 14 polypeptide. In some cases, the donor sequence disclosed herein encodes Cas 14a
polypeptide. In some cases, the donor sequence disclosed herein encodes Casl4b polypeptide. In some cases, the donor sequence disclosed herein encodes Casl4c polypeptide.
[0153] A Cas protein can be from any suitable organism. Examples include Streptococcus pyogenes, Streptococcus thermophilus, Streptococcus sp., Staphylococcus aureus, Nocardiopsis dassonvillei, Streptomyces pristinae spiralis, Streptomyces viridochromo genes, Streptomyces viridochromogenes, Streptosporangium roseum, Streptosporangium roseum, AlicyclobacHlus acidocaldarius, Bacillus pseudomycoides, Bacillus selenitireducens, Exiguobacterium sihiricum, Lactobacillus delbrueckii, Lactobacillus salivarius, Microscilla marina, Burkholderiales bacterium, Polaromonas naphthalenivorans, Polaromonas sp., Crocosphaera watsonii, Cyanothece sp., Microcystis aeruginosa, Pseudomonas aeruginosa, Synechococcus sp., Acetohalobium arabaticum, Ammonifex degensii, Caldicelulosiruptor becscii, Candidatus Desulforudis, Clostridium botulinum, Clostridium difficile, Finegoldia magna, Natranaerobius thermophilus, Pelotomaculum thermopropionicum, Acidithiobacillus caldus, Acidithiobacillus ferrooxidans, Allochromatium vinosum, Marinobacter sp., Nitrosococcus halophilus, Nitrosococcus watsoni, Pseudoalteromonas haloplanktis, Ktedonobacter racemifer, Methanohalobium evestigatum, Anabaena variabilis, Nodularia spumigena, Nostoc sp., Arthrospira maxima, Arthrospira platensis, Arthrospira sp., Lyngbya sp., Microcoleus chthonoplastes, Oscillatoria sp., Petrotoga mobilis, Thermosipho africanus, Acaryochloris marina, Leptotrichia shahii, Leptotrichia wadeii, Leptotrichia wadeii F0279, Rhodobacter capsulatus SB1003, Rhodobacter capsulatus R121, Rhodobacter capsulatus DE442, Lachnospiraceae bacterium NK4A179, Lachnospiraceae bacterium MA2020, Clostridium aminophilum DSM 10710, Paludibacter propionicigenes WB4, Carnobacterium gallinarum DMS4847, Carnobacterium gallinarum DSM4847, and Francisella novicida. In some aspects, the organism is Streptococcus pyogenes (S. pyogenes). In some aspects, the organism is Staphylococcus aureus (S. aureus). In some aspects, the organism is Streptococcus thermophilus (.S'. thermophilus).
[0154] A Cas protein can be derived from a variety of bacterial species including Veillonella atypical, Fusobacterium nucleatum, Filifactor alocis, Solobacterium moorei, Coprococcus catus, Treponema denticola, Peptoniphilus duerdenii, Catenibacterium mitsuokai, Streptococcus mutans, Listeria innocua, Listeria seeligeri. Listeria weihenstephanensis FSL R90317, Listeria weihenstephanensis FSL M60635, Staphylococcus pseudintermedius, Acidaminococcus intestine, Olsenella uli, Oenococcus kitaharae, Bifidobacterium bifidum, Lactobacillus rhamnosus, Lactobacillus gasseri, Finegoldia magna, Mycoplasma mobile, Mycoplasma gallisepticum, Mycoplasma ovipneumoniae, Mycoplasma canis, Mycoplasma synoviae, Eubacterium rectale, Streptococcus thermophilus, Eubacterium dolichum, Lactobacillus coryniformis subsp. Torquens, Ilyobacter polytropus, Ruminococcus albus, Akkermansia muciniphila, Acidothermus cellulolyticus, Bifidobacterium longum, Bifidobacterium dentium, Corynebacterium diphtheria, Elusimicrobium minutum, Nitratifractor salsuginis, Sphaerochaeta globus, Fibrobacter succinogenes subsp. Succinogenes, Bacteroides fragilis, Capnocytophaga ochracea, Rhodopseudomonas palustris, Prevotella micans, Prevotella ruminicola, Flavobacterium columnare, Aminomonas paucivorans, Rhodospirillum rubrum, Candidatus Puniceispirillum marinum,
Verminephrobacter eiseniae, Ralstonia syzygii, Dinoroseobacter shibae, Azospirillum, Nitrobacter hambur gensis, Bradyrhizobium, Wolinella succinogenes, Campylobacter jejuni subsp. Jejuni, Helicobacter mustelae, Bacillus cereus, Acidovorax ebreus, Clostridium perfringens, Parvibaculum lavamentivorans, Roseburia intestinalis, Neisseria meningitidis, Pasteurella multocida subsp. Multocida, Sutterella wadsworthensis, proteobacterium, Legionella pneumophila, Parasutterella excrementihominis, Wolinella succinogenes, and Francisella novicida.
[0155] A Cas protein as disclosed herein can be a wildtype or a modified form of a Cas protein. A Cas protein can be an active variant, inactive variant, or fragment of a wild type or modified Cas protein. A Cas protein can comprise an amino acid change such as a deletion, insertion, substitution, variant, mutation, fusion, chimera, or any combination thereof relative to a wild-type version of the Cas protein. A Cas protein can be a polypeptide with at least about 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity or sequence similarity to a wild type exemplary Cas protein. A Cas protein can be a polypeptide with at most about 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100% sequence identity and/or sequence similarity to a wild type exemplary Cas protein. Variants or fragments can comprise at least about 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity or sequence similarity to a wild type or modified Cas protein or a portion thereof. Variants or fragments can be targeted to a nucleic acid locus in complex with a guide nucleic acid while lacking nucleic acid cleavage activity.
[0156] A Cas protein can comprise one or more nuclease domains, such as DNase domains. For example, a Cas9 protein can comprise a RuvC-like nuclease domain and/or an HNH-like nuclease domain. The RuvC and HNH domains can each cut a different strand of double-stranded DNA to make a double -stranded break in the DNA. A Cas protein can comprise only one nuclease domain (e.g., Cpfl comprises RuvC domain but lacks HNH domain).
[0157] A Cas protein can comprise an amino acid sequence having at least about 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% sequence identity or sequence similarity to a nuclease domain (e.g., RuvC domain, HNH domain) of a wild-type Cas protein.
[0158] A Cas protein can be modified to optimize regulation of gene expression. A Cas protein can be modified to increase or decrease nucleic acid binding affinity, nucleic acid binding specificity, and/or enzymatic activity. Cas proteins can also be modified to change any other activity or property of the protein, such as stability. For example, one or more nuclease domains of the Cas protein can be modified, deleted, or inactivated, or a Cas protein can be truncated to remove domains that are not essential for the function of the protein or to optimize (e.g., enhance or reduce) the activity of the Cas protein for regulating gene expression.
[0159] In some embodiments, the prime editor delivered by the lipid delivery particles of the present disclosure contain a nuclease-null DNA binding protein derived from a DNA nuclease that can induce transcriptional activation or repression of a target DNA sequence. In some embodiments, the donor
sequence encodes a nuclease-null RNA binding protein derived from an RNA nuclease that can induce transcriptional activation or repression of a target RNA sequence. For example, a doner sequence can encode a Cas protein which lacks cleavage activity.
[0160] A Cas protein can be a chimeric protein. For example, a Cas protein can be fused to a heterologous functional domain. A heterologous functional domain can comprise a cleavage domain, an epigenetic modification domain, a transcriptional activation domain, or a transcriptional repressor domain. A Cas protein can also be fused to a heterologous polypeptide providing increased or decreased stability. The fused domain or heterologous polypeptide can be located at the N-terminus, the C-terminus, or internally within the Cas protein.
[0161] The regulation of genes can be of any gene of interest. It is contemplated that genetic homologues of a gene described herein are covered. For example, a gene can exhibit a certain identity and/or homology to genes disclosed herein. Therefore, it is contemplated that a gene that exhibits or exhibits about 50%, 55%, 60%, 65%, 70%, 75%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% homology (at the nucleic acid or protein level) can be modified. It is also contemplated that a gene that exhibits or exhibits about 50%, 55%, 60%, 65%, 70%, 75%, 80%, 81%, 82%, 83%, 84%, 85%, 86%, 87%, 88%, 89%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, 99%, or 100% identity (at the nucleic acid or protein level) can be modified.
[0162] A Cas protein can be provided in any form. For example, a Cas protein can be provided in the form of a protein, such as a Cas protein alone or complexed with a guide nucleic acid. A Cas protein can be provided in the form of a nucleic acid encoding the Cas protein, such as an RNA (e.g., messenger RNA (mRNA)) or DNA.
[0163] The nucleic acid encoding the Cas protein that is part of the prime editor can be codon optimized for efficient translation into protein in a particular cell or organism.
[0164] In some embodiments, a Cas protein is a dead Cas protein. A dead Cas protein can be a protein that lacks nucleic acid cleavage activity.
[0165] A Cas protein can comprise a modified form of a wild type Cas protein. The modified form of the wild type Cas protein can comprise an amino acid change (e.g., deletion, insertion, or substitution) that reduces the nucleic acid-cleaving activity of the Cas protein. For example, the modified form of the Cas protein can have less than 90%, less than 80%, less than 70%, less than 60%, less than 50%, less than 40%, less than 30%, less than 20%, less than 10%, less than 5%, or less than 1% of the nucleic acidcleaving activity of the wild-type Cas protein (e.g., Cas9 from S. pyogenes). The modified form of Cas protein can have no substantial nucleic acid-cleaving activity. When a Cas protein is a modified form that has no substantial nucleic acid-cleaving activity, it can be referred to as enzymatically inactive and/or “dead” (abbreviated by “d”). A dead Cas protein (e.g, dCas, dCas9) can bind to a target polynucleotide but may not cleave the target polynucleotide. In some aspects, a dead Cas protein is a dead Cas9 protein.
[0166] A dCas9 polypeptide can associate with a guide nucleic acid molecule (e.g., PEgRNA) to activate or repress transcription of target DNA. Guide nucleic acid molecules can be introduced into cells expressing the engineered chimeric receptor polypeptide. In some cases, such cells contain one or more
different guide nucleic acid molecules that target the same nucleic acid. In other cases, the guide nucleic acid molecules target different nucleic acids in the cell. The nucleic acids targeted by the guide nucleic acid molecule can be any that are expressed in a cell such as an immune cell. The nucleic acids targeted can be a gene involved in immune cell regulation. In some embodiments, the nucleic acid is associated with cancer. The nucleic acid associated with cancer can be a cell cycle gene, cell response gene, apoptosis gene, or phagocytosis gene. The recombinant guide nucleic acid molecule can be recognized by a CRISPR protein, a nuclease-null CRISPR protein, variants thereof, derivatives thereof, or fragments thereof.
[0167] Enzymatically inactive can refer to a polypeptide that can bind to a nucleic acid sequence in a polynucleotide in a sequence-specific manner, but may not cleave a target polynucleotide. An enzymatically inactive site-directed polypeptide can comprise an enzymatically inactive domain (e.g., nuclease domain). Enzymatically inactive can refer to no activity. Enzymatically inactive can refer to substantially no activity. Enzymatically inactive can refer to essentially no activity. Enzymatically inactive can refer to an activity less than 1%, less than 2%, less than 3%, less than 4%, less than 5%, less than 6%, less than 7%, less than 8%, less than 9%, or less than 10% activity compared to a wild-type exemplary activity (e.g., nucleic acid cleaving activity, wild-type Cas9 activity).
[0168] One or a plurality of the nuclease domains (e.g. , RuvC, HNH) of a Cas protein can be deleted or mutated so that they are no longer functional or comprise reduced nuclease activity (e.g., deactivated or dead Cas, i.e., “dCas”). For example, in a Cas protein comprising at least two nuclease domains (e.g. , Cas9), if one of the nuclease domains is deleted or mutated, the resulting Cas protein, known as a nickase, can generate a single-strand break at a CRISPR RNA (crRNA) recognition sequence within a doublestranded DNA but not a double-strand break. Such a nickase can cleave the complementary strand or the non-complementary strand, but may not cleave both. If all of the nuclease domains of a Cas protein (e.g., both RuvC and HNH nuclease domains in a Cas9 protein; RuvC nuclease domain in a Cpfl protein) are deleted or mutated, the resulting Cas protein can have a reduced or no ability to cleave both strands of a double -stranded DNA. An example of a mutation that can convert a Cas9 protein into a nickase is a D10A (aspartate to alanine at position 10 of Cas9) mutation in the RuvC domain of Cas9 from S. pyogenes. H939A (histidine to alanine at amino acid position 839) or H840A (histidine to alanine at amino acid position 840) in the HNH domain of Cas9 from S. pyogenes can convert the Cas9 into a nickase. An example of a mutation that can convert a Cas9 protein into a dead Cas9 is a D10A (aspartate to alanine at position 10 of Cas9) mutation in the RuvC domain and H939A (histidine to alanine at amino acid position 839) or H840A (histidine to alanine at amino acid position 840) in the HNH domain of Cas9 from S. pyogenes.
[0169] A dead Cas protein can comprise one or more mutations relative to a wild-type version of the protein. The mutation can result in less than 90%, less than 80%, less than 70%, less than 60%, less than 50%, less than 40%, less than 30%, less than 20%, less than 10%, less than 5%, or less than 1% of the nucleic acid-cleaving activity in one or more of the plurality of nucleic acid-cleaving domains of the wildtype Cas protein. The mutation can result in one or more of the plurality of nucleic acid-cleaving domains
retaining the ability to cleave the complementary strand of the target nucleic acid but reducing its ability to cleave the non-complementary strand of the target nucleic acid. The mutation can result in one or more of the plurality of nucleic acid-cleaving domains retaining the ability to cleave the non-complementary strand of the target nucleic acid but reducing its ability to cleave the complementary strand of the target nucleic acid. The mutation can result in one or more of the plurality of nucleic acid-cleaving domains lacking the ability to cleave the complementary strand and the non-complementary strand of the target nucleic acid. The residues to be mutated in a nuclease domain can correspond to one or more catalytic residues of the nuclease. For example, residues in the wild type exemplary .S', pyogenes Cas9 polypeptide such as Asp 10, His840, Asn854 and Asn856 can be mutated to inactivate one or more of the plurality of nucleic acid-cleaving domains (e.g., nuclease domains). The residues to be mutated in a nuclease domain of a Cas protein can correspond to residues AsplO, His840, Asn854 and Asn856 in the wild type .S'. pyogenes Cas9 polypeptide, for example, as determined by sequence and/or structural alignment.
[0170] As examples, residues DIO, G12, G17, E762, H840, N854, N863, H982, H983, A984, D986, and/or A987 (or the corresponding mutations of any of the Cas proteins) can be mutated. For example, e.g., D10A, G12A, G17A, E762A, H840A, N854A, N863A, H982A, H983A, A984A, and/or D986A. Mutations other than alanine substitutions can be suitable.
[0171] A D10A mutation can be combined with one or more of H840A, N854A, or N856A mutations to produce a Cas9 protein substantially lacking DNA cleavage activity (e.g., a dead Cas9 protein). A H840A mutation can be combined with one or more of D10A, N854A, or N856A mutations to produce a site- directed polypeptide substantially lacking DNA cleavage activity. A N854A mutation can be combined with one or more of H840A, D10A, or N856A mutations to produce a site-directed polypeptide substantially lacking DNA cleavage activity. A N856A mutation can be combined with one or more of H840A, N854A, or D10A mutations to produce a site-directed polypeptide substantially lacking DNA cleavage activity.
[0172] In some cases, a dCas9 can be fused to other proteins. In some cases, dCas9 can be fused to SunTag, KRAB, VPS4, P3000, VPR, VP64, V64-p65-Rta, VP160, VP192, HDAC1, DNMT3A, TET1, SPH, KRAB-MeCP2, epigenetic regulators, or other proteins. In some cases, a dCas9 fusion comprises a ZIM3 KRAB-Cas9 fusion. In some cases, a Cas9 fusion can be a paired dCas9 system. In some cases, the dCas9 can be part of a SAM system or REDMAP system. Examples of Cas9 variants and fusion proteins can be found in Li, T. et al,, Sig Transduct Target Ther 8, 36 (2023), which is incorporated in its entirety, [0173] In some embodiments, a Cas protein is a Class 2 Cas protein. In some embodiments, a Cas protein is a type II Cas protein. In some embodiments, the Cas protein is a Cas9 protein, a modified version of a Cas9 protein, or derived from a Cas9 protein. For example, a Cas9 protein lacking cleavage activity. In some embodiments, the Cas9 protein is a Cas9 protein from .S', pyogenes (e.g., SwissProt accession number Q99ZW2). In some embodiments, the Cas9 protein is a Cas9 from S.aureus (e.g., SwissProt accession number J7RUA5). In some embodiments, the Cas9 protein is a modified version of a Cas9 protein from .S', pyogenes or .S'. Aureus. In some embodiments, the Cas9 protein is derived from a
Cas9 protein from .S'. pyogenes or .S'. Aureus. For example, a .S' pyogenes or .S' Aureus Cas9 protein lacking cleavage activity.
[0174] Cas9 can generally refer to a polypeptide with at least about 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100% sequence identity and/or sequence similarity to a wild type exemplary Cas9 polypeptide (e.g. , Cas9 from .S' pyogenes). Cas9 can refer to a polypeptide with at most about 5%, 10%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, 100% sequence identity and/or sequence similarity to a wild type exemplary Cas9 polypeptide (e.g. , from .S' pyogenes). Cas9 can refer to the wildtype or a modified form of the Cas9 protein that can comprise an amino acid change such as a deletion, insertion, substitution, variant, mutation, fusion, chimera, or any combination thereof.
[0175] In some embodiments, a guidable polypeptide domain is a Cas9 or variant thereof. In some embodiments, the Cas9 or variant thereof is a nuclease active Cas9 domain, a nuclease inactive Cas9 domain, or a Cas9 nickase domain or a variant thereof. In some embodiments, a guidable polypeptide domain is Cas9, Casl2e, Casl2d, Casl2a, Casl2bl, Casl3a, Casl2c, or Argonaute (Ago domain), any of which optionally has a nickase activity. In some embodiments, a guidable polypeptide domain comprises an amino acid sequence at least 80%, 85%, 90%, 95%, or 99% identical to any one of sequences listed in Table 8 below. In some embodiments, a guidable polypeptide domain comprises an amino acid sequence at least 80%, 85%, 90%, 95%, or 99% identical to any one of sequences set forth in SEQ ID NOs: 1000- 1020. In some cases, a guidable polypeptide domain is a Cas9 H840A nickase. In some cases, a guidable polypeptide domain is Cas9 D10A nickase. Cas9-H840A. In some cases, a guidable polypeptide domain is a Casl2a/b nickase.
Polymerase
[0176] In some cases, the payload in the present disclosure comprises a polymerase (e.g., reverse transcriptase). A polymerase can comprise a natural or an engineered domain. A polymerase can be capable of synthesizing nucleic acids. A polymerase can be a DNA polymerase or an RNA polymerase. In some cases, a polymerase is a reverse transcriptase. A reverse transcriptase can synthesize DNA from deoxyribonucleotides. In some cases, a reverse transcriptase adds deoxyribonucleotides to the 3’ end of a nucleic acid primer to synthesize DNA. In some cases, a reverse transcriptase uses an RNA template and uses base-pairing interactions to synthesize a DNA strand that is complementary to the RNA template. The reverse transcriptase domain can be a reverse transcriptase from any organism, phage, virus, or an engineered or mutated variant. The reverse transcriptase domain can be a reverse transcriptase derived from or sharing structural or sequencing similarity to a reverse transcriptase in a CRISPR system. The reverse transcriptase can be an M-MLV or HIV reverse transcriptase. The reverse transcriptase can be a human LINE-1 reverse transcriptase or a group II intron reverse transcriptase. The reverse transcriptase can be a human endogenous retrovirus reverse transcriptase.
Transposase domain
[0177] In some cases, a nucleic-acid modifying effector or a nucleic acid-binding moiety comprises a transposase domain. A transposase domain can be a natural or an engineered domain. A transposase domain can be capable of aiding the translocation of a transposable element, a nucleic acid sequence that can change its position within a genome. In some cases, a transposase domain comprises a TnsA, a TnsB, a TnsC, or a TnsD domain. In some cases, a transposase domain comprises a TniQ domain. In some cases, a transposase domain is derived from or shares sequence or structural similarity with a transposase in a CRISPR system (e.g., a CRISPR-associated transposase). In some cases, a transposase domain is derived from or share sequence or structural similarity with a transposase domain from a type I CRISPR- associated transposon (CAST) system. In some cases, transposase domain is derived from or share sequence or structural similarity with a transposase domain from a type V CRISPR-associated transposon (CAST) system. A transposase domain can be capable of binding to a guidable polypeptide domain. In
some cases, a transposase domain is coupled to a guidable polypeptide domain. In some cases, a transposase domain is capable of binding to a type I CRISPR-Cas domain (e.g., a Cascade domain, a Cas8 domain, or a Cas5 domain). In some cases, a transposase domain is capable of binding to a type V CRISPR-Cas domain (e.g., a Casl2 domain). In some cases, a transposase domain is capable of mediating targeted insertion of a nucleic acid into a target nucleic acid. In some cases, a transposase domain is capable of mediating targeted insertion of a nucleic acid that is at least 5 kb, at least 6 kb, at least 7 kb, at least 8kb, at least 9kb, at least lOkb, at least 1 Ikb, at least 12kb, at least 13kb, at least 14kb, or at least 15 kb into a target nucleic acid.
Transcriptional regulatory domain
[0178] In some cases, the payload comprises a transcriptional regulatory domain. A transcriptional regulatory domain can be a natural or an engineered domain. A transcriptional regulatory domain can be capable of regulating, activating, or inhibiting gene expression. For example, a transcriptional repressor can silence gene expression by binding to the promoter of a gene. A transcriptional activator can bind to enhancers or regulatory elements to activate expression of a gene. A transcriptional regulatory domain can comprise a transcription factor. A transcriptional regulatory domain can comprise a transcriptional activation domain or a transcriptional repression domain. For example, a transcriptional activation domain can be or comprise a CAP domain, a VP64 domain, a p65 domain, an Rta domain, a synergistic activation mediator (SAM) domain, a SunTag domain, a VPR domain, a DNA demethylase domain, a histone methyltransferase domain, a histone acetyltransferase domain, or a histone demethylase domain. A transcriptional repression domain can be or comprise a dCas9 domain, a KRAB domain, a Sin3 interacting domain (SID), or a MePC2 domain, a DNA methyltransferase domain, a histone deacetylase domain, a histone methyltransferase domain, or a histone demethylase domain. In some cases, a transcriptional regulatory domain comprises an epigenetic modifying effector domain. For example, an epigenetic modifying effector can be a DNA methyltransferase, a DNA demethylase, a histone methyltransferase, a histone demethylase, a histone acetyltransferase, or a histone deacetylase domain. A DNA methyltransferase domain can be capable of methylating a nucleic acid. A DNA demethylase domain can be capable of demethylating a nucleic acid. A histone methyltransferase domain can be capable of methylating a histone. A histone demethylase domain can be capable of demethylating a histone. A histone acetyltransferase domain can be capable of adding an acetyl group to a histone. A histone deacetylase domain can be capable of removing an acetyl group from a histone.
Zinc finger domain
[0179] In some embodiments, the payload comprises a zinc finger domain. A zinc finger domain can be a natural or an engineered domain. A zinc finger domain can bind to a specific DNA sequence in a target nucleic acid. A zinc finger domain can comprise from 1 to 10, from 2 to 10, from 3 to 10, from 4 to 10, from 5 to 10, from 6 to 10, from 7 to 10, from 8 to 10, from 9 to 10 zinc fingers, from 1 to 8, from 2 to 8, from 3 to 8, from 4 to 8, from 5 to 8, from 6 to 8, from 7 to 8, from 8 to 8, from 9 to 8 zinc fingers. In some cases, a zinc finger domain comprises a two-handed zinc finger domain. A two handed zinc finger domain can comprise two clusters of zinc finger domains that are separated by intervening amino acids. A
two handed zinc finger domain can bind to two noncontiguous target DNA sequences. In some cases, the spacing between the two noncontiguous target sequences comprises from 1 to 15, from 1 to 12, from 1 to 10, from 1 to 8, or from 1 to 5 nucleotides. For example, a two handed type of zinc finger binding protein can be SIP1. A cluster of zinc finger domains in a two handed zinc finger domain can be capable of binding to a unique target nucleic acid sequence.
TALE domain
[0180] In some embodiments, the payload comprises a TALE domain. A TALE domain can be a natural or an engineered domain. A TALE domain can bind to a specific DNA sequence. A TALE domain can comprise one or more effector domains. A TALE effector domain can comprise a central repeat domain comprising tandem repeats. A tandem repeat can comprise repeat variable residues (RVD). One or more RVDs can detect a specific DNA base. Different TALE effector domains may have a different number of repeats and a different order of their repeats. The C-terminal repeat is usually shorter in length (e.g., about 20 amino acids). Sequential repeats and their RVDs can recognize sequential DNA bases.
[0181] A TALE domain described herein can be derived from a TALE effector from a bacterial species. The TALE domain can be engineered to target a given nucleic acid sequence based on their DNA base specificities. The TALE domain can be engineered to remove or add a TALE effector domain. In some cases, the TALE domain corresponds to a perfect match to a nucleic acid target sequence. In some cases, the TALE domain of an epigenetic effector corresponds to one or more mismatches to a target base in the target nucleic acid.
Fusion protein
[0182] In some cases, the payload in the present disclosure comprises a fusion protein. A fusion protein can comprise two or more polypeptide domains of any of the polypeptide domains described elsewhere herein. A fusion protein can be a natural or an engineered fusion protein. In some cases, the two or more polypeptide domains are coupled together. The two or more polypeptide domains can be coupled together directly or coupled together indirectly. For example, a first polypeptide domain can be coupled directly to a second polypeptide domain. Alternatively, the first polypeptide domain can be coupled indirectly to the second polypeptide domain by coupling with a third polypeptide domain that is coupled directly to the second polypeptide domain. In some cases, a first polypeptide domain is coupled to the N-terminus of a second polypeptide domain. In some cases, a first polypeptide domain is coupled to the C-terminus of a second polypeptide domain. In some cases, a first polypeptide domain is coupled to an internal component of a second polypeptide domain. In some cases, the two or more polypeptide domains are covalently linked. In some cases, the two or more polypeptide domains are noncovalently linked. In some cases, the two or more polypeptide domains are coupled together by a linker. For example, a linker may be a peptide linker. A linker can be a rigid linker, which helps maintain a fixed distance between the polypeptide domains that it links. A linker can be a flexible linker, which can allow some flexibility in movement of one polypeptide domain relative to the other polypeptide domain that it is linked to. In some cases, a linker is a cleavable linker. For example, a cleavable linker can comprise a disulfide bond.
Alternatively, a cleavable linker can be an enzymatic cleavable linker, e.g., a linker comprising a protease cleavage site.
[0183] The present disclosure provides fusion proteins comprising a guidable polypeptide domain (e.g., a CRISPR domain). A fusion protein comprising a guidable polypeptide domain (e.g., a CRISPR domain) can comprise one or more of a FokI domain, a deaminase domain, a reverse transcriptase domain, an RNA binding domain, a transcriptional regulatory domain, a plasma membrane recruitment domain, a transmembrane domain, a signaling domain, a receptor domain, a packaging domain, or a targeting domain.
[0184] In some cases, the present disclosure provides a fusion protein comprising a guidable polypeptide domain (e.g., a CRISPR domain) coupled to a deaminase domain. A guidable polypeptide domain (e.g., a CRISPR domain) coupled to a deaminase domain can be used for base editing. A base editor can be capable of editing a nucleic acid sequence in a target nucleic acid molecule. A base editor can be capable of enabling the generation of base conversions or point mutations in a target nucleic acid. For example, a cytosine base editor can comprise a guidable polypeptide domain (e.g., a CRISPR domain) and a cytidine deaminase domain. An adenine base editor can comprise CRISPR domain and an adenosine deaminase domain. In some cases, a base editor enables the conversion of C to G, A to I, or C to U. A cytosine base editor can be capable of enabling the conversion of a C-G base pair to a T-A base pair. A glycosylase base editor can be capable of enabling the conversion of a G-C base pair to a C-G base pair or a G-T base pair. An adenine base editor can be capable of enabling the conversion of an A-T base pair to a G-C base pair. In some cases, a base editor comprises a catalytically inactive guidable polypeptide domain (e.g., a CRISPR domain) (e.g., dCas9, dCasl2a, or dCasl3b). In other cases, the base editor comprises a guidable polypeptide nickase domain (e.g., nCas9). In some cases, the base editor enables a base pair conversion without introducing a double-stranded break. In some cases, the base editor enables base pair conversions in a target window. In some cases, the base editor comprises a targeting window of from 1 to 20 bases, from 1 to 19 bases, from 1 to 18 bases, from 1 to 17 bases, from 1 to 16 bases, from 1 to 15 bases, from 1 to 14 bases, from 1 to 13 bases, from 1 to 12 bases, from 1 to 11 bases, from 1 to 10 bases, from 1 to 9 bases, from 1 to 8 bases, from 1 to 7 bases, from 1 to 6 bases, from 1 to 5 bases, from 1 to 4 bases, from 1 to 3 bases, or from 1 to 2 bases. In some cases, a base editor has a targeting window of from 3 to 10 bases, from 3 to 9 bases, from 3 to 8 bases, from 3 to 7 bases, from 3 to 6 bases, from 3 to 5 bases, or from 3 to 4 bases. In some cases, the guidable polypeptide domain (e.g., a CRISPR domain) is coupled to the N-terminus of a deaminase domain. In some cases, the guidable polypeptide domain (e.g., a CRISPR domain) is coupled to the C-terminus of a deaminase domain. In some cases, the guidable polypeptide domain (e.g., a CRISPR domain) is coupled to an internal component of a deaminase domain.
[0185] In some cases, the fusion protein comprises a guidable polypeptide domain (e.g., a CRISPR domain) coupled to a reverse transcriptase domain. A guidable polypeptide domain (e.g., a CRISPR domain) coupled to a reverse transcriptase can be capable of enabling prime editing. A prime editor can be capable of editing a nucleic acid sequence in a target nucleic acid molecule. A prime editor can be capable of mediating insertion or deletion of a nucleic acid sequence in a target nucleic acid molecule. In
some cases, the prime editor enables a sequence insertion or sequence deletion without introducing a double -stranded break. In some cases, the prime editor introduces a nick at the target site. The prime editor can enable insertion of a template sequence in a target nucleic acid molecule. The template sequence can comprise the desired edit. In some cases, a prime editor reverse transcribes a template sequence to synthesize a complementary strand. In some cases, the synthesized complementary strand is inserted in the target nucleic acid molecule. In some cases, the prime editor uses a primer to carry out reverse transcription. The prime editor can install nucleotides to the 3’ end of a primer strand. In some cases, a primer strand is generated by nicking the target nucleic acid molecule. In some cases, nicking a strand of the target nucleic acid molecule produces a flap with a 3’ OH group. In some cases, the prime editor uses the flap with the 3’ OH group as the primer to carry out reverse transcription. In some cases, a guidable polypeptide domain (e.g., a CRISPR domain) is coupled to the N-terminus of a reverse transcriptase domain. In some cases, a guidable polypeptide domain (e.g., a CRISPR domain) is coupled to the C-terminus of a reverse transcriptase domain. In some cases, a guidable polypeptide domain (e.g., a CRISPR domain) is coupled to an internal component of a reverse transcriptase domain.
[0186] In some cases, the fusion protein comprises a guidable polypeptide domain (e.g., a CRISPR domain) coupled to a transcriptional regulatory domain. In some cases, a guidable polypeptide domain (e.g., a CRISPR domain) is coupled to a transcriptional regulatory domain. In some cases, a guidable polypeptide domain (e.g., a CRISPR domain) is coupled to a transcriptional activation domain. In some cases, a guidable polypeptide domain (e.g., a CRISPR domain) is coupled to a transcriptional repression domain. In some cases, guidable polypeptide domain (e.g., a CRISPR domain) is coupled to a transcriptional regulatory domain. A guidable polypeptide domain (e.g., a CRISPR domain) coupled to a transcriptional regulatory domain can be capable of enabling CRISPR interference (CRISPRi) or CRISPR activation (CRISPRa). In some cases, a guidable polypeptide domain (e.g., a CRISPR domain) can be coupled to a transcriptional regulatory domain such as P3000 or DNMT3. In some cases, a guidable polypeptide domain (e.g., a CRISPR domain) is coupled to the C-terminus of a transcriptional regulatory domain. In some cases, a guidable polypeptide domain (e.g., a CRISPR domain) is coupled to an internal component of a transcriptional regulatory domain. Any of the payloads described herein can further comprise a plasma membrane recruitment domain, transmembrane domain, a signaling domain, a receptor domain, a packaging domain, or a targeting domain. Any of payloads described herein can comprise or be engineered to comprise a protein tag, a peptide tag, or small molecule tag. For example, a payload can comprise a small nuclear localization signal (NLS), a nuclear export signal (NES), a cell penetrating peptide (CPP), a mitochondria penetrating peptide (MPP), a solubility tag, or a fluorescent tag.
Base editor
[0187] In some cases, the payload to be delivered by the lipid containing particles of the present disclosure comprises a nucleobase editor (also termed as “base editor”) or one or more components of a nucleobase editing (also termed as “base editing”) complex.
[0188] The term “base editor (BE), ” or “nucleobase editor (NBE),” as used herein, can refer to an agent comprising a polypeptide that is capable of making a modification to a base (e.g., A, T, C, G, or U) within
a nucleic acid sequence (e.g., DNA or RNA). In some embodiments, the base editor is capable of deaminating a base within a nucleic acid. In some embodiments, the base editor is capable of deaminating a base within a DNA molecule. In some embodiments, the base editor is capable of deaminating an adenosine (A) in DNA. In some embodiments, the base editor is capable of deaminating a cytosine (C) in DNA. In some embodiments, the base editor is capable of converting a guanine (G) in DNA through a glycoylase.
[0189] In some cases, the payload in the present disclosure comprises a deaminase domain. The deaminase domain can be a natural or an engineered domain. A deaminase domain can be capable of carrying out deamination reactions in DNA. A deaminase domain can be capable of enabling the generation of base conversions or point mutations in a target nucleic acid. For example, a deaminase domain can be a cytidine deaminase domain or an adenosine deaminase domain. A cytidine deaminase domain can be capable of converting cytosine to uracil. A cytidine deaminase domain can be capable of enabling the conversion of a C-G base pair to a T-A base pair. For example, a cytidine deaminase can be or comprise a APOBEC1 cytidine deaminase. An adenosine deaminase domain can be capable of converting an adenosine to hypoxanthine. An adenosine deaminase domain can be capable of converting an adenosine to an inosine. An adenosine deaminase can comprise TadA or a TadA mutant. In some embodiments, TadA comprises a monomer. In some embodiments, TadA comprises a heterodimer comprising a wildtype TadA and a mutated Tad A. In some embodiments, TadA comprises a homodimer comprising two wildtype TadA domains or two mutated TadA domains. An adenosine deaminase domain can be capable of enabling the conversion of an A-T base pair to a G-C base pair. A deaminase domain can be a mutated variant. In some cases, a deaminase domain enables the conversion of C to G, A to I, or C to U.
[0190] In some cases, the payload in the present disclosure comprises a glycosylase domain. The glycosylase domain can be a natural or an engineered domain. A glycosylase-based guanine base editor can be designed to remove G, and the AP site generated is repaired by translesion synthesis and/or DNA replication, leading to G-to-C or G-to-T conversion. A glycosylase domain can be capable of enabling the generation of base conversions or point mutations in a target nucleic acid. For example, a glycosylase domain can be a guanine glycosylase domain. Examples of glycosylase base edits can be found in Sun N, et al.. Mol Ther. 2022 Jul 6;30(7):2452-2463 and Huawei Tong, et al., National Science Review, Volume 10, Issue 8, August 2023, each of which is incorporated in its entirety herein.
[0191] In some cases, the base editor disclosed herein comprises a deaminase or a functional domain thereof (“deaminase domain”) that catalyzes deamination reaction.
[0192] The term "deaminase" or "deaminase domain," as used herein, refers to a protein or enzyme that catalyzes a deamination reaction. In some embodiments, the deaminase or deaminase domain is an adenosine deaminase, catalyzing the deamination of adenosine, converting it to the nucleoside hypoxanthine. In some embodiments, the deaminase or deaminase domain is a cytidine deaminase, catalyzing the hydrolytic deamination of cytidine or deoxycytidine to uridine or deoxyuridine, respectively. In some embodiments, the deaminase or deaminase domain is a cytidine deaminase domain,
catalyzing the hydrolytic deamination of cytosine to uracil. In some embodiments, the deaminase or deaminase domain is a naturally-occurring deaminase from an organism, such as a human, chimpanzee, gorilla, monkey, cow, dog, rat, or mouse. In some embodiments, the deaminase or deaminase domain is a variant of a naturally-occurring deaminase from an organism, that does not occur in nature. For example, in some embodiments, the deaminase or deaminase domain is at least 50%, at least 55%, at least 60%, at least 65%, at least 70%, at least 75% at least 80%, at least 85%, at least 90%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, or at least 99.5% identical to a naturally-occurring deaminase from an organism.
[0193] As used herein, an “adenosine deaminase” is an enzyme that catalyzes the deamination of adenosine, converting it to the nucleoside hypoxanthine. Under standard Watson-Crick hydrogen bond pairing, an adenosine base hydrogen bonds to a thymine base (or a uracil in case of RNA).
When adenine is converted to hypoxanthine, the hypoxanthine undergoes hydrogen bond pairing with cytosine. Thus, a conversion of “A” to hypoxanthine by adenosine deaminase will cause the insertion of “C” instead of a “T” during cellular repair and/or replication processes. Since the cytosine “C” pairs with guanine “G”, the adenosine deaminase in coordination with DNA replication causes the conversion of an A«T pairing to a C*G pairing in the double-stranded DNA molecule.
[0194] In some embodiments, the base editor is a chimeric protein comprising a nucleic acid programmable R/DNA binding protein (napR/DNAbp) fused to a deaminase (e.g., cytidine deaminase or adenosine deaminase) domain. The term “nucleic acid programmable D/RNA binding protein (napR/DNAbp)” refers to any protein that can associate (e.g., form a complex) with one or more nucleic acid molecules (i.e., which can broadly be referred to as a “napR/DNAbp-programming nucleic acid molecule” and includes, for example, guide RNA in the case of Cas systems) which direct or otherwise program the protein to localize to a specific target nucleotide sequence (e.g., a gene locus of a genome, or an RNA molecule) that is complementary to the one or more nucleic acid molecules (or a portion or region thereof) associated with the protein, thereby causing the protein to bind to the nucleotide sequence at the specific target site. This term napR/DNAbp embraces CRISPR Cas9 proteins, as well as Cas9 equivalents, homologs, orthologs, or paralogs, whether naturally occurring or non-naturally occurring (e.g., engineered or recombinant), and can include a Cas9 equivalent from any type of CRISPR system (e.g., type II, V, VI), including Cpfl (a type-V CRISPR-Cas systems), C2cl (a type V CRISPR-Cas system), C2c2 (a type VI CRISPR-Cas system) and C2c3 (a type V CRISPR-Cas system). Further Cas- equivalents are described in Makarova et al., “C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector,” Science 2016; 353(6299), the contents of which are incorporated herein by reference. However, the nucleic acid programmable R/DNA binding protein (napR/DNAbp) that can be used in connection with this disclosure are not limited to CRISPR-Cas systems. The present disclosure embraces any such programmable protein, such as the Argonaute protein from Natronobacterium gregoryi (NgAgo) which can also be used for DNA-guided genome editing. NgAgo-guide DNA system does not require a PAM sequence or guide RNA molecules, which means genome editing can be performed simply by the expression of generic NgAgo protein and introduction of
synthetic oligonucleotides on any genomic sequence. See Gao F, Shen X Z, Jiang F, Wu Y, Han C. DNA- guided genome editing using the Natronobacterium gregoryi Argonaute. Nat Biotechnol 2016; 34(7):768- 73, which is incorporated herein by reference.
[0195] In some cases, the napR/DNAbp is derived from a nuclease disclosed herein, such as, Cas9 (e.g., dCas9 and nCas9), CasX, CasY, Casl4, Cpfl, C2cl, C2c2, C2c3, Argonaute protein, or a variant thereof. In some embodiments, the base editor comprises a Cas9 (e.g., dCas9 and nCas9), CasX, CasY, Cpfl, C2cl, C2c2, C2c3, or Argonaute protein fused to a deaminase (e.g, cytidine deaminase or adenosine deaminase). In some embodiments, the base editor comprises a Cas9 nickase (nCas9) fused to an deaminase (e.g., cytidine deaminase or adenosine deaminase). In some embodiments, the base editor comprises a CasX protein fused to a deaminase (e.g, cytidine deaminase or adenosine deaminase). In some embodiments, the base editor comprises a nuclease-inactive Cas9 (dCas9) fused to a deaminase (e.g., cytidine deaminase or adenosine deaminase). In some embodiments, the base editor comprises a CasY protein fused to a deaminase (e.g, cytidine deaminase or adenosine deaminase). In some embodiments, the base editor comprises a Casl4 protein fused to a deaminase (e.g., cytidine deaminase or adenosine deaminase). In some embodiments, the base editor comprises a Cpfl protein fused to a deaminase (e.g., cytidine deaminase or adenosine deaminase). In some embodiments, the base editor comprises a C2cl protein fused to a deaminase (e.g., cytidine deaminase or adenosine deaminase). In some embodiments, the base editor comprises a C2c2 protein fused to a deaminase (e.g., cytidine deaminase or adenosine deaminase). In some embodiments, the base editor comprises a C2c3 protein fused to a deaminase (e.g., cytidine deaminase or adenosine deaminase). In some embodiments, the base editor comprises an Argonaute protein fused to a deaminase (e.g., cytidine deaminase or adenosine deaminase).
[0196] In some embodiments, the adenosine deaminases provided herein are capable of deaminating adenosine. In some embodiments, the adenosine deaminases provided herein are capable of deaminating adenosine in a deoxyadenosine residue of DNA. The adenosine deaminase can be derived from any suitable organism (e.g., E. coli). In some embodiments, the adenosine deaminase is a naturally-occurring adenosine deaminase that includes one or more mutations corresponding to any of the mutations provided herein (e.g., mutations in ecTadA). One of skill in the art will be able to identify the corresponding residue in any homologous protein and in the respective encoding nucleic acid by methods well known in the art, e.g., by sequence alignment and determination of homologous residues. Accordingly, one of skill in the art would be able to generate mutations in any naturally-occurring adenosine deaminase (e.g., having homology to ecTadA) that corresponds to any of the mutations described herein, e.g. , any of the mutations identified in ecTadA. In some embodiments, the adenosine deaminase is from a prokaryote. In some embodiments, the adenosine deaminase is from a bacterium. In some embodiments, the adenosine deaminase is from Escherichia coli, Staphylococcus aureus, Salmonella typhi, Shewanella putrefaciens, Haemophilus influenzae, Caulobacter crescentus, or Bacillus subtilis. In some embodiments, the adenosine deaminase is from E. coli.
[0197] In some cases, the deaminase domain of the base editor disclosed herein is derived from a cytidine deaminase. In some cases, the cytidine deaminase domain is derived from the apolipoprotein B mRNA-editing complex (APOBEC) family deaminase, such as APOBEC 1 deaminase, APOBEC2 deaminase, APOBEC3A deaminase, APOBEC3B deaminase, APOBEC3C deaminase, APOBEC3D deaminase, APOBEC3F deaminase, APOBEC3G deaminase, or APOBEC3H deaminase. In some cases, the cytidine deaminase is a modification of an APOBEC family deaminase. In some cases, the cytidine deaminase is an evolved derivative of an APOBEC family deaminase.
[0198] In some embodiments, the base editor comprises BE1, BE2, BE3, BE4, BE4max, or another base editor variant. In some embodiments, the base editor comprises BE4max (R33A) AUGI-hUNG complex (CGBE1).
[0199] In some embodiments, the base editor is fused to, or further comprises as part of a chimeric protein, an inhibitor of base excision repair, for example, a uracil clycosylase inhibitor (UGI) domain. In some embodiments, the base editor is fused to one, two, or three UGI domains. In some embodiments, the base editor is fused to one or more UGI domains. In some embodiments, a UGI domain reduces off target effects, specifically the conversion of C to G or C to A.
[0200] In some cases, the base editor disclosed herein is a chimeric protein that comprises a structure such as, NEE-fdeaminase domain] -[napR/DNAbp]- [UGI domain]-COOH; NH2-| deaminase domain]- [napR/DNAbp]-[UGI]-[UGI]-COOH; NH2-[deaminase domain] -[napR/DNAbp] -[UGI] -CO OH; NH2- [UGI]-[ deaminase domain]-[napR/DNAbp]-COOH; NH2-[deaminase domain] -[UGI] -[napR/DNAbp] - COOH; NH2-[napR/DNAbp]-[UGI]-[deaminase domain]-COOH; or NH2- [napR/DNAbp] -[deaminase domain]-[UGI]-COOH; wherein each instance of comprises an optional linker.
[0201] In some cases, the base editor is fused to, or further comprises as part of a chimeric protein, a uracil binding protein (UBP). The term “uracil binding protein” or “UBP,” as used herein, refers to a protein that is capable of binding to uracil. In some embodiments, the uracil binding protein is a uracil modifying enzyme. In some embodiments, the uracil binding protein is a uracil base excision enzyme. In some embodiments, the uracil binding protein is a uracil DNA glycosylase (UDG). In some embodiments, a uracil binding protein binds uracil with an affinity that is at least 1%, 2%, 3%, 5%, 10%, 15%, 20%, 30%, 40%, 50%, 60%, 70%, 80%, 90%, or at least 95% of the affinity that a wild type UDG (e.g. , a human UDG) binds to uracil. The term “base excision enzyme” or “BEE,” as used herein, refers to a protein that is capable of removing a base (e.g. , A, T, C, G, or U) from a nucleic acid molecule (e.g. , DNA or RNA). In some embodiments, a BEE is capable of removing a cytosine from DNA. In some embodiments, a BEE is capable of removing a thymine from DNA. Exemplary BEEs include, without limitation UDG Tyrl47Ala, and UDG Asn204Asp as described in Sang et al., “A Unique Uracil-DNA binding protein of the uracil DNA glycosylase superfamily,” Nucleic Acids Research, Vol. 43, No. 17 2015; the entire contents of which are hereby incorporated by reference.
[0202] In some embodiments, the UBP is a uracil modifying enzyme. In some embodiments, the UBP is a uracil base excision enzyme. In some embodiments, the UBP is a uracil DNA glycosylase. In some embodiments, the UBP is any of the uracil binding proteins provided herein. For example, the UBP can
be a UDG, a UdgX, a UdgX*, a UdgX_On, or a SMUG1. In some embodiments, the UBP comprises an amino acid sequence that is at least 75%, 80%, 85%, 90%, 95%, 96%, 97%, 98%, 99%, or 99.5% identical to a uracil binding protein, a uracil base excision enzyme or a uracil DNA glycosylase (UDG) enzyme.
[0203] In some cases, the base editor is fused to, or comprises as a part of the chimeric protein, a nucleic acid polymerase domain (NAP). For instance, the nucleic acid polymerase domain is a eukaryotic nucleic acid polymerase domain. In some cases, the nucleic acid polymerase domain is a DNA polymerase domain. In some cases, the nucleic acid polymerase domain has translesion polymerase activity. In some cases, the nucleic acid polymerase domain is a translesion DNA polymerase. In some cases, the nucleic acid polymerase domain is from Rev7, Revl complex, polymerase iota, polymerase kappa, and polymerase eta. In some cases, the nucleic acid polymerase domain is selected from the group of eukaryotic polymerases consisting of alpha, beta, gamma, delta, epsilon, gamma, eta, iota, kappa, lambda, mu, and nu.
[0204] In some cases, the base editor disclosed herein is a chimeric protein that comprises a structure such as, NIU-fdeaminase domain] -[napR/DNAbp domain]-[UBP]-[NAP]-COOH; NH2-| deaminase domain] - [napR/DNAbp] - [NAP] - [UBP] -COOH; N H2- [deaminase domain] - [NAP] - [napR/DNAbp] - [UBP]-COOH; or NH2-[NAP]-[ deaminase domain]-[napR/DNAbp]-[UBP]-COOH; wherein each instance of comprises an optional linker.
[0205] In some cases, the base editor disclosed herein is complexed with a napR/DNAbp-programming nucleic acid molecule. In some cases, the base editing system disclose herein comprises a base editor and a napR/DNAbp-programming nucleic acid molecule, e.g., the base editor complexed with the napR/DNAbp-programming nucleic acid molecule. In some cases, the lipid containing particles of the present disclosure deliver a base editing system that comprises both a base editor and a napR/DNAbp- programming nucleic acid molecule, e.g., the base editor complexed with the napR/DNAbp-programming nucleic acid molecule. In some cases, a base editor is delivered separately from the napR/DNAbp- programming nucleic acid molecule through lipid containing particles disclosed herein, or together with other delivery methods, into a cell.
[0206] The term “napR/DNAbp-programming nucleic acid molecule” or equivalently “guide sequence” refers the one or more nucleic acid molecules which associate with and direct or otherwise program a napR/DNAbp protein to localize to a specific target nucleotide sequence (e.g., a gene locus of a genome) that is complementary to the one or more nucleic acid molecules (or a portion or region thereof) associated with the protein, thereby causing the napR/DNAbp protein to bind to the nucleotide sequence at the specific target site. An example is a guide RNA of a Cas protein of a CRISPR-Cas genome editing system.
[0207] Exemplary configurations, sequences, and mutations thereof for deaminase domains, napR/DNAbp domains, UGI domains, and whole base editor proteins, and exemplary configurations of a base editing system (e.g., comprising both a base editor and a napR/DNAbp-programming nucleic acid molecule) that can be delivered by a lipid containing particle disclosed herein include those described in
U.S. Patent Publication Nos. US20170121693, US20180073012, US20180312828, US20180170984, US2020010835, US2020172931, US20210230577, US20210198330, US20210277379, US2020399626, US2021371858, US2021380955, US2021277379, US2021301274, US20220127622, US20220313799, US20230055682, US20230159913, US20230086199, US20220127622, US20220313799, US20230279373, and US20230055682, each of which is incorporated herein by reference in its entirety. Exemplary configurations, sequences, and mutations thereof for deaminase domains, napR/DNAbp domains, UGI domains, and whole base editor proteins, that can be delivered by a lipid containing particle disclosed herein also include those described in Komor AC et al. Nature. 2016 May 19;533(7603):420-4; Kim YB et al. Nat Biotechnol. 2017 Apr;35(4):371-376; Rees HA et al. Nat Commun. 2017 Jun 6;8: 15790; Newby GA et al. Mol Ther. 2021 Nov 3;29(11):3107-3124; Huang TP et al. NatProtoc. 2021 Feb;16(2): 1089-1128; Lapinaite A et al. Science. 2020 Jul 31;369(6503):566-571; Anzalone AV et al. Nat Biotechnol. 2020 Jul;38(7):824-844; Rees HA et al. Nat Rev Genet. 2018 Dec;19(12):770-788; Koblan LW et al. Nat Biotechnol. 2018 Oct;36(9): 843-846; and Gaudelli NM et al. Nature. 2017 Nov 23;551(7681):464-471; each of which is incorporated herein by reference in its entirety.
Prime editor
[0208] In some cases, the lipid delivery particles disclosed herein is capable of delivering a payload, such as a prime editing system, or one or more components thereof, such as a ribonucleoprotein (RNP) complex, into a cell in vitro, ex vivo, or in vivo. In some embodiments, the prime editing system, or one or more components thereof, is within the inside cavity of the protein core of the lipid delivery particles disclosed herein.
[0209] Prime editing system is a ‘search-and-replace’ genome editing technology by which the genome of living organisms can be modified. The term "prime editing system" or "prime editor (PE)" refers the compositions involved in genome editing using target-primed reverse transcription (TPRT) describe herein, can comprise a nucleic acid-guided polypeptide, e.g., nucleic acid-guided polypeptide, a nucleic acid polymerase, chimeric proteins (e.g., comprising guidable polypeptide domain and reverse transcriptase), guide nucleic acid molecule (e.g., guide RNAs), and complexes comprising fusion proteins and guide RNAs, as well as accessory elements, such as second strand nicking components and 5' endogenous DNA flap removal endonucleases (e.g., FEN1) for helping to drive the prime editing process towards the edited product formation.
[0210] In some embodiments, the prime editing system disclosed herein comprises a ribonucleoprotein (RNP) complex. In some cases, the RNP complex comprises a prime editor and a guide nucleic acid molecule. In some cases, the prime editor is formed between one or more proteins and one or more polynucleotides. The prime editor can comprise a nucleic acid-guided polypeptide. The guidable polypeptide domain can comprise a nucleic acid-guided polypeptide, for example a nuclease (e.g., a Cas protein). For instance, the prime editor can comprise a fusion protein, comprising a nucleic acid programmable R/DNA binding protein (e.g., a nuclease, such as a Cas protein) and a nucleic acid polymerase (e.g., a reverse transcriptase or any suitable DNA polymerase). In some cases, the nucleic
acid polymerase is coupled to the nucleic acid-guided polypeptide. In some cases, the guide nucleic acid molecule can comprise a guide nucleic acid molecule, e.g., a guide RNA. In some cases, the prime editor is operably linked to the guide nucleic acid molecule via a linker, forming the RNP complex. In some cases, the prime editor is directly linked to the guide nucleic acid molecule, forming the RNP complex. [0211] In a specific instance, prime editing system comprises a fusion protein that comprises an engineered Cas9 nickase and a reverse transcriptase, and the fusion protein is paired with an engineered prime editing guide RNA (PEgRNA). In some cases, the PEgRNA can direct Cas9 to a target site within a host cell where the lipid delivery particles are delivered. In some cases, the peg RNA can encode the information for installing the desired edit. In some cases, the prime editing system can function through a multi-step process: 1) the Cas9 domain can bind and nick the target genomic DNA site, which is specified by a spacer sequence in the PEgRNA; 2) the reverse transcriptase can use the nicked genomic DNA as a primer to initiate synthesis of an edited DNA strand using an engineered extension on the PEgRNA as a template for reverse transcription, which can generate a single -stranded 3' flap containing the edited DNA sequence; 3) cellular DNA repair mechanism can resolve the 3' flap intermediate by the displacement of a 5' flap species that occurs via invasion by the edited 3' flap, excision of the 5' flap containing the original DNA sequence, and ligation of the new 3' flap to incorporate the edited DNA strand, forming a heteroduplex of one edited and one unedited strand; and 4) cellular DNA repair mechanism can replace the unedited strand within the heteroduplex using the edited strand as a template for repair, which completes this editing process. In some embodiments, the prime editing machinery edits a target DNA molecule. In some embodiments, the prime editing machinery edits a target RNA molecule. Examples of targeting RNA molecules using prime editing are described in international patent application WO2021072328 and U.S. Patent Application number US20230357766, each of which is incorporated in its entirety. In other instances, a prime editing system is a multi-flap prime editing system that can simultaneously edit both DNA strands. For example, a dual-flap prime editing system comprises two PEgRNAs, which can be used to target opposite strands of a genomic site and direct the synthesis of two complementary 3’ flaps containing edited DNA sequence. The pair of edited DNA strands (3’ flaps) does not need to directly compete with 5 ’ flaps in endogenous genomic DNA, as the complementary edited strand is available for hybridization instead. In this instance, both strands of the duplex are synthesized as edited DNA, the dual-flap prime editing system obviates the need for the replacement of the non-edited complementary DNA strand. Instead, cellular DNA repair machinery can only excise the paired 5’ flaps (original genomic DNA) and ligate the paired 3’ flaps into the locus.
[0212] In some embodiments, a prime editing system can be paired with a separate Cas9 nickase and a separate gRNA that nicks the DNA at a locus that is different than the locus targeted by the PEgRNA. In some embodiments, one or more prime editing systems can be paired, each targeting a different locus. In some cases, pairing of two prime editing systems, each of which targets a different locus on the same chromosome, can install large insertions, deletions, or modifications. In some cases, pairing of two prime editing systems, each of which targets a different locus, can install large structural modifications. In some embodiments, a prime editor can install up to 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19,
or 20 modifications. In some embodiments, a prime editor can install 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, or more modifications. In some embodiments, a prime editor can install up to about 25, about 30, about 35, about 40, about 45, about 50, about 55, about 60, about 65, about 70, about 75, about 80, about 85, about 90, about 95, or about 100 modifications. In some embodiments, a prime editor can install more than about 25, about 30, about 35, about 40, about 45, about 50, about 55, about 60, about 65, about 70, about 75, about 80, about 85, about 90, about 95, or about 100 modifications. In some cases, a prime editor can install more than 100 modifications. In some embodiments, more than one prime editor can be used to install mutations more than 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, or more modifications. In some embodiments, more than one prime editor can be used to install mutations more than about 25, about 30, about 35, about 40, about 45, about 50, about 55, about 60, about 65, about 70, about 75, about 80, about 85, about 90, about 95, or about 100 modifications. In some embodiments, more than one prime editor can be used to install mutations more than about Ikb, 5kb, lOkb, 20kb, 30kb, 40kb, 50kb, 60kb, 70kb, 80kb, 90kb, or more. [0213] Different variants of prime editors have been developed, such as prime editors (PE) PEI, PE2, PE3, PE4, or PE5, some of which are described in Liu, D. et al., Nature 2019, 576, 149-157 and Huang Z, Liu G. Front Bioeng Biolechnol. 2023 ; 11 : 1039315 , U. S . Patent Application numbers US20210292769, US20230090221, US2022078655, US20230220374, each of which is hereby incorporated by reference herein in its entirety. In some cases, the prime editor comprises a reverse transcriptase (RT) fused with Cas9 H 840A nickase (Cas9n (H840A)) and a prime-editing guide RNA (pegRNA). In some cases, the RT comprises an RNA-dependent DNA polymerase. In some cases, the RT comprises a protein derived from a retrovirus. In some embodiments, the RT comprises Moloney Murine Leukemia Virus (M- MMLV) RT. In some embodiments, the RT comprises a RT from HFV, LtrA, HERV-Kcon, Tel4c, Marathon, Gst-IIC, MA-INT5, or another RT ortholog. In some cases, the RT is modified, mutated, truncated, or evolved. In some cases, the RT comprises a full length RT protein. In some cases, the RT comprises a truncated RT. In some cases, the RT is fused to the Cas protein. In some cases, the RT is fused to the Cas protein at the N terminus to the Cas protein. In some cases, the RT is fused to the Cas protein at the C terminus of the Cas protein. In some cases, the RT is fused to the Cas protein as an inlaid fusion. In some cases, the RT is untethered to the Cas protein. Examples of prime editing architecture are described in Griinewald, J., et al. , Nat Biotechnol 41, 337-343 (2023) and Gao Z, et al., Mol Ther. 2022; 30(9):2942-2951, each of which is incorporated herein in its entirety.
[0214] In some cases, the prime editor comprises (a) a fusion protein having the following N-terminus to C-terminus structure: [NLS]-[Cas9(H840A)]- [linker] -[MMLV RT(wt)] and (b) a PEgRNA. In some cases, the prime editor comprises (a) a fusion protein having the following N-terminus to C-terminus structure: [NLS]-[Cas9(H840A)]-[linker]-[MMLV_RT(D200N)(T330P)(L603W)(T306K) (W313F)] and (b) a PEgRNA. In some cases, the prime editor comprises (a) a fusion protein having the following N- terminus to C-terminus structure: [NLS]-[Cas9(H840A)]-[linker]- [MMLV_RT(D200N)(T330P)(L603W)(T306K) (W313F)]; (b) a PEgRNA; and (c) a nicking guide RNA that introduces a nick in the non-edited DNA strand. In some cases, the addition of nicking guide RNA
increases the chances of the unedited strand to be repaired rather than the edited strand. In some cases, the prime editor comprises (a) a fusion protein having the following N-terminus to C-terminus structure: [NLS]-[Cas9(H840A)]-[linker]-[MMLV_RT(D200N)(T330P)(L603W)(T306K) (W313F)]; (b) a PEgRNA; and (c) a nicking guide RNA that is designed with a spacer that matches only the edited strand but not the original allele before editing, so that the nicking guide RNA is not introduced until after the desired edit is installed. In some cases, the prime editor comprises (a) a fusion protein having the following N-terminus to C-terminus structure: [NLS]-[Cas9(H840A)]-[linker]- [MMLV_RT(D200N)(T330P)(L603W)(T306K) (W313F)]; (b) a PEgRNA; and (c) evading specific DNA mismatch repair (MMR) protein, such as co-expression of a dominant negative MMR protein, such as MLHldn (e.g., MLH1 A754-756). In some cases, the prime editor comprises (a) a fusion protein having the following N-terminus to C-terminus structure: [NLS]-[Cas9(H840A)]-[linker]- [MMLV_RT(D200N)(T330P)(L603W)(T306K) (W313F)]; (b) a PEgRNA; (c) a nicking guide RNA that introduces a nick in the non-edited DNA strand; and (d) evading specific DNA mismatch repair (MMR) protein, such as co-expression of a dominant negative MMR protein, such as MLHldn (e.g., MLH1 A754-756). Evading MMR protein, such as by co-expression of MMR protein MLHldn can increase efficiency of prime editing, as described in International Publication No., WO2023102538 and Chen et al., Cell Volume 184, Issue 22, 28 October 2021, Pages 5635-5652.e29, each of which is hereby incorporated by reference herein in its entirety. An exemplary sequence for MLHldn (MLH1 A754-756) is:
MSFVAGVIRRLDETVVNRIAAGEVIQRPANAIKEMIENCLDAKSTSIQVIVKEGGLKLIQIQDNGT GIRKEDLDIVCERFTTSKLQSFEDLASISTYGFRGEALASISHVAHVTITTKTADGKCAYRASYSD GKLKAPPKPCAGNQGTQITVEDLFYNIATRRKALKNPSEEYGKILEVVGRYSVHNAGISFSVKKQ GETVADVRTLPNASTVDNIRSIFGNAVSRELIEIGCEDKTLAFKMNGYISNANYSVKKCIFLLFINH RLVESTSLRKAIETVYAAYLPKNTHPFLYLSLEISPQNVDVNVHPTKHEVHFLHEESILERVQQHI ESKLLGSNSSRMYFTQTLLPGLAGPSGEMVKSTTSLTSSSTSGSSDKVYAHQMVRTDSREQKLD AFLQPLSKPLSSQPQAIVTEDKTDISSGRARQQDEEMLELPAPAEVAAKNQSLEGDTTKGTSEMS EKRGPTSSNPRKRHREDSDVEMVEDDSRKEMTAACTPRRRIINLTSVLSLQEEINEQGHEVLREM LHNHSFVGCVNPQWALAQHQTKLYLLNTTKLSEELFYQILIYDFANFGVLRLSEPAPLFDLAMLA LDSPESGWTEEDGPKEGLAEYIVEFLKKKAEMLADYFSLEIDEEGNLIGLPLLIDNYVPPLEGLPIF ILRLATEVNWDEEKECFESLSKECAMFYSIRKQYISEESTLSGQQSEVPGSIPNSWKWTVEHIVYK ALRSHILPPKHFTEDGNILQLANLPDLYKVF. In some cases, other strategies for evading MMR protein can be adopted, such as installing silent mutations next to the desired edit or co-expressing an antibody targeting the MMR protein. In some cases, the foregoing prime editor comprises (a) a fusion protein having the following N-terminus to C-terminus structure: [bipartite NLSI- [Cas9(R221K)(N394K)(H840A)]-[linker]-[MMLV_RT(D200N)(T330P)(L603W)]-[bipartite NLS]- [NLS] instead. In some cases, the components in the foregoing prime editors are packaged in a single lipid delivery particle. In some cases, the components in the foregoing prime editors are packaged in two or more lipid deliver particles that are delivered to the recipient cell simultaneously. In some cases, the
components in the foregoing prime editors are packaged in two or more lipid deliver particles that are delivered to the recipient cell sequentially.
[0215] In some cases, the prime editing system can comprise a flap endonuclease (e.g., FEN1 or variant thereof) that is delivered as a part of the lipid delivery particle (e.g. , fused to a plasma membrane recruitment element as a chimeric protein). The flap endonuclease can comprise naturally occurring enzymes that process the removal of 5' flaps formed during cellular processes, including DNA replication. The flap endonuclease includes those described in Patel et al., Nucleic Acids Research, 2012, 40(10): 4507-4519 and Tsutakawa et al., Cell, 2011, 145(2): 198-211, each of which is incorporated herein by reference in its entirety.
[0216] Additional elements that can be delivered as a part of the prime editing system via the lipid delivery particles (e.g., fused to the nucleic acid-guided polypeptide, or fused to plasma membrane recruitment element) described herein include inhibitor of base repair (e.g., proteins that inhibit a nucleic acid repair enzyme, for example, a base excision repair enzyme), uracil glycosylase inhibitor domains (e.g., protein that inhibits a uracil-DNA glycosylase base-excision repair enzyme), epitope tags, and reporter gene sequences, including those described in International Publication No. WO2023205744, which is incorporated herein by reference in its entirety.
Epigenetic editor
[0217] In some cases, the payload to be delivered by the lipid containing particles of the present disclosure comprises an epigenetic editor or one or more components of an epigenetic editing complex (e.g. , comprising an epigenetic editor and a nucleic acid molecule that guides the epigenetic editor to bind and/or modify one or more specific target sequences).
[0218] In some cases, the epigenetic editor or epigenetic editing complex disclosed herein has epigenetic activities, such as, methyltransferase activity, demethylase activity, dismutase activity, alkylation activity, depurination activity, oxidation activity, pyrimidine dimer forming activity, integrase activity, transposase activity, recombinase activity, polymerase activity, ligase activity, helicase activity, photolyase activity or glycosylase activity, acetyltransferase activity, deacetylase activity, kinase activity, phosphatase activity, ubiquitin ligase activity, deubiquitinating activity, adenylation activity, deadenylation activity, SUMOylating activity, deSUMOylating activity, ribosylation activity, deribosylation activity, myristoylation activity, remodelling activity, protease activity, oxidoreductase activity, transferase activity, hydrolase activity, lyase activity, isomerase activity, synthase activity, synthetase activity, or demyristoylation activity. In some cases, the epigenetic editor or epigenetic editing complex disclosed herein has a chromosome modification enzyme, or a functional domain that has the functional activity equivalent to a chromosome modification enzyme, such as a methylase, demethylase, acetylase, deacetylase, deaminase, phosphorylase, dephosphorylase, histone modifying enzyme, or nucleotide modifying enzyme. In some cases, the epigenetic editor or epigenetic editing complex disclosed herein has a histone modifying enzyme, or a functional domain that has the functional activity equivalent to a histone modifying enzyme. In some cases, the epigenetic editor or epigenetic editing complex disclosed
herein has a nucleotide modifying enzyme, or a functional domain that has the functional activity equivalent to a nucleotide modifying enzyme.
[0219] In some embodiments, the epigenetic editor or epigenetic editing system comprises a protein domain that represses expression of the target gene. For example, the epigenetic editor or epigenetic editing system can comprise a functional domain derived from a zinc finger repressor protein. Sequences of exemplary functional domains of an epigenetic editor or epigenetic editing system that can reduce or silence target gene expression are provided can be found in PCT/US2021/030643 and Tycko et al. (Tycko J, DelRosso N, Hess GT, Aradhana, Baneqee A, Mukund A, Van MV, Ego BK, Yao D, Specs K, Suzuki P, Marinov GK, Kundaje A, Bassik MC, Bintu L. High-Throughput Discovery and Characterization of Human Transcriptional Effectors. Cell. 2020 Dec 23;183(7):2020-2035.el6. doi:
10. 1016Zj.cell.2020. i l.024. Epub 2020 Dec 15. PMID: 33326746; PMCID: PMC8178797.), each of which is incorporated here by reference in its entirety.
[0220] In some embodiments, the epigenetic editor or epigenetic editing system makes an epigenetic modification at a target gene that activates expression of the target gene. In some embodiments, the epigenetic editor or epigenetic editing system modifies the chemical modification of DNA or histone residues associated with the DNA at a target gene harboring the target sequence, thereby activating or increasing expression of the target gene. In some embodiments, the epigenetic editor or epigenetic editing system comprises a DNA demethylase, a DNA dioxygenase, a DNA hydroxylase, or a histone demethylase domain.
Nucleic acids and polynucleotides
[0221] In some cases, the lipid delivery particle of the present disclosure comprises a payload comprising a nucleic acid. The nucleic acid as a payload can comprise or be composed of one or more nucleotides. Nucleotides are referred to by their commonly accepted single-letter codes: A represents adenine, C represents cytosine, G represents guanine, T represents thymine, U represents uracil, I represents inosine. Unless otherwise indicated, nucleotide sequences are written from left to right in a 5' to 3' orientation. In some cases, the nucleic acid as a payload comprises a polynucleotide. The nucleic acid as a payload can comprise DNA or RNA. In some cases, the nucleic acid as a payload comprises or encodes a gene. The nucleic acid as a payload can comprise or encode any of the polynucleotides described elsewhere herein. The nucleic acid as a payload can be a vector encoding any of the polypeptide domains described elsewhere herein. In some cases, the nucleic acid as a payload is an engineered polynucleotide.
[0222] In some cases, the payload does not comprise a repair template. In some cases, the double stranded break is repaired through non-homologous end joining. In some cases, the payload comprises a repair template. The repair template can be double -stranded or single-stranded. The repair template can comprise a template sequence comprising a desired edit to be introduced in a target nucleic acid molecule. In some cases, the repair template is a homology-directed repair template. A homology-directed repair template can comprise a homology arm that is homologous to a sequence in the target nucleic acid. In
some cases, the payload comprises a DNA-synthesis template comprising a DNA-synthesis template sequence. The DNA-synthesis template can comprise a desired edit to be introduced in a target nucleic acid molecule. The DNA-synthesis template can be a template for a DNA polymerase or a reverse transcriptase to carry out DNA synthesis. For example, in prime editing, a prime editor can use the DNA synthesis template sequence to synthesize a DNA strand that is complementary to the DNA synthesis template sequence. In some cases, the DNA strand is inserted into the target nucleic acid. In some cases, the nucleic acid comprising the DNA synthesis template sequence also comprises a primer-binding sequence. A primer-binding sequence can be complementary to a sequence in a primer strand to which a DNA polymerase or reverse transcriptase can add nucleotides. In some cases, the primer strand is part of a target nucleic acid molecule. A primer-binding sequence can be complementary to a sequence in the target nucleic acid.
[0223] In some cases, the payload comprises a double-stranded DNA containing a desired gene sequence to be inserted in the target nucleic acid molecule. In some cases, the double-stranded DNA is configured to couple to a transposase domain. In some cases, the payload is delivered in the same particle as the transposase domain. In some cases, the payload is delivered in a separate particle as the transposase domain.
[0224] In some cases, the payload comprises a polynucleotide that is configured to bind to a guidable polypeptide domain. In some cases, the polynucleotide directs a guidable polypeptide domain to a sequence in a target nucleic acid molecule. In some cases, the polynucleotide comprises a scaffold segment configured to bind to a guidable polypeptide domain (e.g., Cas9 or Casl2). In some cases, polynucleotide comprises a spacer sequence that is complementary to a target sequence in the target nucleic acid molecule and is capable of hybridizing to the target sequence. The polynucleotide can be a natural molecule or an engineered or synthetic molecule. The polynucleotide can be derived or share sequence or structural similarities to CRISPR RNA (crRNA), a tracrRNA, or a scoutRNA encoded in a CRISPR system. In some cases, the polynucleotide is engineered to be a single RNA guide (sgRNA) comprising elements of the crRNA and the tracrRNA. In some cases, the polynucleotide comprises a scaffold segment and a spacer sequence. The scaffold segment can be configured to bind to a guidable polypeptide domain. The scaffold segment can be specific to a specific type of guidable polypeptide (e.g., Cas9 or Casl2). In some cases, the spacer sequence is programmed to be any sequence. In some cases, the spacer sequence is programmed to a sequence complementary to a target nucleic acid sequence.
[0225] In some cases, the payload comprises a polynucleotide that is a guide nucleic acid molecule for a prime editing system, e.g., a prime editing guide RNA (PEgRNA). In some cases, the PEgRNA is capable of (i) identifying a target nucleotide sequence to be edited, and (ii) encoding new genetic information that replaces the targeted sequence. In some cases, a guide nucleic acid molecule for a prime editing system comprises two or more guide RNAs. In some cases, a guide nucleic acid molecule for a prime editing system comprises a nicking guide RNA. In some cases, a guide RNA comprises (A) a primer binding site, (B) a clamp segment, (C) a sequence encoding new genetic information that replaces the targeted sequence, (D) an aptamer, (E) spacer, or (F) scaffold, or any combinations thereof. In some cases, a guide
RNA comprises a sequence encoding new genetic information that replaces the targeted sequence, a spacer, and scaffold. In some cases, a guide RNA comprises a spacer and scaffold. In some cases, the guide nucleic acid molecule is heterologous to the cell or host receiving the lipid delivery particle. [0226] In some cases, the PEgRNA comprises an extended single guide RNA (sgRNA) containing a primer binding site (PBS) and a template sequence for nucleic acid polymerase (e.g., reverse transcriptase or DNA polymerase). For example, a PEgRNA can comprise an architecture corresponding to 5'-[spacer]- [guide RNA core] -[extension arm]-3'. The spacer sequence can comprise about 20 nucleotides in length. The spacer sequence can bind to a protospacer in a target nucleic acid molecule. The spacer sequence can guide the nucleic acid-guided polypeptide (e.g., Cas9) to the target nucleic acid molecule. The guide RNA core can be responsible for binding of the nucleic acid-guided polypeptide (e.g., Cas9). The extension arm can comprise a primer binding site, an edit template, and a homology arm, in a 3' to 5' direction. The PEgRNA can further comprise, optionally, a 3 ’ end modifier region, 5 ’ end modifier region, a transcriptional signal at the 3’ end. The PEgRNA can optionally comprise a secondary structure, such as, hairpins, stem/loops, toe loops, RNA-binding protein recruitment domains (e.g., the MS2 aptamer which recruits and binds to the MS2cp protein). In some cases, the PEgRNA comprises an aptamer and the prime editor further comprises an aptamer binding protein (e.g. , fused to Cas protein or reverse transcriptase). Guide RNAs including an aptamer include those described in International Publication No. W02023205708, which is hereby incorporated herein by reference in its entirety. Homology arm can encode a portion of a resulting reverse transcriptase-encoded single strand DNA flap to be integrated into the target DNA site by replacing the endogenous strand. The portion of the single strand DNA flap encoded by the homology arm is complementary to the non-edited strand of the target DNA sequence, which facilitates the displacement of the endogenous strand and annealing of the single strand DNA flap in its place, thereby installing the edit. The edit template can comprise a sequence corresponding to new genetic information that replaces the targeted sequence, i.e., a single strand RNA of the PEgRNA that codes for a complementary single strand DNA that is either the sense or the antisense strand of the new genetic information that replaces the targeted sequence and which is incorporated into the genomic DNA target locus through the prime editing process.
[0227] In some cases, during genome editing, the primer binding site allows the 3 ’ end of the nicked DNA strand to hybridize to the PEgRNA, while the reverse transcriptase template serves as a template for the synthesis of edited genetic information. A prime editing system can allow DNA synthesis based on the reverse transcriptase template at a nick site a single 3' flap, which becomes integrated into a target nucleic acid on the same strand. In other embodiments, a prime editing system can be a multi-flap prime editing system that generate pairs or multiple pairs of 3' flaps on different strands, which form duplexes comprising desired edits and which become incorporated into target nucleic acid molecules, e.g., at specific loci or edit sites in a genome. In some embodiments, the pairs or multiple pairs of 3' flaps form duplexes because they comprise reverse complementary sequences which anneal to one another once generated by the prime editors described herein. The duplexes can be incorporated into the target site by cell -driven mechanisms that naturally replace the endogenous duplex sequences located between adjacent
nick sites. In certain embodiments, the new duplex sequences can be introduced at one or more locations (e.g., at adjacent genomic loci or on two different chromosomal locations), and can comprise one or more sequences of interest, e.g., protein-encoding sequence, peptide-encoding sequence, or RNA-encoding sequence.
[0228] In some cases, the payload comprises a polynucleotide comprising a scaffold segment, a spacer sequence, a DNA synthesis template, and a primer-binding sequence. In some cases, the scaffold segment and a spacer sequence are on a first nucleic acid molecule and the DNA synthesis template and the primer-binding sequence are on a second nucleic acid molecule.
[0229] In some cases, the guide RNA further comprises a clamp segment. In some cases, the guide RNA comprising, from 3’ to 5’, a primer binding site, a sequence encoding at least a portion of the first recombinase recognition sequence, a clamp segment, scaffold, and spacer. The clamp segment comprises a sequence that, after being reverse transcribed is at least partially complementary to a genomic site close to the primer binding site and where the spacer binds. Without wishing to be bound by a certain theory, the clamp segment can enhance integration efficiency of the new genetic material that replaces the target sequence at the double-stranded target DNA sequence relative to a guide RNA without the clamp segment. The clamp segment can allow for a reduced number of nucleotides in the primer binding site need to bind its genomic site and facilitate reverse transcription, which in turn enables design of a guide RNA that is shorter than conventional guide RNAs used for other gene editing methods. The clamp segment is described in International Publication No. WO2023215831, which is hereby incorporated herein by reference in its entirety.
[0230] In some cases, a guide RNA can complete the insertion of new genetic material that replaces the target sequence without another guide RNA when delivered to a cell together with a prime editor described herein. The guide RNA can complete the insertion of the new genetic material that replaces the target sequence with a second guide RNA that is a nicking guide RNA when delivered together with a prime editor described herein.
[0231] In some cases, a guide RNA comprises two or more guide RNAs. In some cases, the two or more guide RNAs comprise a first guide RNA encoding at least a first portion of new genetic material that replaces the target sequence. In some cases, the two or more guide RNAs comprise a second guide RNA encoding at least a second portion of the new genetic material that replaces the target sequence. In some cases, the first guide RNA and the second guide RNA work in a pair and collectively encode the new genetic material that replaces the target sequence, thereby inserting the new genetic material that replaces the target sequence into the genome of a cell receiving the lipid delivery particles in a site-specific manner. In some cases, the first and the second portion of the new genetic material that replaces the target sequence have at least 6bp overlap. In some cases, the first portion of the new genetic material that replaces the target sequence is 46 bp. In some cases, the first portion of the new genetic material that replaces the target sequence is 42 bp. In some cases, the first portion or the second portion of the new genetic material that replaces the target sequence is 36 bp, 38 bp, 40 bp, 42 bp, 44 bp, or 46 bp. The first guide RNA comprises a first spacer. The second guide RNA comprises a second spacer. The first spacer
and the second spacer bind to two genomic target sites that are within 5-100 bp from each other. When the two or more guide RNAs are delivered to a cell together with a prime editor, the double strand DNA between the two genomic target sites are deleted and the full sequence of the new genetic material that replaces the target sequence is inserted instead. The deletion can be mediated by the following steps: (a) reverse transcription of the sequence encoding the first portion of the new genetic material that replaces the target sequence in the first guide RNA and the sequence encoding the second portion of the new genetic material that replaces the target sequence in the second guide RNA, wherein the first and the second portion of the new genetic material that replaces the target sequence having at least 6bp overlap,
(b) annealing of the two overlapped portion of the new genetic material that replaces the target sequence,
(c) synthesis of the second strand comprising the full sequence of the new genetic material that replaces the target sequence, (d) excision of the original DNA sequence, and (e) ligation of the pair nicks. The mechanism, process, and components of this process include those described in International Publication Nos. WO2023122764, W02023205710, and WO2023225670, each of which is hereby incorporated herein by reference in its entirety.
[0232] In some cases, the payload comprises a polypeptide domain described herein coupled to a polynucleotide domain described herein. In some cases, the payload comprises a polypeptide domain described herein complexed to a polynucleotide domain described herein. In some cases, the payload comprises a ribonucleoprotein. For example, the payload may comprise a guidable polypeptide domain complexed to a polynucleotide configured to bind to the guidable polypeptide domain (e.g., Cas9 complexed with an RNA guide).
[0233] Any of the payloads described herein can further comprise a plasma membrane recruitment element, a transmembrane domain, a signaling domain, a receptor domain, a packaging domain, or a targeting domain. Any of payloads described herein can comprise or be engineered to comprise a protein tag, a peptide tag, or small molecule tag. For example, a payload can comprise a nuclear localization signal (NLS), a nuclear export signal (NES), a cell penetrating peptide (CPP), a mitochondria penetrating peptide (MPP), a solubility tag, a fluorescent tag, or any combinations thereof.
Other Payloads
[0234] In some cases, the payload in the lipid delivery particle of the present disclosure comprises a recombinant protein. The payload can be a diagnostic imaging agent, such as a contrast agent. In some cases, the payload comprises a therapeutic agent, including, but not limited to, a nuclease, a recombinase, a growth factor, an antibody, a chimeric antigen receptor, a T cell receptor, a cytokine, a cytokine inhibitor or agonist, a transcription factor, an organelle, a nucleic acid molecule, a therapeutic DNA, a therapeutic RNA, a retrotransposon, a reverse transcriptase, an oligonucleotide, an aptazyme, an aptamer, or a ribozyme, a generic or specific kinase inhibitor, a small molecule drug, an immunomodulator, a tumor suppressor, a developmental regulator, a cancer vaccine, an anesthetic, an enzyme, a hormone, a ligand, a receptor, a T cell receptor, a transposon, a retrotransposon, a DNA polymerase, a RNA dependent DNA polymerase, a homing endonuclease, interferons, chemokines, insulin, growth factors, an antisense oligonucleotide, an RNAi, a shRNA, and any combination thereof. The payload can be a
prophylactic agent. In some cases, the payload comprises a biomarker. The payload can also comprise an exogenous antigen or an enzyme. In some cases, the payload comprises a metabolite molecule. In some cases, the payload comprises a lipid molecule. In some cases, the payload comprises a structural protein. In some cases, the payload comprises a hormone or a hormonal protein.
PRODUCTION OF LIPID DELIVERY PARTICLES
[0235] In some aspects, provided herein are composition, methods of production, methods of purification related to the lipid delivery particles provided herein. In some cases, the lipid delivery particles can be produced from producer cell lines that are either transiently transfected with at least one plasmid or stably expressing constructs that have been integrated into the producer cell line genomic DNA.
[0236] Producer cell lines can be generated by stably integrating genetic material with a gene of interest into a host cell line. In some cases, the genetic material is transiently expressed in a producer cell line. In some cases, the genetic material is expressed via viral methods. In some cases, the genetic material is expressed via non-viral methods. In some cases, a producer cell line grows in a serum-free medium or in suspension. A producer cell line can be grown in serum-free medium and suspension simultaneously. In some cases, producer cell lines can be generated with adherent cells (e.g., cells cultured in media and attached to a substrate).
[0237] Producer cells can be used to produce the lipid delivery particles described herein. In some cases, generating a producer cell line comprises transfecting cells (e.g., cells of a mammalian cell type) with genetic material of the present disclosure, culturing the cells to produce the lipid delivery particles, obtaining a media from the mammalian cell producing the lipid delivery particles, collecting and filtering the harvested media, and, optionally, purifying the lipid delivery particles to retain structural integrity. In some cases, the method of producing the lipid delivery particle further comprises providing new media to promote transient production of the lipid delivery particles. In some cases, the mammalian cell type includes a HT1080 cell, a COS cell, a HeLa cell, a Chinese Hamster Ovary (CHO) cell, or a HEK 293 cell. HEK293 cells are cells derived from human embryonic kidney cells grown in tissue culture. In some cases, the HEK293 cell is a HEK293, 293E, 293T, 293F, 293FT, or 293T Gesicle cell. The producer cell line can be transformed with a viral vector or non-viral method in any number of means including calcium phosphate and the like.
[0238] Following transfection, the cells can be cultured under conditions for production of lipid delivery particles. Exemplary culturing conditions can include refeeding cells in appropriate media, addition of CO2, and humidity. In some cases, culturing conditions includes addition of antibiotics, anti-fungals, and/or growth factors. The medium can be harvested after 24, 48, 72, or 96 hours, or at any appropriate time point to allow sufficient production of the lipid delivery particles.
[0239] Optionally, the lipid delivery particles in the media can be isolated and collected using any number of techniques known in the art. In some cases, the lipid delivery particles are purified, wherein the lipid delivery particles are washed or resuspended in an appropriate buffer or media or at particular concentration.
[0240] In an aspect, disclosed herein are methods of manufacturing producer cell lines that comprise the lipid delivery particles of the present disclosure. Adherent cells can be first transfected to produce lipid delivery particles. In some cases, transfection occurs by the addition or expression of exogenous nucleic acid sequences via non-viral methods (e.g., by electroporation, microinjection, or a chemical system such as DEAE-dextran or cationic polymers). In some cases, transfection occurs by the addition or expression of exogenous nucleic acid sequences via viral methods (e.g. , by infecting the cells with a viral vector, such as an adenoviral vector, adeno-associate viral vector, a lentiviral vector, a herpes viral vector, or a HSV vector). In some cases, the cells are from a HEK293 cell line (e.g., HEK293, 293E, or 293T). In some cases, to transfect DNA into the host cells, the cells are cultured in a medium. In some cases, cells can be cultured in the medium for 5, 10, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24 or 25 hours. In some cases, cells can be cultured in the medium for between 10-20 hours. In some cases, cells can be cultured in the medium for 18 hours.
[0241] Following incorporation into the transfection medium, cells are transferred to a new solution. In some cases, the new solution is new media. In some cases, the new media promotes the production of the lipid delivery particles. In some cases, the cells incorporate into the new media for between 10-50 hours. In some cases, the cells incorporate into the new media for 10, 20, 30, 35, 40, 45, or 50 hours. In some cases, the cells incorporate into the new media for 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39 or 40 hours. The media can then be harvested. The harvested media can be filtered, and the lipid delivery particles can be collected. Filtration can comprise microfiltration and/or depth filtration. In some cases, the lipid delivery particles can undergo further purification and/or concentration methods that maintain the structural integrity of the particles.
[0242] In some aspects, provided herein is a method of loading a lipid delivery particle with components such as a payload. RNA and protein from a producer cell can get packaged and/or incorporated into lipid delivery vehicles of the present disclosure. In some cases, the components of the lipid delivery particles, such as a payload, is loaded via the packaging and assembly process of the lipid delivery particle. For instance, the payload can be a polypeptide or protein that is packaged into the lipid delivery particle as a part of a chimeric protein as disclosed herein.
[0243] In some cases, the payload is assembled into the lipid delivery particle as an independent entity, e.g., not as a part of a chimeric protein. In other embodiments, the lipid delivery particle provided herein is loaded with a payload by utilizing any suitable method for delivering a biological or chemical payload through a lipid membrane, such as nucleofection, electroporation, lipid-based, polymer-based, or CaCF transfection, sonication, freeze thaw, incubation at various temperatures, or heat shock of lipid delivery particles mixed with payload. In some cases, the nucleic acid molecules, such as a template RNA described herein, are loaded into the lipid delivery particle by direct loading, such as electroporation of the lipid delivery particle in vitro. In some cases, the nucleic acid molecules are loaded into the lipid delivery particle by binding to a nucleic acid binding protein (e.g., Cas protein) that is part of the lipid delivery particle or is already loaded into the lipid delivery particle.
[0244] There can be more than one type of loading techniques utilized for loading payloads (e.g., for loading more than one type of payloads) into the lipid delivery particle. For instance, in some cases, a first payload is a polypeptide that is assembled into the lipid delivery particle as a part of a chimeric protein, and a second payload is a separate protein or nucleic acid (RNA or DNA) that interacts with (e.g., binds) the first payload, and thus is loaded into the lipid delivery particle via the interaction between the first payload and the second payload. Alternatively, the second payload can be loaded into the lipid delivery particle via a transfection-like technique or any other suitable method.
[0245] In aspects, also provided herein are methods of using a lipid delivery particle or pharmaceutical composition according to some embodiments of the present disclosure, comprising contacting a cell with the lipid delivery particle described herein. In some cases, the cell is a mammalian cell, such as a human cell. In some cases, the cell is within a subject in need of treatment for a disease or a condition. In some cases, contact comprising administering the lipid delivery particle described herein to the subject, such as via injections.
[0246] In aspects, also provided herein are methods of administering a lipid delivery particle, systems, or pharmaceutical compositions according to some embodiments of the present disclosure. In some cases, the method comprises administering the lipid delivery particle, system, or pharmaceutical composition described herein to a subject in need thereof, such as via injections.
[0247] In aspects, also provided herein are methods of producing a lipid delivery particle or pharmaceutical composition according to some embodiments of the present disclosure. In some cases, the method comprises contacting a producer cell with compositions described herein.
Methods of purification
[0248] In an aspect, described herein are methods of purifying lipid delivery particles. In some cases, the lipid delivery particles are produced from producer cell lines that are either transiently transfected with at least one plasmid or stably expressing constructs that have been integrated into the producer cell line genomic DNA. In some cases, the producer cell culture medium is harvested 24-, 48-, 72-, or 96-hours post-transfection. In some cases, the producer cell culture medium is harvested between 40- and 48-hours post-transfection. The harvested medium can undergo centrifugation steps to remove producer cell debris while maintaining the structural integrity of the lipid delivery particle. In some cases, during harvesting, the producer cell medium is centrifuged, e.g., at 500g for 5 minutes. The clarified lipid delivery particle containing supernatant can then be collected and filtered. In some cases, the lipid delivery particles are further concentrated. In some cases, the lipid delivery particles are further concentrated by ultracentrifugation. In some cases, the lipid delivery particles are concentrated 50-fold, 100-fold, 200- fold, 500-fold, 1000-fold, 2000-fold, 3000-fold, or 5000-fold. In some cases, the concentrated lipid delivery particles are resuspended, e.g., in cold PBS. In some cases, the concentrated lipid delivery particles are frozen, e.g., frozen at a rate of -l°C/min and stored at -80°C.
[0249] In some cases, the purification methods can comprise chromatographic methods (e.g., anion exchange chromatography), ultrafiltration methods (e.g, tangential flow filtration), clarifying normal flow filtration, and/or sterilizing membrane filtration. Anion exchange chromatography can separate
substances based on net-surface charge, using an ion-exchange resin. Tangential flow filtration can separate molecules using ultrafiltration membranes. In some cases, the membrane pore size used for tangential flow filtration can retain a biological product of a size less than 1000 kDa, less than 750 kDa, less than 500 kDa, less than 250 kDa, less than 200 kDa, less than 150 kDa, less than 100 kDa, or less than 50 kDa. Normal flow filtration assists in the clarification of biofluid by convecting the substance directly toward a membrane under an applied pressure. In some cases, normal flow filtration can comprise a membrane pore size of greater than 0.1 pm, greater than 0.2 pm, greater than 0.3 pm, greater than 0.4 pm, greater than 0.5 pm, greater than 0.6 pm, greater than 0.7 pm, greater than 0.8 pm, greater than 0.9 pm, greater than 1.0 pm, greater than 1.5 pm, or greater than 2.0 pm. In some cases, normal flow filtration can comprise a membrane pore size of 0.2 pm, 0.45 pm, 0.8 pm, 1.2 pm, or 2.0 pm. Sterilizing membrane filtration can be used to sterilize heat-sensitive liquid without exposure to denaturing hear. In some cases, sterilizing membrane filtration can comprise a membrane pore size of about 0.1 pm, about 0.2 pm, about 0.3 pm, about 0.4 pm, or about 0.5 pm. In some cases, sterilizing membrane filtration can comprise a membrane pore size of about 0.2 pm or 0.22 pm.
COMPOSITIONS AND SYSTEMS
[0250] In aspects, also provided herein are nucleic acid molecules that encode one or more of the components of the lipid delivery particles of the present disclosure. For instance, a nucleic acid molecule encoding the chimeric protein is provided. A nucleic acid molecule encoding the envelope protein is also provided.
[0251] In aspects, provided herein are compositions or systems that include nucleic acid molecules that encode one or more of the components of the lipid delivery particles of the present disclosure. A composition can comprise a first nucleic acid sequence encoding a chimeric protein comprising a Pleckstrin Homology domain comprising at least 60% at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99. 1%, at least 99.2%, at least 99.3%, at least 99.4%, at least 99.5%, at least 99.6%, at least 99.7%, at least 99.8%, or at least 99.9% sequence identity to the sequence of SEQ ID NO: 569; and at least two selected from the group of amino acid substitutions consisting of E17K, R25C, T81Y, T101C, and a combination of K142A, Hl 43 A, R144A (142-144A) relative to the sequence of SEQ ID NO: 569; and a protein payload.
[0252] A composition can comprise a first nucleic acid sequence encoding a chimeric protein comprising a Pleckstrin Homology domain comprising at least 60% at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99. 1%, at least 99.2%, at least 99.3%, at least 99.4%, at least 99.5%, at least 99.6%, at least 99.7%, at least 99.8%, or at least 99.9% sequence identity to SEQ ID NO: 613; and at least one amino acid substitution selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K relative to SEQ ID NO: 613; and a protein payload.
[0253] The compositions or systems can be used for producing a lipid delivery particle of the present disclosure, for instance, by transfecting or otherwise delivering the nucleic acid molecules in the
compositions or systems into a producer cell. The nucleic acid molecules can be expressed in the producer cell, the result of which assemble, package, and subsequently cause the producer cell to release the lipid delivery particle.
[0254] In some cases, a lipid delivery particle of the present disclosure facilitates gene editing efficiency greater than 40%, greater than 50%, greater than 60%, or more. In some cases, a lipid delivery particle of the present disclosure facilitates gene editing efficiency greater than 70%. In some cases, a lipid delivery particle of the present disclosure facilitates gene editing efficiency comprising 8-fold increase of base editing efficiency when compared to a conventional VLP (e.g. , the VLPs described in Mangeot, P. E. et al. Genome editing in primary cells and in vivo using viral-derived Nanoblades loaded with Cas9-sgRNA ribonucleoproteins. Nat. Commun. 10, 45 (2019).). In some cases, a lipid delivery particle of the present disclosure facilitates gene editing efficiency comprising 8-fold increase of prime editing efficiency when compared to a conventional VLP. In some cases, a lipid delivery particle of the present disclosure exhibits reduced immunogenicity in transduced target cells. In some cases, a lipid delivery particle of the present disclosure produces reduced off-target genome editing in target cells when delivering genome editing system into the target cells when compared to a conventional VLP. In some cases, a lipid delivery particle of the present disclosure leads to more than 100-fold reduction in Cas-independent off-target editing when compared to a conventional VLP. In some cases, a lipid delivery particle of the present disclosure leads to at least 10-fold, such as 12- to 900-fold, lower Cas-dependent off-target editing when compared to a conventional VLP.
PHARMACEUTICAL COMPOSITION
[0255] In an aspect, disclosed herein is a pharmaceutical formulation comprising the lipid delivery particle disclosed herein and optionally further comprising a pharmaceutically acceptable carrier, excipient, or additive. The term “pharmaceutical formulation”, as used herein, refers to a composition formulated for pharmaceutical use. The terms such as “excipient,” “carrier,” “pharmaceutically acceptable carrier” or the like are used interchangeably herein. Pharmaceutical formulations comprise an immunologically effective amount of one or more cells, vectors, lipid delivery particles, or compositions disclosed herein, and optionally one or more other components which are pharmaceutically acceptable. In some cases, the pharmaceutical formulation comprises additional agents, e.g., for specific delivery, increasing half-life, or other therapeutic benefit. In some cases, the pharmaceutical formulation may comprise one or more of dimethylsulfoxide (DMSO), dextrose, water, succinate, poly I: poly C, poly-L- lysine, carboxymethylcellulose, and/or chloride.
[0256] As used herein, a “pharmaceutically acceptable carrier” is an agent that is compatible with the other ingredients of the formulation and not injurious to the tissue of the subject (e.g., physiologically compatible, sterile, physiologic pH, etc.) In some cases, a pharmaceutically acceptable carrier comprises any vehicle, such as a liquid or solid fdler, diluent, excipient, manufacturing aid (e.g., lubricant, talc magnesium, calcium or zinc stearate, or steric acid), or solvent encapsulating material, involved in carrying or transporting the compound from one site (e.g. , the delivery site) of the body, to another site (e.g., organ, tissue or portion of the body).
[0257] Some exemplary materials which can serve as pharmaceutically-acceptable carriers include: (1) sugars, such as lactose, glucose and sucrose; (2) starches, such as com starch and potato starch; (3) cellulose, and its derivatives, such as sodium carboxymethyl cellulose, methylcellulose, ethyl cellulose, microcrystalline cellulose and cellulose acetate; (4) powdered tragacanthin; (5) malt; (6) gelatin; (7) lubricating agents, such as magnesium stearate, sodium lauryl sulfate and talc; (8) excipients, such as cocoa butter and suppository waxes; (9) oils, such as peanut oil, cottonseed oil, safflower oil, sesame oil, olive oil, com oil and soybean oil; (10) glycols, such as propylene glycol; (11) polyols, such as glycerin, sorbitol, mannitol and polyethylene glycol (PEG); (12) esters, such as ethyl oleate and ethyl laurate; (13) agar; (14) buffering agents, such as magnesium hydroxide and aluminum hydroxide; (15) alginic acid; (16) pyrogen-free water; (17) isotonic saline; (18) Ringer's solution; (19) ethyl alcohol; (20) pH buffered solutions; (21) polyesters, polycarbonates and/or polyanhydrides; (22) bulking agents, such as polypeptides and amino acids; (23) serum alcohols, such as ethanol; and (24) other non-toxic compatible substances employed in pharmaceutical formulations. Wetting agents, coloring agents, release agents, coating agents, sweetening agents, flavoring agents, perfuming agents, preservative and antioxidants can also be present in the formulation.
[0258] Pharmaceutical formulation disclosed herein can comprise one or more pH buffering compounds to maintain the pH of the formulation at a predetermined level that reflects physiological pH, such as in the range of about 5.0 to about 8.0. The pH of the pharmaceutical formulation can be about 4, about 5, about 6, about 7, about 8 or about 9. The pH buffering compound used in the aqueous liquid formulation can be an amino acid or mixture of amino acids, such as histidine or a mixture of amino acids such as histidine and glycine. The pH buffering compound can be an agent which does not chelate calcium ions. Exemplary pH buffering compounds include imidazole and acetate ions. The pH buffering compound can be present in any amount suitable to maintain the pH of the formulation at a predetermined level.
[0259] The pharmaceutical formulations described herein can be prepared by any method known or hereafter developed in the art of pharmacology. In general, such preparatory methods include the step of bringing the active ingredient(s) into association with an excipient and/or one or more other accessory ingredients, and then, optionally, shaping and/or packaging the product into a desired single- or multidose unit. Pharmaceutical formulations can additionally comprise a pharmaceutically acceptable excipient, which, as used herein, includes any and all solvents, dispersion media, diluents, or other liquid vehicles, dispersion or suspension aids, surface active agents, isotonic agents, thickening or emulsifying agents, preservatives, solid binders, lubricants, and the like, as suited to the particular dosage form desired.
METHOD OF TREATMENT, PREVENTION, OR DIAGNOSIS
[0260] A lipid delivery particle provided herein can find use in a variety of fields and methods. In some cases, the lipid delivery particle of the present disclosure can be used to deliver one or more payloads, such as a ribonucleoprotein complex to a cell. In some cases, the target cells to which the lipid delivery particles are delivered are in vitro cells, ex vivo cells, or in vivo cells. The lipid delivery particles of the
present disclosure can be applicable for delivery of freights into a variety of cell types, such as, animal cells, plant cells, bacteria cells, algal cells, or fungal cells.
[0261] In aspects, also provided herein are methods of treating a subject by administering a lipid delivery particle described herein, a system described herein, a composition described herein, or pharmaceutical composition according to some embodiments of the present disclosure.
[0262] In some cases, the present disclosure provides methods of treating, preventing, or diagnosing a condition, disease, or disorder. In some cases, a composition, kit, or method described herein can be used to treat, prevent, or diagnose a condition, disease, or disorder. The condition, disease, or disorder can comprise a cancer, an immune disorder, an autoimmune disorder, a metabolic disorder, a hormonal disorder, an inflammatory disorder, a developmental disorder, a reproductive disorder, an imprinting disorder, a genetic disorder, a neurological disorder, or a neurodegenerative disorder. In some cases, the condition, disease, or disorder comprises a liver disorder, an eye disorder, a heart disorder, a kidney disorder, a skin disorder, a blood disorder, a fibrotic disorder, a skeletal disorder, or a muscle order. In some cases, the condition, disease, or disorder is caused by a genetic mutation (e.g, an insertion, deletion, or point mutation). In some cases, the condition, disease, or disorder is hereditary. In some cases, the condition, disease, or disorder is caused by a virus or bacteria or fungus. In some cases, the condition, disease, or disorder is caused by aberrant gene expression. In some cases, the condition, disease, or disorder is a result of age. In some embodiments, the condition, disease, or disorder is chronic.
[0263] The subject in the method of present disclosure can be an animal. In some embodiments, the subject is an animal cell. In some embodiments, the subject is a mammal. In some embodiments, the subject is a human. In some embodiments, the subject is an aquaculture animal (fish, crabs, shrimp, oysters etc.), a mammal. In some embodiments, the animals cell is from, for example, a pet or zoo animal (cats, dogs, lizards, birds (e.g., parrots), lions, tigers and bears etc.), from a farm or working animal (horses, cows (e.g., dairy and beef cattle) pigs, chickens, turkeys, hens or roosters, goats, sheep, etc.), or a human. In some embodiments, the target cell as disclosed herein is in a subject to whom the method of the present disclosure is applicable.
[0264] The methods described herein can be therapeutic or veterinary methods for treating a subject. In some embodiments, the methods described herein are used to treat a disease resulting from a nonfunctional, poorly functional, or poorly expressed protein or gene product, for instance, resulting from a genetic mutation in one or more cells of the subject. In some embodiments, the methods described herein are used to treat a genetic disease (e.g. , a mutation, a substitution, a deletion, an expansion, or a recombination), a monogenic disease, an inherited metabolic disease, a cancer, a neurodegenerative disease, a cardiovascular disease, a pulmonary disease, a renal disease, a liver disease, a genetic disease, a vascular disease, ophthalmic disease, musculoskeletal disease, lymphatic disease, auditory and inner ear disease, a metabolic disease, an inflammatory disease, an autoimmune disease, or an infectious disease. In some cases, provided herein are pharmaceutical compositions and methods for treating a retinal disease, e.g., Leber congenital amaurosis, by administering a pharmaceutical composition formulated for subretinal injection.
KIT
[0265] In aspects, also provided herein are kits comprising the unit doses containing the lipid delivery particles, systems, compositions or pharmaceutical compositions of the present disclosure. In some embodiments, the kit comprises the lipid delivery particles, compositions, or pharmaceutical formulations of the present disclosure; and an informational medium containing instructions for administering the lipid delivery particle, composition, or pharmaceutical formulation to a subject. The kit can include a label indicating the intended use of lipid delivery particle, composition, or pharmaceutical formulation in the kit. Label can include any writing, or recorded material supplied on or with the kit, or which otherwise accompanies the kit.
[0266] A kit of the present disclosure can include, alternatively or additionally, diagnostic agents and/or other therapeutic agents. In some cases, the kit includes cells or pharmaceutical formulations of the present disclosure and a diagnostic agent that can be used in a diagnostic method for diagnosing a condition, disease, or disorder in a subject.
METHODS OF ADMINISTERING
[0267] In some cases, the composition or pharmaceutical formulation described herein is prepared for administration to a subject. In some cases, the pharmaceutical formulation is prepared to induce a therapeutic or prophylactic effect in a subject. Suitable routes of administrating the pharmaceutical formulation described herein include transdermal, intravesical, intravenous, intravascular, intraosseus, topical, subcutaneous, intradermal, intralesional, intraarticular, intraperitoneal, transmucosal, gingival, intradental, intracochlear, transtympanic, intraorgan, epidural, intrathecal, intramuscular, periocular, intratumoral, intracerebral, intravitreal, and intracerebroventricular administration. In some cases, the pharmaceutical formulation described herein is administered locally to a diseased site (e.g., site of infection or tumor site). In some cases, the pharmaceutical composition described herein is delivered in a controlled release system. In some cases, a pump is used. In some cases, polymeric materials is used for controlled release. In some cases, the pharmaceutical composition described herein is administered to a subject by injection, by means of a catheter, by means of a suppository, or by means of an implant, the implant being of a porous, non-porous, or gelatinous material, including a membrane, such as a sialastic membrane, or a fiber. In some cases, the pharmaceutical formulation is formulated in accordance with routine procedures as a formulation adapted for intravenous or subcutaneous administration to a subject. In some cases, pharmaceutical formulations for administration by injection are solutions in sterile isotonic use as solubilizing agent and a local anesthetic such as lignocaine to ease pain at the site of the injection. Generally, the ingredients can be supplied either separately or mixed together in unit dosage form, for example, as a dry lyophilized powder or water free concentrate in a hermetically sealed container such as an ampoule or sachets indicating the quantity of active agent. In some cases, if the pharmaceutical is to be administered by infusion, it is dispensed with an infusion bottle containing sterile pharmaceutical grade water or saline. In some cases, if the pharmaceutical formulation is administered by injection, an ampoule of sterile water for injection or saline is provided so that the ingredients can be mixed prior to administration.
[0268] A pharmaceutical formulation as described herein can be administered or packaged as a unit dose, for example, in reference to a pharmaceutical formulation to physically discrete units suitable as unitary dosage for the subject, each unit containing a predetermined quantity of active material calculated to produce the desired therapeutic effect in association with the required diluent, carrier, or vehicle.
EXAMPLES
[0269] The following examples are provided to further illustrate some embodiments of the present disclosure but are not intended to limit the scope of the disclosure; it will be understood by their exemplary nature that other procedures, methodologies, or techniques known to those skilled in the art can alternatively be used.
Example 1: Screens to identify Pleckstin Homology domains with specific properties
[0270] A screen of Pleckstin Homology domains will be performed to identify mutations that confer a PH domain with enhanced or altered properties. To generate the PH domain variants, mutations will be introduced through mutagenesis and/or gene editing-based methods. Unedited PH domains from alternative sources will also be screened. Non-limiting examples of enhanced or altered properties that will be measured include recruitment of a payload into the lipid delivery particle, the amount of a payload that can be encapsulated in the lipid delivery particle, release of a payload from the lipid delivery particle into a target cell, trafficking of the lipid delivery particle or payload protein to a subcellular locale in a target cell, or suitability of the PH domain for combination with other proteins, protein domains, or protein components.
[0271] To identify mutations that enhance or alter recruitment of a payload into the lipid delivery particle, the lipid delivery particle pseudo typed with VSVg or other envelope proteins can be formed with different PH domains. A single PH domain will be fused to a reporter protein, such as green fluorescent protein (GFP), or another enzyme with measurable functional activity such as Cas9 or ABE8e. The PH domain-GFP and/or PH domain-Cas9 chimeric proteins will be packaged into lipid delivery particles. A range of cell lines, including HEK293T, HeLa, HepG2, ARPE-19, BEAS-2B, induced pluripotent stem cells (iPSC), or SH-SYSY cells, will be treated with the packaged lipid delivery particles that were previously purified. The purification and concentration of the lipid delivery particle will be performed by filtration methods, such as PVDF filtration, and ultracentrifugation. Recruitment of a payload into the lipid delivery particle using different PH domains will be measured by flow cytometry, microscopy, or sequencing.
[0272] To identify mutations that enhance or alter the amount of a payload that can be encapsulated into the lipid delivery particle using different PH domains, a single PH domain will be fused to a reporter protein, such as GFP, or another enzyme with measurable functional activity such as Cas9 or ABE8e. The PH domain-GFP and/or PH domain-Cas9 chimeric proteins will be packaged into lipid delivery particles. A range of cell lines, including HEK293T, HeLa, HepG2, ARPE-19, BEAS-2B, or SH-SYSY cells, will be treated with the packaged lipid delivery particles that were previously purified. The purification and concentration of the lipid delivery particle will be performed by filtration methods, such as PVDF
filtration, and ultracentrifugation. The amount of a payload that can be encapsulated into the lipid delivery particle using different PH domains will be measured and/or quantified by flow cytometry, microscopy, or sequencing.
[0273] To identify mutations that enhance or alter release of a payload into a target cell using different PH domains, a single PH domain will be fused to a reporter protein, such as GFP, or another enzyme with measurable functional activity such as Cas9 or ABE8e. The PH domain-GFP and/or PH domain-Cas9 chimeric proteins will be packaged into lipid delivery particles. A range of cell lines, including HEK293T, He La, HepG2, ARPE-19, BEAS-2B, or SH-SYSY cells, will be treated with the packaged lipid delivery particles that were previously purified. The purification and concentration of the lipid delivery particle will be performed by filtration methods, such as PVDF filtration, and ultracentrifiigation. The amount of a payload that can be released into the cell via the lipid delivery particle using different PH domains will be measured and/or quantified by flow cytometry, microscopy, or sequencing.
[0274] To identify mutations that enhance or alter trafficking of a payload protein to a subcellular locale in a target cell using different PH domains, a single PH domain will be fused to a reporter protein, such as GFP, or another enzyme with measurable functional activity such as Cas9 or ABE8e. The PH domain- GFP and/or PH domain-Cas9 chimeric proteins will be packaged into lipid delivery particles. A range of cell lines, including HEK293T, HeLa, HepG2, ARPE-19, BEAS-2B, or SH-SYSY cells, will be treated with the packaged lipid delivery particles that were previously purified. The purification and concentration of the lipid delivery particle will be performed by filtration methods, such as PVDF filtration, and ultracentrifugation. The amount of a payload protein that can be trafficked into a subcellular locale using different PH domains will be measured and/or quantified by flow cytometry, microscopy, or sequencing.
[0275] To identify mutations that enhance or alter suitability of an PH domain for combination with other protein domains using different PH domains, a single PH domain will be fused to a reporter protein, such as GFP, another enzyme with measurable functional activity such as Cas9 or ABE8e, or a range of other protein domains. The PH domain-GFP and/or PH domain-Cas9 chimeric proteins will be packaged into lipid delivery particles. A range of cell lines, including HEK293T, HeLa, HepG2, ARPE-19, BEAS- 2B, or SH-SYSY cells, will be treated with the packaged lipid delivery particles that were previously purified. The purification and concentration of the lipid delivery particle will be performed by filtration methods, such as PVDF filtration, and ultracentrifugation. The suitability of an PH domain for combination with other protein domains using different PH domains will be measured and/or quantified by flow cytometry, microscopy, sequencing, a combination thereof, or using another method.
Example 2: Gene editing efficiency using different PH domains
[0276] Fifty PH domains were tested for gene editing capacity as determined by the percentage of sequencing reads expressing edits (any derivation from the wildtype sequence at a specific locus was counted as an editing event). Lipid delivery particles were generated and delivered to two cell lines, HEK293 (FIGs. 2A and 2B) and K562 (FIGs. 2C and 2D) at a high and low dose. Different PH domain- Cas9 chimeric proteins were generated and encapsulated as payloads in the lipid delivery particle for
delivery into HEK293 and K562 cells. Particles were concentrated lOOx by PEG precipitation prior to transduction of recipient cells. The lipid delivery vehicles were introduced to the cells at a high dose and a low dose.
[0277] A Cas9 chimeric protein lacking a PH domain (No PH) and an untreated sample (NT) was included as a negative control. The lipid delivery particles were transduced. The cells were lysed, genomic DNA was extracted, amplicons inducing the target site were generated, and the efficiency of the Cas9 fused to different PH domains was assessed through quantification of edits installed at the target editing site. The results are summarized in FIGs. 2A-2D.
[0278] The treated and control samples were harvested for genomic DNA using Qiagen Blood & Cell Culture DNA kit. The DNA was used as a template for PCR, wherein HEXA target specific primers flank the insertion locus. PCR was performed using a high fidelity, proof-reading Taq polymerase (Prime Star, Takara) to isolate the target region of interest. The PCR inserts was purified and prepared for a next generation sequencing run using a library preparation kit (Nextera XT, Illumina). The library was run on the Illumina MiSeq and editing efficiency of each guide RNA and the PH domains was determined (FIGs. 2A-2D).
Example 3: Generation of a lipid delivery particle comprising a Pleckstin Homology domain, an envelope, and gene editing machinery
[0279] A nucleic acid molecule will be generated that encodes a chimeric protein that comprises one of the PH domains disclosed herein and a payload protein. In this example, the nucleic acid construct will encode the PH domain AKT1 E17K/R25C (SEQ ID NO. 575). Another nucleic acid molecule that will be generated encodes an envelope and target specific guide RNA. In this example, the payload protein comprises a Cas domain. The nucleic acid molecules encoding the chimeric protein, the envelope, and the target specific guide RNA are introduced into HEK293T producer cells to generate the lipid delivery particles comprising protein payload.
Example 4. Delivery of a lipid delivery particle comprising a Pleckstin Homology domain, an envelope, and gene editing machinery to a target cell
[0280] The lipid delivery particle comprising the protein payload generated in Example 2 will be introduced into a target cell. In this example, the target cell will be an induced pluripotent stem cell derived neurons model (iPSC-N). The protein payload will comprise a guide RNA that targets alpha subunit of the enzyme [3-hexosaminidase (HEXA) located on human chromosome 15. Multiple guide RNAs will be designed to target HEXA in exon 11, the locus that encodes a pathogenic 4 base pair insertion. The guide RNAs will be analyzed for specificity, efficiency, secondary structure, and potential to induce off-target mutations. The PH domains SEQ ID NO: 569-596 will be modified for delivery to neurons and efficiency of the gene editing protein payload.
[0281] To determine the efficiency of the lipid delivery particle on correcting the pathogenic mutation in cardiomyocytes, nucleic acid molecules encoding the chimeric protein, the envelope, and the target specific guide RNA will be introduced into HEK293T producer cells. The lipid delivery particles will be isolated and purified using filtration methods and centrifugation. The iPSC-Ns expressing the 4 base pair
insertion mutation will be treated with the lipid delivery particle and cultured for 48-72 hours. A control iPSC-Ns expressing the 4 bae pair insertion will be treated with the lipid delivery particle comprising a scramble guide RNA particle and cultured for 48-72 hours. The treated and control iPSC-Ns are harvested for genomic DNA using Qiagen Blood & Cell Culture DNA kit. The DNA will be used as a template for PCR, wherein HEXA target specific primers flank the insertion site. PCR will be performed using a high fidelity, proof-reading Taq polymerase (Prime Star, Takara) to isolate the target region of interest. The PCR inserts will be purified and prepared for a next generation sequencing run using a library preparation kit (Nextera XT, Illumina). The library will then be run on the Illumina MiSeq and editing efficiency of each guide RNA and the PH domains will be determined.
Example 5. Delivery of a lipid delivery particle to a subject with genetic disease
[0282] The guide RNA and PH domain will then be provided to a subject with familial hypertrophic cardiomyopathy caused by a pathogenic insertion in HEXA. The lipid delivery particle will be provided to the subject intravenously or through cerebral spinal fluid. The gene editing payload protein will be delivered to the target neurons, where the guide RNA delivers the gene editing machinery to exon 11 of HEXA, and removes the pathogenic insertion. A sample from the subject will be taken after treatment to assess if the genome alteration is sufficient to restore normal function of the gene or to block pathogenic function of the pathogenic gene. A control sample from the subject will be taken from the subject prior to treatment.
[0283] The treated and control samples will be harvested for genomic DNA using Qiagen Blood & Cell Culture DNA kit. The DNA will be used as a template for PCR, wherein HEXA target specific primers flank the insertion locus. PCR will be performed using a high fidelity, proof-reading Taq polymerase (Prime Star, Takara) to isolate the target region of interest. The PCR inserts will be purified and prepared for a next generation sequencing run using a library preparation kit (Nextera XT, Illumina). The library will then be run on the Illumina MiSeq and editing efficiency of each guide RNA and the PH domains will be determined.
Example 6. Plextrin Homology Domain Screen Improves Delivery of Gene Editing Machinery [0284] While preferred embodiments of the present disclosure have been shown and described herein, it will be obvious to those skilled in the art that such embodiments are provided by way of example only. Numerous variations, changes, and substitutions will now occur to those skilled in the art without departing from the disclosure. It should be understood that various alternatives to the embodiments of the present disclosure may be employed in practicing the present disclosure. It is intended that the following claims define the scope of the present disclosure and that methods and structures within the scope of these claims and their equivalents be covered thereby.
[0285] While preferred embodiments of the present disclosure have been shown and described herein, it will be obvious to those skilled in the art that such embodiments are provided by way of example only. Numerous variations, changes, and substitutions will now occur to those skilled in the art without departing from the disclosure. It should be understood that various alternatives to the embodiments of the present disclosure may be employed in practicing the present disclosure. It is intended that the following
claims define the scope of the present disclosure and that methods and structures within the scope of these claims and their equivalents be covered thereby.
Claims
1. A chimeric protein comprising:
(a) a Pleckstrin Homology domain comprising:
(i) at least 60% at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99.1%, at least 99.2%, at least 99.3%, at least 99.4%, at least 99.5%, at least 99.6%, at least 99.7%, at least 99.8%, or at least 99.9% sequence identity to the sequence of SEQ ID NO: 569; and
(ii) at least two selected from the group of amino acid substitutions consisting of E17K, R25C, T81Y, T101C, and a combination of K142A, H143A, R144A (142-144A) relative to the sequence of SEQ ID NO: 569; and
(b) a protein payload.
2. The chimeric protein of claim 1, wherein the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K and R25C; E17K and T81Y; E17K and T101C; E17K and 142-144A; R25C and T81Y; R25C and T101C; R25C and 142-144A; T81Y and T101C; T81Y and 142- 144A; and T101C and 142-144A.
3. The chimeric protein of claim 2, wherein the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 572, 573, 574, 575, 576, 577, 578, 579, or 580.
4. The chimeric protein of claim 1, wherein the chimeric protein comprises at least three mutations selected from the group consisting of E17K, R25C, T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569.
5. The chimeric protein of claim 4, wherein the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K, R25C, and T81Y; E17K, R25C, and T101C; E17K, R25C, and 142-144A; E17K, T81Y, and T101C; E17K, T81Y, and 142-144A; E17K, T101C, and 142-144A; R25C, T81Y, and T101C; R25C, T81Y, and 142-144A; R25C, T101C, and 142-144A; and T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569.
6. The chimeric protein of claim 5, wherein the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 581, 582, 583, 584, 585,586, 587, 588, 589, or 590.
7. The chimeric protein of claim 1, wherein the chimeric protein comprises at least four selected from the group consisting of E17K, R25C, T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569.
8. The chimeric protein of claim 7, wherein the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K, R25C, T81Y, and T101C; E17K, R25C, T81Y, and 142- 144A; R25C, T81Y, T101C, and 142-144A; R25C, T81Y, T101C, and 142-144A; and E17K, T81Y, T101C, and 142-144A, relative to the sequence of SEQ ID NO: 569.
9. The chimeric protein of claim 8, wherein the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 591, 592, 593, 594, or 595.
10. The chimeric protein of claim 1, wherein the chimeric protein comprises at least the mutations E17K, R25C, T81Y, TIOIC, and 142-144A, relative to the sequence of SEQ ID NO: 569.
11. The chimeric protein of claim 10, wherein the chimeric protein comprises the sequence of SEQ ID NO: 596.
12. The chimeric protein of any one of claims 1-11, further comprising one or more nuclear export sequences.
13. The chimeric protein of claim 12, wherein the one or more nuclear export sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 5.
14. The chimeric protein of any one of claims 1-13, further comprising one or more nuclear localization sequences.
15. The chimeric protein of claim 14, wherein the one or more nuclear localization sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 6.
16. The chimeric protein of any one of claims 1-15, wherein the Pleckstrin Homology domain is coupled to the protein payload.
17. The chimeric protein of claim 16, wherein the Pleckstrin Homology domain is reversibly coupled to the protein payload.
18. The chimeric protein of claim 17, wherein the protein payload is reversibly coupled to the Pleckstrin Homology domain by a cleavable linker.
19. The chimeric protein of claim 18, wherein the cleavable linker is cleavable by a protease.
20. The chimeric protein of any one of claims 16-19, wherein the protein payload is coupled to a C- terminal of the Pleckstrin Homology domain.
21. The chimeric protein of any one of claims 16-19, wherein the protein payload is coupled to an N- terminal of the Pleckstrin Homology domain.
22. The chimeric protein of any one of claims 1-21, wherein the protein payload comprises one or more viral proteins.
23. The chimeric protein of claim 22, wherein the one or more viral proteins comprise one or more retroviral proteins.
24. The chimeric protein of claim 23, wherein the one or more retroviral proteins comprises a retroviral Gag protein.
25. The chimeric protein of any one of claims 1-21, wherein the protein payload comprises one or more non-viral proteins.
26. The chimeric protein of claim 25, wherein the one or more non-viral proteins comprises one or more mammalian proteins.
27. The chimeric protein of claim 26, wherein the one or more mammalian proteins comprises an Arc protein.
28. The chimeric protein of any one of claims 1-27, wherein the protein payload comprises a gene editing protein.
29. The chimeric protein of claim 28, wherein the gene editing protein comprises a prime editing protein.
30. The chimeric protein of claim 29, wherein the prime editing protein is coupled to a target-specific prime editing guide RNA.
31. The chimeric protein of any one of claims 28-30, wherein the gene editing protein comprises a CRISPR system.
32. The chimeric protein of claim 31, wherein the CRISPR system comprises a Cas domain.
33. The chimeric protein of any one of claims 28-32, wherein the gene editing protein is coupled to a target-specific guide RNA.
34. The chimeric protein of any one of claims 1-27, wherein the protein payload comprises an epigenetic editing protein.
35. The chimeric protein of claim 34, wherein the epigenetic editing protein is coupled to a target-specific guide RNA.
36. The chimeric protein of any one of claims 1-27, wherein the protein payload comprises a recombinase protein.
37. The chimeric protein of any one of claims 1-27, wherein the protein payload comprises an integrase protein.
38. A nucleic acid molecule encoding the chimeric protein of any one of claims 1-37.
39. A lipid delivery particle comprising the chimeric protein of any one of claims 1-37.
40. The lipid delivery particle of claim 39, further comprising an envelope, wherein the envelope comprises a lipid bilayer encasing a lumen, wherein the chimeric protein is located in the lumen.
41. The lipid delivery particle of claim 39 or claim 40, further comprising a target ligand.
42. The lipid delivery particle of claim 41, wherein the target ligand reversibly couples to a target receptor.
43. The lipid delivery particle of claim 42, wherein the target receptor comprises a cell surface protein on a target cell.
44. The lipid delivery particle of claim 43, wherein the envelope couples to the target cell at least partially through coupling of the target ligand and the target receptor on the target cell.
45. The lipid delivery particle of claim 44, wherein coupling of the target ligand and the target receptor on the target cell induces release of the protein payload in the target cell.
46. The lipid delivery particle of claim 45, wherein release of the protein payload in the target cell follows fusion of the envelope with cell membrane of the target cell.
47. A method of delivering a payload to a target cell, the method comprising: contacting the target cell with the lipid delivery particle of any one of claims 39-46.
48. A chimeric protein comprising:
(a) a Pleckstrin Homology domain comprising at least 60%, at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%,
at least 96%, at least 97%, at least 98%, at least 99%, at least 99.1%, at least 99.2%, at least 99.3%, at least 99.4%, at least 99.5%, at least 99.6%, at least 99.7%, at least 99.8%, or at least 99.9% sequence identity to any one of the sequences set forth in Table 4B; and
(b) a protein payload.
49. The chimeric protein of claim 48, wherein the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 597, 598, 599, 600, 601, 602, 603, 604, 605, 606, 607, 608, 609, 610, 611, 612, 613, 614, 615, 616, 617, 618, 619, 620, 621, 622, 623, 624, 625, 626, 627, 628, 629, 630, 631, 632, 633,
634, 635, 636, 637, 638, 639, 640, 641, 642, 643, 644, 645, 646, 647, 648, 649, 650, 651, 652, 653, 654,
655, 656, 657, 658, 659, 660, 661, 662, 663, 664, 665, 666, 667, 668, 669, 670, 671, 672, 673, 674, 675,
676, 677, 678, 679, 680, 681, 682, 683, 684, 685, 686, 687, 688, 689, 690, 691, 692, 693, 694, 695, 696,
697, 698, 699, 700, 701, 702, 703, 704, 705, 706, 707,708 ,709, 710, 711, 712, 713, 714, 715, 716, 717, 718, 719, 720, 721, or 722.
50. The chimeric protein of claim 48 or claim 49, further comprising one or more nuclear export sequences.
51. The chimeric protein of claim 50, wherein the one or more nuclear export sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 5.
52. The chimeric protein of any one of claims 48-51, further comprising one or more nuclear localization sequences.
53. The chimeric protein of claim 52, wherein the one or more nuclear localization sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 6.
54. The chimeric protein of any one of claims 48-53, wherein the Pleckstrin Homology domain is coupled to the protein payload.
55. The chimeric protein of claim 54, wherein the Pleckstrin Homology domain is reversibly coupled to the protein payload.
56. The chimeric protein of claim 55, wherein the protein payload is reversibly coupled to the Pleckstrin Homology domain by a cleavable linker.
57. The chimeric protein of claim 56, wherein the cleavable linker is cleavable by a protease.
58. The chimeric protein of any one of claims 54-57, wherein the protein payload is coupled to C-terminal of the Pleckstrin Homology domain.
59. The chimeric protein of any one of claims 54-57, wherein the protein payload is coupled to N- terminal of the Pleckstrin Homology domain.
60. The chimeric protein of any one of claims 48-59, wherein the protein payload comprises one or more viral proteins.
61. The chimeric protein of claim 60, wherein the one or more viral proteins comprise one or more retroviral proteins.
62. The chimeric protein of claim 61, wherein the one or more retroviral proteins comprises a retroviral Gag protein.
63. The chimeric protein of any one of claims 48-62, wherein the protein payload comprises one or more non-viral proteins.
64. The chimeric protein of claim 63, wherein the one or more non-viral proteins comprises one or more mammalian proteins.
65. The chimeric protein of claim 64, wherein the one or more mammalian proteins comprises an Arc protein.
66. The chimeric protein of any one of claims 48-65, wherein the protein payload comprises a gene editing protein.
67. The chimeric protein of claim 66, wherein the gene editing protein comprises a prime editing protein.
68. The chimeric protein of claim 67, wherein the prime editing protein is coupled to a target-specific prime editing guide RNA.
69. The chimeric protein of any one of claims 66-68, wherein the gene editing protein comprises a CRISPR system.
70. The chimeric protein of claim 69, wherein the CRISPR system comprises a Cas domain.
71. The chimeric protein of any one of claims 66-70, wherein the gene editing protein is coupled to a target-specific guide RNA.
72. The chimeric protein of any one of claims 48-65, wherein the protein payload comprises an epigenetic editing protein.
73. The chimeric protein of claim 72, wherein the epigenetic editing protein is coupled to a target-specific guide RNA.
74. The chimeric protein of any one of claims 48-65, wherein the protein payload comprises a recombinase protein.
75. The chimeric protein of any one of claims 48-65, wherein the protein payload comprises an integrase protein.
76. A nucleic acid molecule encoding the chimeric protein of any one of claims 48-75.
77. A lipid delivery particle comprising the chimeric protein of any one of claims 48-75.
78. The lipid delivery particle of claim 77, further comprising an envelope, wherein the envelope comprises a lipid bilayer encasing a lumen, wherein the chimeric protein is located in the lumen.
79. The lipid delivery particle of claim 77 or claim 78, further comprising a target ligand.
80. The lipid delivery particle of claim 79, wherein the target ligand reversibly couples to a target receptor.
81. The lipid delivery particle of claim 80, wherein the target receptor comprises a cell surface protein on a target cell.
82. The lipid delivery particle of claim 81, wherein the envelope couples to the target cell through coupling of the target ligand and the target receptor on the target cell.
83. The lipid delivery particle of claim 82, wherein coupling of the target ligand and the target receptor on the target cell induces release of the protein payload in the target cell.
84. The lipid delivery particle of claim 83, wherein release of the protein payload in the target cell follows endocytosis of the lipid delivery particle.
85. A method to deliver a payload to a target cell, the method comprising: delivering the lipid delivery particle of any one of claims 77-84 to a target cell.
86. A chimeric protein comprising:
(a) a Pleckstrin Homology domain comprising:
(i) at least 60% at least 65%, at least 70%, at least 75%, at least 80%, at least 85%, at least 90%, at least 91%, at least 92%, at least 93%, at least 94%, at least 95%, at least 96%, at least 97%, at least 98%, at least 99%, at least 99.1%, at least 99.2%, at least 99.3%, at least 99.4%, at least 99.5%, at least 99.6%, at least 99.7%, at least 99.8%, or at least 99.9% sequence identity to the sequence of SEQ ID NO: 613; and
(ii) at least one amino acid substitution selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K relative to the sequence of SEQ ID NO: 613; and
(b) a protein payload.
87. The chimeric protein of claim 86, wherein the chimeric protein comprises the sequence set forth in any one of SEQ ID NOs: 603, 604, 605, 606, or 607.
88. The chimeric protein of claim 86, wherein the chimeric protein comprises at least two mutations selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613.
89. The chimeric protein of claim 88, wherein the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K and E58K; E17K and E52K; E17K and E53K; E17K and E55K; E17K and E65K; E58K and E52K; E58K and E53K; E58K and E55K; E58K and E65K; E52K and E53K; E52K and E55K; E52K and E65K; E53K and E55K; and E53K and E65K, relative to the sequence of SEQ ID NO: 613.
90. The chimeric protein of claim 89, wherein the chimeric protein comprises the sequence of SEQ ID NO: 608.
91. The chimeric protein of claim 86, wherein the chimeric protein comprises at least three mutations selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613.
92. The chimeric protein of claim 91, wherein the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K, E58K, and E52K; E17K, E58K, and E53K; E17K, E58K, and E55K; E17K, E58K, and E65K; E17K, E52K, and E53K; E17K, E52K, and E55K; E17K, E52K, and E65K; E17K, E53K, and E55K; E17K, E53K, and E65K; E58K, E52K, and E53K; E58K, E52K, and E55K; E58K, E52K, and E65K; E58K, E53K, and E55K; E58K, E53K, and E65K; E52K, E53K, and E65K; and E52K, E53K, and E55K, relative to the sequence of SEQ ID NO: 613.
93. The chimeric protein of claim 92, wherein the chimeric protein comprises the sequence of SEQ ID NO: 609.
94. The chimeric protein of claim 86, wherein the chimeric protein comprises at least four mutations selected from the group consisting of E17K, E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613.
95. The chimeric protein of claim 94, wherein the chimeric protein comprises at least one mutation group selected from the group consisting of: E17K, E58K, E52K, and E53K; E17K, E58K, E52K, and E55K; E17K, E58K, E52K, and E65K; E58K, E52K, E53K, and E55K; E58K, E52K, E53K, and E65K; E58K, E52K, E53K, and E65K, relative to the sequence of SEQ ID NO: 613.
96. The chimeric protein of claim 95, wherein the chimeric protein comprises the sequence of SEQ ID NO: 610.
97. The chimeric protein of claim 86, wherein the chimeric protein comprises at least five selected from the group consisting of: E17K, E58K, E52K, E53K, and E55K; E17K, E58K, E52K, E53K, and E65K; E17K, E58K, E52K, E55K, and E65K; E17K, E58K, E53K, E55K, and E65K; E17K, E52K, E53K, E55K, and E65K; and E58K, E52K, E53K, E55K, and E65K, relative to the sequence of SEQ ID NO: 613.
98. The chimeric protein of claim 97, wherein the chimeric protein comprises the sequence of SEQ ID NO: 611.
99. The chimeric protein of claim 86, wherein the chimeric protein comprises the mutations E17K, E58K, E52K, E53K, E55K, and E65K , relative to the sequence of SEQ ID NO: 613.
100. The chimeric protein of claim 99, wherein the chimeric protein comprises the sequence of SEQ ID NO: 612.
101. The chimeric protein of any one of claims 86-100, further comprising one or more nuclear export sequences.
102. The chimeric protein of claim 101, wherein the one or more nuclear export sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 5.
103. The chimeric protein of any one of claims 86-102, further comprising one or more nuclear localization sequences.
104. The chimeric protein of claim 103, wherein the one or more nuclear localization sequences comprises about 80%, 85%, 90%, 91%, 92%, 93%, 94%, 95%, 96%, 97%, 98%, or 99% sequence identity to any one of the sequences set forth in Table 6.
105. The chimeric protein of any one of claims 86-104, wherein the Pleckstrin Homology domain is coupled to the protein payload.
106. The chimeric protein of claim 105, wherein the Pleckstrin Homology domain is reversibly coupled to the protein payload.
107. The chimeric protein of claim 106, wherein the protein payload is reversibly coupled to the Pleckstrin Homology domain by a cleavable linker.
108. The chimeric protein of claim 107, wherein the cleavable linker is cleavable by a protease.
109. The chimeric protein of any one of claims 105-108, wherein the protein payload is coupled to C- terminal of the Pleckstrin Homology domain.
110. The chimeric protein of any one of claims 105-108, wherein the protein payload is coupled to N- terminal of the Pleckstrin Homology domain.
111. The chimeric protein of any one of claims 86-110, wherein the protein payload comprises one or more viral proteins.
112. The chimeric protein of claim 111, wherein the one or more viral proteins comprise one or more retroviral proteins.
113. The chimeric protein of claim 112, wherein the one or more retroviral proteins comprises a retroviral Gag protein.
114. The chimeric protein of any one of claims 86-113, wherein the protein payload comprises one or more non-viral proteins.
115. The chimeric protein of claim 114, wherein the one or more non-viral proteins comprises one or more mammalian proteins.
116. The chimeric protein of claim 115, wherein the one or more mammalian proteins comprises an Arc protein.
117. The chimeric protein of any one of claims 86-116, wherein the protein payload comprises a gene editing protein.
118. The chimeric protein of claim 117, wherein the gene editing protein comprises a prime editing protein.
119. The chimeric protein of claim 118, wherein the prime editing protein is coupled to a target-specific prime editing guide RNA.
120. The chimeric protein of any one of claims 117-119, wherein the gene editing protein comprises a CRISPR system.
121. The chimeric protein of claim 120, wherein the CRISPR system comprises a Cas domain.
122. The chimeric protein of any one of claims 117-121, wherein the gene editing protein is coupled to a target-specific guide RNA.
123. The chimeric protein of any one of claims 86-116, wherein the protein payload comprises an epigenetic editing protein.
124. The chimeric protein of claim 123, wherein the epigenetic editing protein is coupled to a targetspecific guide RNA.
125. The chimeric protein of any one of claims 86-116, wherein the protein payload comprises a recombinase protein.
126. The chimeric protein of any one of claims 86-116, wherein the protein payload comprises an integrase protein.
127. A nucleic acid molecule encoding the chimeric protein of any one of claims 86-126.
128. A lipid delivery particle comprising the chimeric protein of any one of claims 86-126.
129. The lipid delivery particle of claim 128, further comprising an envelope, wherein the envelope comprises a lipid bilayer encasing a lumen, wherein the chimeric protein is located in the lumen.
130. The lipid delivery particle of claim 128 or claim 129, further comprising a target ligand.
131. The lipid delivery particle of claim 130, wherein the target ligand reversibly couples to a target receptor.
132. The lipid delivery particle of claim 131, wherein the target receptor comprises a cell surface protein on a target cell.
133. The lipid delivery particle of claim 132, wherein the envelope couples to the target cell through coupling of the target ligand and the target receptor on the target cell.
134. The lipid delivery particle of claim 133, wherein coupling of the target ligand and the target receptor on the target cell induces release of the protein payload in the target cell.
135. The lipid delivery particle of claim 134, wherein release of the protein payload in the target cell follows endocytosis of the lipid delivery particle.
136. A method to deliver a payload to a target cell, the method comprising: delivering the lipid delivery particle of any one of claims 128-135 to a target cell.
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| US202363506811P | 2023-06-07 | 2023-06-07 | |
| US63/506,811 | 2023-06-07 |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2024254530A1 true WO2024254530A1 (en) | 2024-12-12 |
Family
ID=93796419
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/US2024/033111 Ceased WO2024254530A1 (en) | 2023-06-07 | 2024-06-07 | Pleckstrin holomogy domains for delivery of protein payloads to a target cell |
Country Status (1)
| Country | Link |
|---|---|
| WO (1) | WO2024254530A1 (en) |
Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2015171543A1 (en) * | 2014-05-05 | 2015-11-12 | California Institute Of Technology | Mutant akt-specific capture agents, compositions, and methods of using and making |
| WO2017053629A2 (en) * | 2015-09-22 | 2017-03-30 | William Marsh Rice University | Light-controlled gene delivery with virus vectors through incorporation of optogenetic proteins and genetic insertion of non-conformationally constrained peptides |
| WO2022020800A2 (en) * | 2020-07-24 | 2022-01-27 | The General Hospital Corporation | Enhanced virus-like particles and methods of use thereof for delivery to cells |
-
2024
- 2024-06-07 WO PCT/US2024/033111 patent/WO2024254530A1/en not_active Ceased
Patent Citations (3)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| WO2015171543A1 (en) * | 2014-05-05 | 2015-11-12 | California Institute Of Technology | Mutant akt-specific capture agents, compositions, and methods of using and making |
| WO2017053629A2 (en) * | 2015-09-22 | 2017-03-30 | William Marsh Rice University | Light-controlled gene delivery with virus vectors through incorporation of optogenetic proteins and genetic insertion of non-conformationally constrained peptides |
| WO2022020800A2 (en) * | 2020-07-24 | 2022-01-27 | The General Hospital Corporation | Enhanced virus-like particles and methods of use thereof for delivery to cells |
Non-Patent Citations (1)
| Title |
|---|
| DATABASE GenBank 5 August 2021 (2021-08-05), "Connector enhancer of kinase suppressor of ras 2, partial [Galemys pyrenaicus]", XP093250729, Database accession no. KAG8508693.1 * |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| US11912985B2 (en) | Methods and compositions for simultaneous editing of both strands of a target double-stranded nucleotide sequence | |
| US12509680B2 (en) | Methods and compositions for prime editing nucleotide sequences | |
| US20250064979A1 (en) | Self-assembling virus-like particles for delivery of prime editors and methods of making and using same | |
| US20240067940A1 (en) | Methods and compositions for editing nucleotide sequences | |
| JP2024544013A (en) | Compositions and methods for effective in vivo delivery | |
| US20260009027A1 (en) | Prime editing-mediated readthrough of premature termination codons (pert) | |
| US20240382620A1 (en) | Genome editing compositions and methods for treatment of usher syndrome type 3 | |
| JP2025519070A (en) | Compositions and methods for efficient in vivo delivery | |
| WO2024138033A2 (en) | Compositions and methods for delivery of nucleic acid editors | |
| WO2024178144A1 (en) | Methods and compositions for editing nucleotide sequences | |
| WO2024254550A1 (en) | Compositions and methods for promoting payload delivery in cells | |
| WO2023028180A2 (en) | Genome editing compositions and methods for treatment of retinopathy | |
| WO2024254530A1 (en) | Pleckstrin holomogy domains for delivery of protein payloads to a target cell | |
| WO2024254518A2 (en) | Compositions of lipid delivery particles and method of use thereof | |
| WO2024254544A2 (en) | Compositions and methods for promoting payload delivery in cells | |
| WO2023081787A2 (en) | Genome editing compositions and methods for treatment of fanconi anemia | |
| WO2025235617A1 (en) | Fusogenic lipid particles for delivery of payload in cells | |
| WO2025064678A2 (en) | Prime editing-mediated readthrough of frameshift mutations (perf) |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 24820184 Country of ref document: EP Kind code of ref document: A1 |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |