WO2014007273A1 - 有用微生物および目的物質の製造方法 - Google Patents
有用微生物および目的物質の製造方法 Download PDFInfo
- Publication number
- WO2014007273A1 WO2014007273A1 PCT/JP2013/068205 JP2013068205W WO2014007273A1 WO 2014007273 A1 WO2014007273 A1 WO 2014007273A1 JP 2013068205 W JP2013068205 W JP 2013068205W WO 2014007273 A1 WO2014007273 A1 WO 2014007273A1
- Authority
- WO
- WIPO (PCT)
- Prior art keywords
- gene
- compound
- promoter
- acid
- strain
- Prior art date
- Legal status (The legal status is an assumption and is not a legal conclusion. Google has not performed a legal analysis and makes no representation as to the accuracy of the status listed.)
- Ceased
Links
Images
Classifications
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N1/00—Microorganisms; Compositions thereof; Processes of propagating, maintaining or preserving microorganisms or compositions thereof; Processes of preparing or isolating a composition containing a microorganism; Culture media therefor
- C12N1/20—Bacteria; Culture media therefor
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/11—DNA or RNA fragments; Modified forms thereof; Non-coding nucleic acids having a biological activity
- C12N15/52—Genes encoding for enzymes or proenzymes
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N15/00—Mutation or genetic engineering; DNA or RNA concerning genetic engineering, vectors, e.g. plasmids, or their isolation, preparation or purification; Use of hosts therefor
- C12N15/09—Recombinant DNA-technology
- C12N15/63—Introduction of foreign genetic material using vectors; Vectors; Use of hosts therefor; Regulation of expression
- C12N15/70—Vectors or expression systems specially adapted for E. coli
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12N—MICROORGANISMS OR ENZYMES; COMPOSITIONS THEREOF; PROPAGATING, PRESERVING, OR MAINTAINING MICROORGANISMS; MUTATION OR GENETIC ENGINEERING; CULTURE MEDIA
- C12N9/00—Enzymes; Proenzymes; Compositions thereof; Processes for preparing, activating, inhibiting, separating or purifying enzymes
- C12N9/10—Transferases (2.)
- C12N9/1048—Glycosyltransferases (2.4)
- C12N9/1051—Hexosyltransferases (2.4.1)
- C12N9/1055—Levansucrase (2.4.1.10)
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P13/00—Preparation of nitrogen-containing organic compounds
- C12P13/04—Alpha- or beta- amino acids
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P13/00—Preparation of nitrogen-containing organic compounds
- C12P13/04—Alpha- or beta- amino acids
- C12P13/06—Alanine; Leucine; Isoleucine; Serine; Homoserine
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P13/00—Preparation of nitrogen-containing organic compounds
- C12P13/04—Alpha- or beta- amino acids
- C12P13/08—Lysine; Diaminopimelic acid; Threonine; Valine
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P13/00—Preparation of nitrogen-containing organic compounds
- C12P13/04—Alpha- or beta- amino acids
- C12P13/14—Glutamic acid; Glutamine
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P13/00—Preparation of nitrogen-containing organic compounds
- C12P13/04—Alpha- or beta- amino acids
- C12P13/20—Aspartic acid; Asparagine
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P7/00—Preparation of oxygen-containing organic compounds
- C12P7/40—Preparation of oxygen-containing organic compounds containing a carboxyl group including Peroxycarboxylic acids
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P7/00—Preparation of oxygen-containing organic compounds
- C12P7/40—Preparation of oxygen-containing organic compounds containing a carboxyl group including Peroxycarboxylic acids
- C12P7/42—Hydroxy-carboxylic acids
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P7/00—Preparation of oxygen-containing organic compounds
- C12P7/40—Preparation of oxygen-containing organic compounds containing a carboxyl group including Peroxycarboxylic acids
- C12P7/44—Polycarboxylic acids
-
- C—CHEMISTRY; METALLURGY
- C12—BIOCHEMISTRY; BEER; SPIRITS; WINE; VINEGAR; MICROBIOLOGY; ENZYMOLOGY; MUTATION OR GENETIC ENGINEERING
- C12P—FERMENTATION OR ENZYME-USING PROCESSES TO SYNTHESISE A DESIRED CHEMICAL COMPOUND OR COMPOSITION OR TO SEPARATE OPTICAL ISOMERS FROM A RACEMIC MIXTURE
- C12P7/00—Preparation of oxygen-containing organic compounds
- C12P7/40—Preparation of oxygen-containing organic compounds containing a carboxyl group including Peroxycarboxylic acids
- C12P7/44—Polycarboxylic acids
- C12P7/46—Dicarboxylic acids having four or less carbon atoms, e.g. fumaric acid, maleic acid
Definitions
- the present invention relates to a microorganism for efficiently producing useful substances such as organic acids and amino acids, and a method for producing useful substances using the microorganisms.
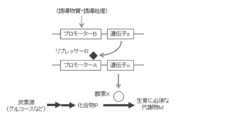
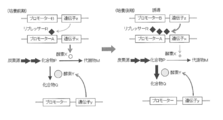
- FIG. 1 in order to efficiently produce compound P in a biosynthetic pathway or metabolic pathway in which a metabolite M essential for growth is produced from a carbon source via compound P that is an intermediate metabolite, A method for obtaining an auxotrophic mutant deficient in the activity of enzyme X that converts compound P to metabolite M has been used frequently.
- the target compound P is produced using the mutant strain, it is necessary to add the metabolite M or the final product generated from the metabolite M to the medium, and there is a problem that the manufacturing cost increases.
- the metabolite M or the final product produced from the metabolite M is controlled by artificially placing the transcription of the gene x encoding the enzyme X under the control of the repressor to control the amount of metabolite M produced.
- the transcription amount of gene x is controlled using another promoter that is not subject to the original promoter of gene x but is repressed by a repressor. There have been no reports of cases where the production amount of compound P was improved by doing so.
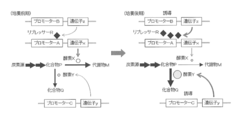
- a useful compound is produced using a microorganism, in a biosynthetic pathway or a metabolic pathway in which a metabolite M essential for growth is produced from a carbon source via a compound P that is an intermediate metabolite, from a compound P on the pathway
- a method of producing the useful compound by a branched route is often used. Even when this method is used, a method of obtaining an auxotrophic mutant deficient in the activity of the enzyme X has been often used.
- a metabolite M or a final product generated from the metabolite M is added to the medium. There is a problem that the manufacturing cost is high.
- An object of the present invention is to produce a compound P using a microorganism having a biosynthetic pathway or a metabolic pathway that generates a metabolite M essential for growth from a carbon source via a compound P that is an intermediate metabolite.
- a method for obtaining a strain capable of efficiently accumulating compound P in a medium in which the amount of conversion from P to metabolite M is reduced and no metabolite M or a final product generated from metabolite M is added (FIG. 1).
- it is to provide a method for producing compound Q by enzyme Y based on accumulated compound P (FIG. 3).
- the present inventors solved the above-mentioned problems in research for constructing a strain that efficiently produces DHS as a target substance using Corynebacterium glutamicum ATCC13032 strain (hereinafter abbreviated as ATCC13032 strain). Tried to do.
- ATCC13032 strain Corynebacterium glutamicum ATCC13032 strain
- the gene qsuB (cg0502) encoding DHS dehydrase (enzyme Y) that enhances the expression of a plurality of enzymes involved in the biosynthetic pathway from glucose to DHS (compound P) and further promotes the conversion from DHS to protocatechuic acid.
- This NSH ⁇ aroE3 strain was confirmed to produce 0.5 g / L protocatechuic acid by culturing in a medium supplemented with three kinds of aromatic amino acids and shikimic acid.
- the NSH ⁇ aroE3_vanE3 strain was constructed by integrating DNA into which the aroE3 gene was linked to the vanA (cg2616) gene promoter (hereinafter abbreviated as vanA promoter).
- This NSH ⁇ aroE3_vanE3 strain was found to be capable of growing in a synthetic medium without addition of three kinds of aromatic amino acids and shikimic acid, and produced 16.6 g / L of protocatechuic acid, thereby solving the problems of the present invention.
- a strain that controls the expression of the aroE3 gene with a repressor encoded by the rhcR (cg1308) gene (hereinafter abbreviated as RhcR repressor) was constructed.
- RhcR repressor a repressor encoded by the rhcR
- the qsuD (cg0504) gene encoding shikimate dehydrogenase that promotes the conversion of DHS to shikimate (metabolite M) and qsuB encoding DHS dehydrase
- the NSH ⁇ aroE3 ⁇ qsuB ⁇ qsuD strain in which a deletion mutation was also introduced into the gene was constructed, it was confirmed that the NSH ⁇ aroE3 ⁇ qsuB ⁇ qsuD strain could not grow unless three kinds of aromatic amino acids, vitamin K2 and paraaminobenzoic acid were added.
- NSH ⁇ aroE3 ⁇ qsuB ⁇ qsuD strain was confirmed to produce 7.7 ⁇ g / L DHS by culturing in a medium containing these additives.
- the NSH ⁇ aroE3_vanE3 ⁇ qsuB ⁇ qsuD strain was constructed by introducing deletion mutations into the qsuD gene and the qsuB gene based on the NSH ⁇ aroE3_vanE3 strain by the same procedure as the construction of the NSH ⁇ aroE3 ⁇ qsuB ⁇ qsuD strain.
- the Pben-vanR strain was constructed by integrating the benA (cg2637) gene promoter and the vanR gene encoding the vanR repressor into the chromosome.
- the present inventors have found that 15.4 g / L DHS can be produced in addition to growing in a synthetic medium that does not contain any substances, and have solved the problems of the present invention.
- the present invention relates to the following (1) to (24).
- prokaryotes having all the properties described in (a) to (d) below.
- the promoter B is selected from the group consisting of ferulic acid, vanillic acid, vanillin, benzoic acid, 3-hydroxybenzoic acid, resorcinol, 4-hydroxybenzoic acid, 2,4-dihydroxybenzoic acid, fructose and sucrose.
- the prokaryotic organism according to any one of (1) to (3), wherein the prokaryotic organism is a promoter induced by addition of a compound.
- the gene z encoding the protein of the repressor R is the vanR (cg2615) gene, pcaR (cg2624) gene, or rhcR (cg1308) gene of Corynebacterium glutamicum ATCC13032 strain, The prokaryotic organism according to any one of (1) to (4).
- the promoter A is a promoter of vanA (cg2616) gene, promoter of pobA (cg1226) gene, promoter of pcaH (cg2631) gene or promoter of rhcH (cg1309) gene of Corynebacterium glutamicum ATCC13032 strain.
- a method for producing compound P comprising culturing the prokaryote according to any one of (1) to (6) in the presence of a carbon source in a state where transcription of promoter A is suppressed.
- the prokaryotic organism according to any one of (1) to (6) is cultured in the presence of a carbon source in a state where transcription of promoter A is suppressed, and then a treatment for inducing transcription of promoter B is added.
- the production method of the compound P characterized by reducing the production amount of the metabolite M by increasing the expression level of the protein of repressor R by this.
- the substitution mutation in the promoter region that promotes transcription of the gene y described in the property (e) indicates that the entire promoter or a part of the promoter is the original promoter and promoter A that promotes transcription of the gene.
- the promoter C is selected from the group consisting of ferulic acid, vanillic acid, vanillin, benzoic acid, 3-hydroxybenzoic acid, resorcinol, 4-hydroxybenzoic acid, 2,4-dihydroxybenzoic acid, fructose and sucrose.
- a method for producing compound Q comprising culturing the prokaryote according to any one of (11) to (16) in the presence of a carbon source in a state where transcription of promoter A is suppressed.
- the prokaryotic organism according to any one of (11) to (16) is cultured in the presence of a carbon source in a state where transcription of promoter A is suppressed, and then a treatment for inducing transcription of promoter B is added.
- the production method of compound Q characterized by decreasing the production amount of the metabolite M and increasing the production amount of compound Q by increasing the expression level of the protein of repressor R by this.
- a method for producing compound Q characterized by increasing the expression level of the protein of repressor R by adding to decrease the production amount of metabolite M and increase the production amount of compound Q.
- the production method of compound Q according to any one of (17) to (19), wherein the production amount of compound Q is increased by increasing.
- the combination of the compound P, the compound Q, and the metabolite M is any one of (j) to (w) below, and any one of (17) to (20) The manufacturing method of the compound Q as described in.
- Compound P is 3-dehydroshikimic acid, Compound Q is protocatechuic acid, and Metabolite M is shikimic acid.
- Compound P is chorismic acid, compound Q is anthranilic acid, and metabolite M is prefenic acid.
- (M) Compound P is prefenic acid, Compound Q is allogenic acid, and Metabolite M is phenylpyruvic acid.
- N Compound P is prefenic acid, Compound Q is phenylpyruvic acid, and metabolite M is 4-hydroxyphenylpyruvic acid or allogenic acid.
- O Compound P is 2-oxoisovaleric acid, Compound Q is 2-isopropylmalic acid, and Metabolite M is valine.
- (P) Compound P is 2-oxoisovaleric acid, Compound Q is valine, and Metabolite M is 2-isopropylmalic acid.
- Compound P is glutamic acid, Compound Q is ⁇ -glutamyl phosphoric acid, and Metabolite M is N-acetylglutamic acid.
- R Compound P is glutamic acid, Compound Q is N-acetylglutamic acid, and Metabolite M is ⁇ -glutamylphosphoric acid.
- S Compound P is aspartic acid, Compound Q is asparagine, and metabolite M is ⁇ -alpartyl phosphate.
- T Compound P is aspartic acid ⁇ -semialdehyde, Compound Q is 2,3-dihydrodipicolinic acid, and Metabolite M is homoserine.
- Compound P is homoserine, compound Q is O-acetylhomoserine, and metabolite M is homoserine phosphate.
- V Compound P is homoserine, compound Q is homoserine phosphate, and metabolite M is O-acetylhomoserine.
- W Compound P is serine, Compound Q is O-acetylserine, and Metabolite M is glycine.
- microorganisms that efficiently produce useful organic compounds such as DHS and protocatechuic acid can be obtained based on a carbon source such as glucose. Moreover, if the microorganism of this invention is used, DHS, a protocatechuic acid, etc. can be efficiently manufactured using the culture medium which does not contain an aromatic amino acid.
- the target compound P on the biosynthetic pathway from the carbon source to the metabolite M essential for growth is produced by controlling the expression of the enzyme by the repressor R which is the product of the gene z, and the compound P is converted from the compound P to the compound Q.
- enzyme X is normally produced by releasing the transcriptional repression of promoter A by repressor R.
- enzyme X is restored by reversing the transcriptional repression of promoter A by repressor R.
- the figure which shows that the expression level of is reduced significantly and the compound P accumulate
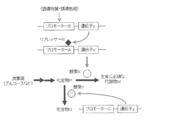
- enzyme X is normally produced by releasing the transcriptional repression of promoter A by repressor R.
- the repression of promoter A by repressor R is restored to restore enzyme X.
- stores is converted into the compound Q by the effect
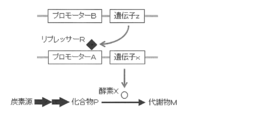
- the amount of repressor R is small, so that enzyme X is normally produced by transcription of promoter A.
- the amount of repressor R increases, so the expression level of enzyme X is greatly reduced.
- stores converts into the compound Q by the effect
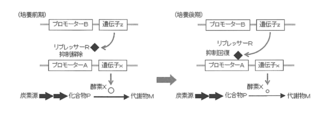
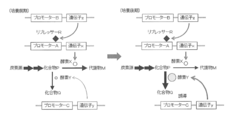
- the expression of enzyme X is kept low by repressing transcription of promoter A by repressor R, and the production amount of metabolite M is restricted, so that a small amount of compound P can be produced while growing microorganisms.
- the latter figure shows that the production amount of compound Q is increased by inducing and expressing enzyme Y that converts compound P to compound Q to increase the expression amount of enzyme Y.
- the expression of enzyme X is kept low by repressing transcription of promoter A by repressor R, and the production amount of metabolite M is restricted, so that a small amount of compound P can be produced while growing microorganisms.
- the repressor R is induced and expressed to increase the accumulation amount of the compound P
- the enzyme Y that converts the accumulated compound P into the compound Q is induced and expressed to increase the expression amount of the enzyme Y,
- the microorganism of the present invention is a microorganism used for efficiently producing a target compound P which is an intermediate metabolite on a biosynthetic pathway from a carbon source to a metabolite M essential for growth, and has the following four properties.
- the microorganism is preferably a prokaryotic organism in which a transcription repression system by repressor R is developed for gene expression control.
- A Among the enzyme groups involved in biosynthesis from the carbon source that can be used by the microorganism to the compound P, the activity of any one or more of the enzymes is enhanced compared to the wild-type prokaryotic strain.
- the activity of the enzyme X is deficient or reduced by having a mutation due to substitution, deletion or addition.
- the property (a) can be imparted to the microorganism used in the present invention. That is, by introducing a mutation into the translation initiation region of the gene encoding the enzyme protein, introducing a mutation into the promoter region of the gene, replacing the promoter with another strong promoter, cloning into a multicopy plasmid, etc.
- the amount of compound P produced can be increased by increasing the amount.
- the amount of compound P produced can also be increased by introducing an amino acid mutation that cancels the feedback inhibition into the enzyme protein. Mutation introduction into these DNAs or proteins can be performed by mutagenesis treatment or the like, but can also be performed using recombinant DNA technology or homologous recombination technology.
- the activity of the enzyme X can be deleted using a mutagenesis treatment or a recombinant DNA technique.
- the translation region of the wild type gene x is used using a homologous recombination technique.
- a deletion mutation is introduced in the inside. In this case, if the entire translation region is deleted, downstream gene expression may be inhibited due to the polar effect. Therefore, the translation region lacks the translation region in-frame, leaving the 5 'and 3' portions. It is desirable to introduce a mutation.
- the activity of enzyme X is also lost by introducing a mutation by substitution, deletion or addition of one or more bases in the vicinity of the translation initiation region of gene x or in the promoter region that promotes transcription of gene x. be able to.
- the property (b) and the property (c) may be simultaneously imparted by replacing the original promoter region that promotes transcription of the gene x with the DNA of the promoter A described in the property (c).
- Property (c) can be imparted by placing DNA encoding the protein of active enzyme X under the control of promoter A, whose transcription is repressed by repressor R.
- the promoter A is a promoter different from the original promoter that promotes transcription of the gene x, and may be any promoter as long as transcription is suppressed by the repressor R.
- the promoter A repressed by the repressor R in the Escherichia coli K-12 strain is a pL promoter or pR promoter that undergoes transcriptional repression by the cI857 gene product of ⁇ phage, and lac that undergoes transcriptional repression by the lacI gene product.
- Examples include promoters or promoters of the gal operon that are subject to transcriptional repression by the galR gene product.
- vanR repressor the vanA promoter that undergoes transcriptional repression by the product of the vanR (cg2615) gene (hereinafter referred to as vanR repressor) (hereinafter abbreviated as VanR repressor), rhcR (Cg1308) promoter (hereinafter abbreviated as rhcH promoter) of rhcH (cg1309) gene (hereinafter abbreviated as rhcH gene) subjected to transcriptional repression by the product (hereinafter abbreviated as rhcR gene) of gene (cg1308),
- the promoter (hereinafter abbreviated as pcaI promoter) of the pcaI (cg2623) gene hereinafter referred to as the pcaI gene
- the microorganism of the present invention has a low expression level of enzyme X in a normal state due to the addition of the properties (b) and (c).
- the microorganism since the microorganism has a mutation that increases the amount of the compound P due to the addition of the property (a), the microorganism can produce an amount of the metabolite M necessary for growth.
- the growth of the microorganism can be improved by releasing the transcriptional repression by repressor R and increasing the expression level of enzyme X.
- Property (d) is not only a wild-type expression unit of gene z encoding the protein of repressor R, but also an expression unit of gene z in which gene z is placed under the control of an inducible promoter B different from the promoter of the gene. It can also be given by forming. In addition to the wild type expression unit of gene z, the expression unit of gene z placed under the control of the inducible promoter B may be incorporated into the microorganism of the present invention.
- the repressor R any repressor can be used as long as it is a prokaryotic-derived repressor. However, it is preferable to use a repressor that can release the suppression by adding an inducer or changing temperature.
- Escherichia coli K-12 strain as the microorganism of the present invention, for example, the cI857 gene of ⁇ phage that is induced by culturing at 38 ° C. or higher, the lacI gene that is induced by addition of lactose, A repressor encoded by a galR gene that is induced by addition of galactose or a tetR gene that is induced by addition of tetracycline can be used.
- the suppression is released by the addition of VanR repressor, resorcinol or 2,4-dihydroxybenzoic acid that is released by the addition of ferulic acid, vanillic acid or vanillin.
- RhcR repressor, or PcaR repressor whose inhibition is released by the addition of 4-dihydroxybenzoic acid can be used.
- promoter B any promoter can be used as long as it is an inducible promoter derived from prokaryotes. Specifically, when gene z is expressed in Escherichia coli K-12, the ⁇ phage pL promoter, pR promoter, lac promoter, gal operon promoter, trp operon promoter, araBAD promoter, or the like is used. be able to.
- vanA promoter which is induced by addition of ferulic acid, vanillic acid or vanillin, rhcH promoter which is induced by addition of resorcinol or 2,4-dihydroxybenzoic acid, 4-hydroxybenzoic acid Induced by addition of pcaI promoter induced by addition of acid, nagI (cg3351) gene (hereinafter referred to as nadI gene) promoter induced by addition of 3-hydroxybenzoic acid, benzoic acid Promoter of benA (cg2637) gene (hereinafter referred to as benA gene) (hereinafter referred to as benA promoter), promoter of cg2118 gene or ptsS (cg2925) gene (hereinafter referred to as benA promoter) , PtsS gene Promoter can be used for).
- the microorganism of the present invention has an enzyme activity that metabolizes compound P in addition to enzyme X, and there is a possibility that the accumulated amount of the compound will decrease in the production of compound P, mutagenesis treatment or recombinant DNA It is preferable to delete or reduce the activity of any one or more of the metabolic enzymes of compound P possessed by the microorganism, depending on the technique.
- Corynebacterium Corynebacterium
- Brevibacterium Brevibacterium
- Arthrobacter Arthrobacter
- Nocardioides Oida CEA d Nocardioidaceae
- micro Agrobacterium Microbacterium
- Streptomyces Seth Streptomyces Seth
- Amiko Kakuto-flops cis Amycolatopsis
- Rhodococcus Rhodococcus
- Kineokokkasu Kinineococcus
- Acinetobacter Acinetobactor
- Pseudomonas Pseudomonas
- Pantoea Pantoea
- Klebsiella Klebsiella
- Prokaryotes belonging to any of the genus Escherich
- the method for producing compound P using the microorganism of the present invention includes, for example, a method for producing target compound P by culturing in a medium containing a carbon source while suppressing transcription of promoter A.
- a method for producing target compound P by culturing in a medium containing a carbon source while suppressing transcription of promoter A since the transcription amount of the promoter A controlling the expression of the enzyme X is suppressed to a low level by the repressor R, the production amount of the metabolite M is limited, and the compound P is efficiently produced. (FIG. 4).
- the production process of the compound P is carried out in the first culture phase, the second culture phase and the second culture phase. It is also possible to produce the target compound P by dividing. That is, after inoculating the microorganism in a medium containing a carbon source, transcriptional repression is canceled by adding an inducer or increasing or decreasing the culture temperature according to the nature of the repressor R, thereby reducing the expression level of enzyme X. Increase and continue culture.
- the amount of compound P produced can be increased by restoring the suppression by the repressor R at an appropriate time and suppressing the expression of the enzyme X (FIG. 5).
- suppression by repressor R can be recovered by adjusting the amount of the inducer added and depleting the inducer in the later stage of culture.
- promoter A which is induced by an increase or decrease in culture temperature
- the suppression by repressor R can be recovered by returning the culture temperature to a temperature at which suppression by repressor R occurs.
- the target compound P is produced by dividing the production process of the compound P into two phases, ie, the first culture phase and the second culture phase.
- the inducible promoter B is activated by adding an inducer or raising or lowering the culture temperature depending on the nature of the inducible promoter B.
- the compound when the production method using a microorganism of the present invention is used, the compound can be easily obtained by changing the culture method using the same microorganism according to the growth state of the microorganism or the effect of compound P on the microorganism. P production can be maximized.
- any carbon source can be used as long as it can be used by the microorganism of the present invention.
- sugars such as glucose, sucrose and fructose
- alcohols such as ethanol and methanol
- organic acids such as citric acid, malic acid and succinic acid, glycerol, molasses and the like can be used.
- Compound P may be any organic compound as long as it is a metabolite on the biosynthetic pathway that generates a metabolite essential for growth from a carbon source. Examples include DHS, glutamic acid, aspartic acid, and serine.
- DHS can be produced by controlling the expression of shikimate dehydrogenase (enzyme X) that produces shikimate (metabolite M). DHS is useful as an antioxidant (United States States Patent 5,821,266) and can be used as a raw material for producing protocatechuic acid and shikimic acid using microorganisms.
- Glutamic acid which is a useful amino acid, is mino acid-N-acetyltransferase (enzyme X) that produces N-acetylglutamic acid (metabolite M) or ⁇ -glutamyl kinase (enzyme X) that produces ⁇ -glutamylphosphate (metabolite M). ) To control the expression.
- Aspartic acid which is a useful amino acid, expresses the expression of aspartokinase (enzyme X) that produces ⁇ -alpartyl phosphate (metabolite M) or asparagine synthase (enzyme X) that produces asparagine (metabolite M). Produced by controlling.
- Serine a useful amino acid, regulates the expression of serine hydroxymethyltransferase (enzyme X) that produces glycine (metabolite M) or serine transferase (enzyme X) that produces O-acetylserine (metabolite M) Can be produced.
- ATCC13032 strain that efficiently produces DHS will be described in detail.
- GB accession number BX927147 from the GenBank (hereinafter abbreviated as GB) database of National Center for Biotechnology Information (hereinafter abbreviated as NCBI).
- NCBI National Center for Biotechnology Information
- a strain derived from the ATCC 13032 strain producing DHS is a microorganism having the following properties (aa) to (dd).
- Bb a translational region or translational regulatory region of a wild-type gene (gene x) encoding a protein of shikimate dehydrogenase (enzyme X) that converts DHS into shikimate (metabolite M), or transcription of the gene
- gene x a wild-type gene
- enzyme X protein of shikimate dehydrogenase
- the ability to convert DHS to shikimic acid is deficient or reduced by having mutations due to substitution, deletion or addition of one or more bases in the promoter region to be promoted.
- the ATCC13032 strain has aroE3 (cg1835) gene (DNA region of complementary strand base numbers 1,726,078 to 1,726,908) and aroE1 (cg1283) gene (complementary strand base numbers of 1,182,337 to 1,183,143) as genes encoding active shikimate dehydrogenase. Region) and qsuD (cg0504) gene (DNA region of nucleotide numbers 446,537 to 447,388), but introduced mutations into the aroE3 gene, which plays a major role in the conversion of DHS to shikimate,
- the property bb can also be imparted by deletion or reduction of the expression of the active shikimate dehydrogenase of the gene.
- the qsuD gene and / or the aroE1 gene may be imparted with the property bb by introducing a mutation in which the expression of the active shikimate dehydrogenase of the gene is deleted or reduced.
- Good. The transcription of DNA encoding active enzyme X is controlled by promoter A, which is different from the original promoter that promotes transcription of gene x and whose transcription is suppressed by the protein of repressor R.
- Dd One or more copies of the gene z encoding the protein of the repressor R are contained, and transcription of the gene is controlled by the original promoter of the gene and / or an inducible promoter B different from the promoter.
- the repressor R includes, as described above, a VanR repressor whose deregulation is released by addition of ferulic acid, vanillic acid or vanillin, a RhcR repressor whose deregulation is released by addition of resorcinol or 2,4-dihydroxybenzoic acid.
- a PcaR repressor that is derepressed by the addition of 4-dihydroxybenzoic acid can be used.
- promoter A that is repressed by repressor R in the ATCC13032 strain is a vanA promoter that is repressed by VanR repressor, a rhcH promoter that is repressed by RhcR repressor, or transcription repressed by PcaR repressor. And the promoter of the pcaI gene.
- Promoter B is induced by addition of ferulic acid, vanillic acid or vanillin, vanA promoter which is induced by addition of resorcinol or 2,4-dihydroxybenzoic acid, induction by rhcH promoter, which is induced by addition of 4-hydroxybenzoic acid
- rhcH promoter which is induced by addition of 4-hydroxybenzoic acid
- Use pcaI promoter, nagI promoter that is induced by addition of 3-hydroxybenzoic acid benA promoter that is induced by addition of benzoic acid
- promoter of cg2118 gene that is induced by addition of either fructose or sucrose, or ptsS promoter be able to.
- the property (bb) And the property (cc) may be simultaneously imparted.
- a mutation that enhances the pathway for generating DHS from DAHP or a mutation that increases the amount of DAHP generated may be introduced.
- DAHP is converted to 3-dehydroquinic acid (hereinafter abbreviated as DHQ) by 3-dehydroquinic acid synthase, and DHQ is further converted to DHS by DHQ dehydrase. Therefore, a mutation occurs in the transcriptional / translational regulatory region of the gene aroB (cg1827) gene (hereinafter referred to as aroB gene) encoding DHQ synthase and / or the aroD (cg0503) gene (hereinafter referred to as aroD gene) encoding DHQ dehydrase.
- aroB gene the transcriptional / translational regulatory region of the gene aroB gene cg1827 gene
- aroD gene the aroD gene cg0503 gene
- the expression level of DHQ synthase and / or DHQ dehydrase can be enhanced and the amount of DHS can be increased.
- the amount of DHS can be increased by replacing the original promoter that promotes transcription of the aroB gene and
- the DHS amount can be increased by increasing the DAHP amount.
- Methods for increasing the amount of DAHP include cancellation of feedback inhibition by aromatic amino acids for DAHP synthase, enhancement of the expression level of DAHP synthase, or erythrose 4-phosphate (hereinafter referred to as E4P) which is a substrate of DAHP synthase. Abbreviated) or phosphoenolpyruvate (hereinafter abbreviated as PEP).
- DAHP synthase is often feedback-inhibited by tryptophan, tyrosine or phenylalanine. Since it is known that feedback inhibition can be canceled by introducing amino substitution mutations into DAHP synthase, aroF (cg1129) gene (hereinafter referred to as aroF gene) and aroG (cg2391) gene (hereinafter referred to as aroG gene) Mutations can be introduced into the DAHP synthase gene such as to increase the amount of DAHP.
- Examples of a method for enhancing the expression level of DAHP synthase include a method for introducing a mutation into the transcriptional / translational regulatory region of a gene encoding DAHP synthase.
- the amount of DAHP can be increased by replacing the original promoter that promotes transcription of the gene encoding DAHP synthase with the Tu promoter.
- a tkt gene encoding transketolase (hereinafter referred to as tkt gene), a tal (cg1776) gene encoding transaldolase (hereinafter referred to as cg1776).
- Gene or zwf (cg1778) gene encoding glucose-6-phosphate 1-dehydrogenase, etc. to introduce the mutation into the transcriptional / translational regulatory region of the enzyme gene involved in the pentose phosphate pathway There is a way to enhance it.
- a mutation is made in the transcription / translation regulatory region of pps (cg0644) gene encoding PEP synthase or pck (cg3169) gene encoding PEP carboxykinase. And a method for enhancing the expression of the gene.
- the amount of PEP can be increased by introducing a mutation into the translation region of the gene or the transcription / translation regulatory region of the gene to suppress the conversion of PEP.
- mutations may be introduced into a single gene, but in order to further improve the amount of DHS produced, it is preferable to introduce mutations into a plurality of genes.
- a strain derived from the ATCC13032 strain having the properties (aa) to (dd) described above can produce DHS based on a carbon source such as glucose. Since the aroD gene encoding DHQ dehydrase forms an operon with the qsuB (cg0502) gene encoding the DHS dehydrase that converts DHS into protocatechuic acid (hereinafter referred to as the qsuB gene), it imparts a property (aa) Therefore, when the expression of the aroD gene is increased by promoter replacement, the expression of the qsuB gene is also increased, so that a part of the accumulated DHS is converted to protocatechuic acid. In order to suppress the conversion to protocatechuic acid, it is desirable to introduce a mutation into the translation region of the qsuB gene or the transcription / translation control region of the gene.
- the microorganism of the present invention has the following property (e) in addition to the above four properties (a) to (d), by using the microorganism, it is converted from a carbon source to a metabolite M essential for growth.
- the target compound Q can be efficiently produced by converting the compound P on the biosynthetic route to the compound Q.
- substitution of one or more bases in the translation region of gene y encoding the protein of enzyme Y that converts compound P to compound Q, the translational regulatory region of the gene or the promoter region that promotes transcription of gene y By having a mutation due to deletion or addition, the ability to convert compound P to compound Q is enhanced, or the expression of gene y is controlled differently from the wild type.
- the expression level of the enzyme Y is often not sufficient in the wild type state, one or more bases are contained in the translational regulatory region of the gene y or the promoter region that promotes transcription of the gene y. Introduce mutation by substitution, deletion, or addition to enhance the expression level of enzyme Y, or to give property (e) by placing the transcription of gene y under control different from that of wild type promoter can do.
- the first enzyme in the branch pathway is often subject to feedback inhibition by the final product, the activity of enzyme Y is enhanced by releasing the feedback inhibition by introducing a mutation into the translation region of gene y.
- the property (e) can also be imparted.
- the property (e) is imparted by replacing the entire promoter or a part of the promoter that promotes transcription of the gene y with DNA of a promoter C that is different from the original promoter that promotes transcription of the gene and the promoter A. be able to.
- promoter C a promoter different from the original promoter of the gene y and different from the promoter A can be used.
- promoter C a constitutive expression promoter can be used, but an inducible promoter is preferably used.
- inducible promoter any of the promoters listed as examples of the promoter B used for imparting the above property (d) can also be used.
- promoter C ⁇ phage pL promoter or pR promoter, lac promoter, gal operon promoter, trp operon promoter, araBAD promoter, etc., or vanA promoter that is induced by addition of ferulic acid, vanillic acid, or vanillin RhcH promoter induced by addition of resorcinol or 2,4-dihydroxybenzoic acid, pcaI promoter induced by addition of 4-hydroxybenzoic acid, nagI promoter induced by addition of 3-hydroxybenzoic acid, benzoic acid
- vanA promoter that is induced by addition, or the promoter of the cg2118 gene or the ptsS promoter that is induced by addition of either fructose or sucrose can be used.
- Promoter C used for imparting property (e) may be the same as promoter B used for imparting property (d).
- the promoter C may be a promoter that undergoes transcriptional control by a protein that controls transcription of the promoter B.
- the microorganism of the present invention has an enzyme activity for metabolizing Compound Q, and there is a possibility that the accumulated amount of the compound is reduced in the production of Compound P, mutagenesis treatment or recombinant DNA technology is used. It is preferable that the activity of any one or more metabolic enzymes of compound P possessed by the microorganism is deficient or reduced.
- Examples of the method for producing compound Q using the microorganism of the present invention include a method for producing compound Q by culturing in a medium containing a carbon source while suppressing transcription of promoter A.
- the transcription amount of the promoter A that controls the expression of the enzyme X is suppressed by the repressor R, the production amount of the metabolite M is limited, and the compound Q is efficiently converted from the compound P. It can be produced well (Fig. 7).
- the production process of compound Q is carried out in the first culture phase, the second culture phase, and the second culture phase. It is preferable to produce compound Q by dividing. That is, after inoculating the microorganism in a medium containing a carbon source, the expression level of enzyme X is reduced by releasing transcriptional repression by adding an inducer or increasing or decreasing the culture temperature according to the nature of the repressor R. Increase and continue culture. Thereafter, the amount of compound Q produced from compound P can be increased by restoring inhibition by repressor R at an appropriate time and inhibiting the expression of enzyme X (FIG. 8).
- suppression by repressor R can be recovered by adjusting the amount of the inducer added and depleting the inducer in the later stage of culture. .
- the suppression by repressor R can be recovered by returning the culture temperature to a temperature at which suppression by repressor R occurs.
- the compound Q is produced by dividing the production process of the compound Q into two phases of the first culture phase and the second culture phase. That is, after inoculating the microorganism in a medium containing a carbon source and growing the cells, the inducible promoter B is activated by adding an inducer or raising or lowering the culture temperature depending on the nature of the inducible promoter B.
- the production amount of the presser R By increasing the production amount of the presser R, the production of the metabolite M can be suppressed and the production amount of the compound Q from the compound P can be increased (FIG. 9).
- the production process of compound Q is cultured using a microorganism in which the expression of enzyme Y is placed under the control of inducible promoter C.
- Compound Q is preferably produced by dividing it into two phases, the first period and the latter period of culture. That is, after inoculating the microorganism in a medium containing a carbon source and growing the bacterial cells, depending on the nature of the inducible promoter C, the transcription of the inducible promoter C is activated by adding an inducer or raising or lowering the culture temperature.
- the production amount of compound Q from compound P can be increased (FIG. 10).
- inducible promoter C activation of inducible promoter C is activated late in the culture to increase the amount of enzyme Y produced, and inducible promoter B is activated to increase the amount of repressor R produced, thereby suppressing the production of metabolite M. Then, the amount of compound Q produced from compound P may be increased (FIG. 11).
- promoter C and promoter B the same promoter, the induction of promoter C can be performed at the same time as the induction of promoter B.
- promoter C and promoter B are selected from a group of promoters that are transcriptionally controlled by the same control protein, the induction of promoter C is the same as the induction of promoter B even if they are different promoters. Can be done at the time.
- the compound when the production method using a microorganism of the present invention is used, the compound can be easily obtained by changing the culture method using the same microorganism according to the growth state of the microorganism or the influence of compound Q on the microorganism. Q production can be maximized.
- the compound Q is produced by a pathway branched from the compound P in a biosynthetic pathway or metabolic pathway that produces a metabolite M essential for growth from a carbon source via the compound P. Any compound can be produced if it can produce
- Compound P may be any organic compound as long as it is a metabolite on the biosynthetic pathway that generates a metabolite essential for growth from a carbon source. Examples include DHS, chorismic acid, prefenic acid, 2-oxoisovaleric acid, glutamic acid, aspartic acid, aspartic acid ⁇ -semialdehyde, homoserine, and serine.
- Compound Q may be any organic compound as long as it is a substance obtained by converting compound P with enzyme Y.
- protocatechuic acid for example, protocatechuic acid, anthranilic acid, 4-hydroxyphenylpyruvic acid, allogenic acid, phenylpyruvic acid, 2-isopropylmalic acid, valine, ⁇ -glutamylphosphoric acid, N-acetylglutamic acid, asparagine, 2,3-dihydrodipicolinic acid and O -Acetyl homoserine and the like.
- compound P When compound P is DHS, shikimate (metabolite M) is produced by shikimate dehydrogenase (enzyme X), but protocatechuic acid (compound P) can be produced by DHS dehydrase (enzyme Y).
- Protocatechuic acid can be used as a raw material for the production of 2-pyrone-4,6-dicarboxylic acid, which is expected as a raw material for new plastics, using microorganisms, as well as a raw material for pharmaceuticals, agricultural chemicals, fragrances and the like.
- anthranilic acid is a synthetic raw material such as tryptophan, which is an aromatic amino acid. If the microorganism has a tryptophan biosynthetic pathway, tryptophan can be directly produced.
- phenylpyruvic acid (metabolite M) is produced by prefenic anhydrase (enzyme X), but the microorganism of the present invention is prefenic acid dehydrogenase in Escherichia coli K-12 strain or the like.
- 4-Hydroxyphenylpyruvic acid (Compound Q) can be produced by (Enzyme Y).
- 4-Hydroxyphenylpyruvic acid is a raw material for synthesizing tyrosine, an aromatic amino acid. If the microorganism has a tyrosine biosynthesis pathway from 4-hydroxyphenylpyruvic acid, tyrosine can also be directly produced.
- phenylpyruvic acid is produced by prefenic acid dehydrase (enzyme X).
- prefenic acid dehydrase enzyme X
- prefenic acid aminotransferase Allogenic acid (compound Q) can be produced by enzyme Y). Allogenic acid is a raw material for synthesizing tyrosine, an aromatic amino acid, and if the microorganism has a tyrosine biosynthesis pathway from allogenic acid, tyrosine can also be directly produced.
- 4-hydroxyphenylpyruvic acid is produced by the prefenic acid dehydrogenase (enzyme X) in the Escherichia coli K-12 strain or the like of the microorganism of the present invention.
- Allogeneic microorganisms such as Corynebacterium glutamicum produce allogenic acid (metabolite M) by prephenic acid dehydrogenase (enzyme X), but produce phenylpyruvic acid (compound Q) by prefenic acid dehydrating enzyme (enzyme Y). can do.
- Phenylpyruvic acid can be a raw material for the synthesis of phenylalanine, an aromatic amino acid. If the microorganism has a biosynthesis pathway for phenylalanine, phenylalanine can be directly produced.
- compound P When compound P is 2-oxoisovaleric acid, valine (metabolite M) is produced by branched-chain amino acid aminotransferase (enzyme X), but 2-isopropylmalic acid synthase (enzyme Y) produces 2-isopropylmalic acid. (Compound Q) can be produced.
- 2-Isopropylmalic acid is a raw material for the synthesis of leucine, and if the microorganism has a leucine biosynthesis pathway, leucine can also be produced directly.
- compound Q When compound P is 2-oxoisovaleric acid, 2-isopropylmalate (metabolite M) is produced by 2-isopropylmalate synthase (enzyme X), but valine is produced by branched-chain amino acid aminotransferase (enzyme Y). (Compound Q) can be produced.
- N-acetylglutamic acid (metabolite M) is produced by amino acid-N-acetyltransferase (enzyme X), but ⁇ -glutamylphosphate (compound Q) is produced by ⁇ -glutamyl kinase (enzyme Y).
- ⁇ -glutamyl phosphate serves as a raw material for proline synthesis, and proline can also be produced directly if the microorganism has a biosynthesis pathway for proline.
- N-acetylglutamic acid serves as a raw material for synthesizing arginine. If the microorganism has an arginine biosynthetic pathway, arginine can also be produced directly.
- compound Q When compound P is aspartate ⁇ -semialdehyde, homoserine (metabolite M) is produced by homoserine dehydrogenase (enzyme X), but 2,3-dihydrodipicolinic acid is produced by dihydroxypyrophosphate synthase (enzyme Y). (Compound Q) can be produced. 2,3-dihydrodipicolinic acid is a raw material for synthesizing lysine, and if the microorganism has a biosynthetic pathway for synthesizing lysine from 2,3-dihydrodipicolinic acid, lysine can also be produced directly.
- homoserine phosphate serves as a raw material for the synthesis of threonine and isoleucine, and if the microorganism has an isoleucine biosynthetic pathway, threonine and isoleucine can also be produced directly.
- O-acetylserine is a raw material for synthesizing cysteine. If the microorganism has a cysteine biosynthesis pathway, it can directly produce cysteine.
- the gene encoding enzyme Y involved in the production of compound Q may be a gene derived from the microorganism of the present invention, or may be a gene derived from a prokaryotic or eukaryotic organism different from the microorganism of the present invention. Good.
- the gene product may be used for the production of protocatechuic acid.
- Escherichia coli K-12 strain Has no DHS dehydrase gene, it is necessary to use DHS dehydrase genes derived from other microorganisms.
- the strain derived from the ATCC 13032 strain producing protocatechuic acid is a microorganism having the following property (ee) in addition to the properties (aa) to (dd) described above.
- ee Different from the original promoter that promotes transcription of the qsuB gene (gene y; base numbers 444, 184 to 446,040) encoding the protein of DHS dehydrase (enzyme Y) that converts DHS to protocatechuic acid, and also different from promoter A Promoter C controls the transcription of the qsuB gene.
- a strain derived from the ATCC13032 strain having the properties (aa) to (ee) described above can produce protocatechuic acid based on a carbon source such as glucose, but it is produced because the strain has protocatechuate dioxygenase. There is a possibility that the amount of protocatechuic acid is reduced by decomposing the produced protocatechuic acid.
- pcaG (cg2630) gene encoding protocatechuic acid 4,5-dioxygenase alpha subunit protein (complementary chain base number 2,511,382 to 2,511,996) and / or protocatechuic acid 4,5-dioxygenase beta subunit Introduces mutations by substitution, deletion or addition of one or more bases into the translation region of the pcaH (cg2631) gene encoding the protein (complementary strand base numbers 2,512,008 to 2,512,700) or the transcription / translation control region of the gene. By doing so, it is preferable to delete or reduce the degradation activity of the enzyme.
- this strain has parahydroxybenzoic acid hydroxylase which hydroxylates 5-position of protocatechuic acid and catalyzes the conversion to gallic acid, a part of the produced protocatechuic acid is converted to gallic acid.
- substitution of one or more bases in the translation region of the pobB (cg1226) gene (complementary strand base number 1,126,301 to 1,127,488) encoding the parahydroxybenzoate hydroxylase protein or the transcription / translation control region of the gene It is preferable that the hydroxylation activity of the parahydroxybenzoate hydroxylase is deleted or reduced by introducing a mutation caused by deletion or addition.
- the microorganism of the present invention is a genus Corynebacterium, Brevibacterium, Arthrobacter, Nocardioidaeae, Microbacterium, Streptomyces, Amicokactopsis, Any microorganism can be used as long as it belongs to the genus Rhodococcus, Kineococcus, Acinetobacter, Pseudomonas, Pantoae, Klebsiella, and Escherichia.
- the method for introducing mutations in the ATCC13032 strain will be described in detail.
- Escherichia coli K-12 strain is mainly used as a host bacterium used for introducing DNA cloning and mutation.
- the above Escherichia coli K-12 and ATCC13032 strains are cultured by known methods commonly used for culturing Escherichia coli and Corynebacterium bacteria, respectively. After culturing, the chromosomal DNA of the microorganism is isolated and purified by a known method (for example, the method described in Current Protocols in Molecular Biology). Obtaining a fragment containing DNA encoding the above promoter or metabolic enzyme from the chromosomal DNA by restriction enzyme digestion, polymerase chain reaction (PCR), or hybridization using synthetic DNA. Can do.
- PCR polymerase chain reaction
- any plasmid vector, phage vector, etc. can be used as long as it is a self-replicating vector in Escherichia coli K-12.
- pUC19 Gene, 33, 103 (1985)
- pUC18 pBR322
- the like can be used.
- Escherichia coli used as a host for the above vector can be any microorganism belonging to Escherichia coli, and specifically, Escherichia coli (XL) -XL1-Blue MRF '[manufactured by Stratagene Strategies, 5, 81 (1992)], Escherichia coli JM109, Escherichia coli BL21, and the like.
- the DNA is introduced into microorganisms belonging to the genera, Klebsiella and Escherichia, a vector capable of autonomous replication in these microorganisms is used.
- a vector capable of autonomous replication in these microorganisms is used.
- the recombinant DNA can be introduced into the host microorganism using a shuttle vector capable of autonomous replication in both microorganisms of both the microorganism and the Escherichia coli K-12 strain.
- shuttle vectors capable of autonomous replication in Corynebacterium bacteria include pAM330 (see JP-A-58-67699), pHM1519 (see JP-A-58-77895), and the like.
- pAM330 see JP-A-58-67699
- pHM1519 see JP-A-58-77895
- DNA fragments capable of autonomously replicating plasmids in Corynebacterium are extracted from these vectors and inserted into the Escherichia coli-Corynebacterium shuttle vector, both Escherichia coli and Corynebacterium are obtained.
- the plasmid pHM1519 replication region of the Corynebacterium glutamicum ATCC13058 strain is inserted into the pHSG298 (manufactured by Takara Bio Inc.), an Escherichia coli vector, as a replication region for replicating the plasmid in the Corynebacterium strain.
- the shuttle vector can be constructed.
- the shuttle vector can be retained.
- the ATCC13032 strain and the Corynebacterium glutamicum ATCC13058 strain can be obtained from an independent administrative agency, the National Institute for Product Evaluation Technology and the Biological Genetic Resources Department (hereinafter abbreviated as NITE).
- any method for introducing DNA can be used as long as it is a method for introducing DNA into the host cell, for example, a method using calcium ions [Proc. Natl. Acad. Sci. USA, 69, 2110 (1972) Electroporation method [Nucleic Acids Res., 16, 6127 (1988)].
- DNA can also be introduced into Corynebacterium glutamicum by the method of van der Rest et al. [Appl. Microbiol. Biotechnol., 52, 541 (1999)].
- DNA can be extracted from the transformant obtained as described above, and the base sequence of the DNA of the present invention contained in the DNA can be determined.
- a commonly used base sequence analysis method for example, dideoxy method [Proc.tlNatl. Acad. Sci. USA, 74, 54631977 (1977)] or 3730xl type DNA analyzer (manufactured by Applied Biosystems), etc. Can be used.
- the target DNA can also be prepared by chemical synthesis using a 8905 type DNA synthesizer manufactured by Perceptive Biosystems based on the DNA base sequence determined above.
- Strains with the desired phenotype include treatment with mutagens such as N-methyl-N'-nitro-N-nitrosoguanidine and ethylmethanesulfonic acid, mutation treatment by UV and radiation irradiation, and natural mutation. Is obtained by introducing a mutation.
- the strain clones the DNA of the region into which the mutation is introduced, and introduces the mutation into the DNA in vitro (in vitro).
- the plasmid is introduced into the strain, or between the plasmid and the chromosomal DNA.
- Substitution mutations can also be introduced by homologous recombination.
- the type of mutation to be introduced may be any of a base substitution mutation, a deletion mutation, an insertion mutation, or a substitution mutation with a large DNA fragment, as long as the desired phenotype appears. As long as it is above, it may be a mutation of any size.
- the above-described method is used as a method for introducing a mutation unless otherwise specified.
- the microorganism of the present invention when it is necessary to regulate the expression level of a gene other than a promoter or transcription control protein, adjustment of the distance between the ribosome binding sequence and the translation initiation codon, modification of the sequence around the translation initiation codon, or The translation initiation efficiency of the gene can be regulated by modifying the base sequence in the translation region.
- the DNA for introducing the mutation thus obtained is usually inserted into a plasmid that can be replicated in Escherichia coli, and then introduced into the host strain Corynebacterium glutamicum using the DNA introduction method described above. To do.
- the plasmid is a temperature-sensitive vector
- one cross-type homologous set is obtained by high-temperature culture selection, and when it has an antibiotic resistance marker such as kanamycin resistance that can be expressed in a host bacterium, the antibiotic resistance selection. Sort out replacements.
- W Jager et al. In order to obtain a mutant strain in which the DNA is replaced by double cross-type homologous recombination in a chromosomal region homologous to the DNA fragment, usually developed by W Jager et al. [J. Bacteriol.
- a method using a levansucrase gene (sacB gene) of Bacillus subtilis 168 strain can be used. Sucrose resistance selection is used. Using this method, Corynebacterium glutamicum expressing the sacB gene of Bacillus subtilis 168 strain is lethal in a medium containing sucrose. Strains can be obtained by sucrose resistance selection.
- the culture of the microorganism of the present invention can be performed in a normal nutrient medium containing a carbon source, a nitrogen source, an inorganic salt, various vitamins, etc.
- a carbon source examples include sugars such as glucose, sucrose, and fructose, ethanol, Alcohols such as methanol, organic acids such as citric acid, malic acid, and succinic acid, glycerol, and molasses are used.
- the nitrogen source for example, ammonia, ammonium sulfate, ammonium chloride, ammonium nitrate, urea or the like is used alone or in combination.
- inorganic salts examples include potassium monohydrogen phosphate, potassium dihydrogen phosphate, and magnesium sulfate.
- nutrients such as various vitamins such as peptone, meat extract, yeast extract, corn steep liquor, casamino acid, and biotin can be added to the medium.
- Cultivation is usually performed under aerobic conditions such as aeration agitation and shaking.
- the culture temperature is not particularly limited as long as the microorganism of the present invention can grow, and the pH during the culture is not particularly limited as long as the microorganism of the present invention can grow.
- the pH adjustment during the culture can be performed by adding an acid or an alkali.
- the target organic compound in the culture solution after completion of the culture is obtained by removing insoluble components such as bacterial cells from the culture solution by centrifugation or the like, if necessary, followed by extraction with an organic solvent such as ethyl acetate, activated carbon, etc.
- the target organic compound can be collected by using a method such as an ion exchange resin, a crystallization method, a precipitation method such as salting out, a distillation method or the like alone or in combination.
- ATCC13032 strain that efficiently produces DHS as compound P as the microorganism of the present invention and the method for producing DHS using the microorganism will be specifically described with reference to Examples.
- examples of the construction of a strain of ATCC13032 strain that efficiently produces protocatechuic acid as compound Q as the microorganism of the present invention and the production method of protocatechuic acid by the microorganism will be specifically described with reference to Examples. The present invention is not limited to these examples.
- Example 1 Comparison of DHS productivity between Pben-vanR strain and NSH ⁇ aroE3 ⁇ qsuB ⁇ qsuD Phen-vanR strain (Reference Example 11) in which the expression of the aroE3 gene is suppressed by the VanR repressor, and the ability to convert DHS to shikimate and DHS by in-frame deletion mutation of the qsuB, qsuD and aroE3 genes NSH ⁇ aroE3 ⁇ qsuB ⁇ qsuD strain (Reference Example 10) that lost the ability to convert protocatechuic acid into protocatechuic acid was cultured in a 3 L jar fermenter, and DHS productivity test was performed. These strains have higher DHS production than wild strains because the transcription of four genes (aroF, aroG, aroB, and aroD genes) involved in the synthesis of DHS from DAHP is controlled by the Tu promoter. is doing.
- Pre-culture and pre-culture media for these strains include CGXII media (ammonium sulfate 20 g / L, urea 5 g / L, potassium dihydrogen phosphate 1 g / L, dipotassium hydrogen phosphate 1 g / L, Magnesium sulfate heptahydrate 0.25 g / L, calcium chloride 10 mg / L, ferrous sulfate heptahydrate 10 mg / L, manganese sulfate pentahydrate 10 mg / L, zinc sulfate 7 water Japanese 1mg / L, Copper sulfate 0.2mg / L, Nickel chloride hexahydrate 0.02mg / L, Biotin 0.2mg / L, Glucose 20g / L, Protocatechuic acid 30mg / L, pH7.0) Using.
- CGXII media ammonium sulfate 20 g / L, urea
- CGCF medium which is a CGXII medium containing 2.1 g / L citric anhydride and 1.0 g / L iron sulfate / 7H 2 O, was used.
- DHS and protocatechuic acid were performed using this method unless otherwise specified.
- NSH ⁇ aroE3 ⁇ qsuB ⁇ qsuD strain is extremely poor in CGXII medium and CGCF medium, so 0.5 g / L tryptophan, 0.4 g / L phenylalanine, 0.45 g / L tyrosine, 17 mg / L shikimic acid, 45 mg / L vitamin K2 and 14 mg / L paraaminobenzoic acid were added to these media and cultured.
- After starting the main culture collect the culture medium in a timely manner, dilute it 1000 times with 5% aqueous acetonitrile, and analyze it under the separation conditions shown in Table 1 using high performance liquid chromatography (Waters, LCT TM Premier TM XE). Was calculated based on the ratio of the DHS peak area with a DHS aqueous solution (DHS purchased from Sigma) as a control.
- DHS DHS aqueous solution
- the DHS production of the Pben-vanR strain and the NSH ⁇ aroE3 ⁇ qsuB ⁇ qsuD strain was maximized at 86 hours and 62 hours after the start of the culture, respectively, and the maximum DHS production of these strains was 15.4 g / L and 7.7 ⁇ g / L, respectively.
- the Pben-vanR strain that can artificially control the expression of the aroE gene with a VanR repressor is a medium that does not contain aromatic amino acids as compared to the NSH ⁇ aroE3 ⁇ qsuB ⁇ qsuD strain that is cultured in a medium that contains aromatic amino acids. It became clear that DHS productivity was excellent even when cultured in the medium.
- Example 2 Protocatechuic acid production using NSH ⁇ aroE3 and NSH ⁇ aroE3_vanE3 strains
- NSH ⁇ aroE3 strain having an in-frame deletion mutation in the aroE3 gene see Reference Example 7
- VanR repressor based on the strain The NSH ⁇ aroE3_vanE3 strain (see Reference Example 8) modified so as to express the aroE3 gene was cultured in a 3 L jar fermenter and the production amount of protocatechuic acid was compared.
- the culture solution obtained by culturing these strains in CGXII medium was diluted with CGXII medium so that the absorbance at 600 nm was 3.0, and then inoculated into CGCF medium (0.9 L) in a 3 L jar fermenter by 1%.
- the NSH ⁇ aroE3 strain has a very poor growth in CGXII medium and CGCF medium due to deletion mutation of aroE3 gene, so 50 mg / L tryptophan, 50 mg / L phenylalanine, 50 mg / L tyrosine and 17 mg / L L shikimic acid was added to these media and cultured.
- glucose was added at a rate of 2.0 ⁇ g / hour and further cultured for 62.5 hours.
- the culture solution is collected in a timely manner, and the culture solution is diluted 1000 times with 5% acetonitrile aqueous solution and analyzed under the separation conditions shown in Table 1 using high performance liquid chromatography (Water, LCT TM Premier TM XE).
- the protocatechuic acid aqueous solution with a known concentration was purchased from Wako Pure Chemical Industries, Ltd. was used as a control to calculate the ratio of the protocatechuic acid peak area.
- quantification of procatechuic acid was measured using this method.
- the protocatechuic acid production of the NSH ⁇ aroE3 and NSH ⁇ aroE3_vanE3 strains was maximized at 48 hours and 62.5 hours after the start of culture, respectively, and the maximum production of protocatechuic acid of these strains was 0.5 g / L and 16.6 g / L, respectively. Met. From this result, when a strain that controls the expression of the aroE gene with a VanR repressor is cultured in a medium that does not contain aromatic amino acids and shikimic acid, protocatechuic acid of NSH ⁇ aroE3 strain cultured in a medium that contains aromatic amino acids and shikimic acid It was revealed that protocatechuic acid was produced approximately 33 times as much as the production amount.
- Example 3 Production of Protocatechuic Acid Using NSH ⁇ aroE3_vanE3 Strain, NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB Strain and NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB-vanR Strain of NsuS 3 L jars of NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB strain (Reference Example 12) and NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB-vanR strain (Reference Example 13) bred so that expression of qsuB gene and vanR gene can be induced by addition of benzoic acid
- the production of protocatechuic acid was compared by culturing with a fermenter.
- the qsuB gene was induced by adding 5 mM final concentration of benzoic acid at 14.5 hours after the start of the main culture.
- the qsuB and vanR genes were induced by adding 5 mM final concentration of benzoic acid.
- glucose was added at a rate of 3.5 g / hour from the start of the main culture at 14.5 hours, 15.5 hours, and 18.5 hours, respectively. The culture was terminated after 62 hours.
- Example 4 Protocatechuic acid production using NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB-vanR and tkt Culturing the above-described NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB-vanR strain (Reference Example 13) and the tkt strain (Reference Example 14) with enhanced transketolase expression of the strain using 3 L jar fermenters as follows: The production of protocatechuic acid was compared.
- FIG. Production of protocatechuic acid using control system with RhcR repressor All the above examples were production experiments in which the expression of aroE3 gene was controlled by VanR repressor. Difference in protocatechuic acid productivity by the presence or absence of transcription induction of nagI promoter using Pnag-qsuB-rhcR strain (Reference Example 17) that controls expression and expresses qsuB and rhcR genes under the control of nagI promoter was examined as follows. When inducing transcription of the nagI promoter of the Pnag-qsuB-rhcR strain, 3-hydroxybenzoic acid was added to 5 mM at 16.5 hours after the start of the main culture, and glucose was added at 4.9 hours at 20.5 hours.
- the mixture was added at a rate of g / hour, and the culture was terminated after 62.5 hours from the start of the culture.
- the transcription of the nagI promoter of the Pnag-qsuB-rhcR strain was not induced, the culture was terminated 62.5 hours after the start of the culture without adding 3-hydroxybenzoic acid. as a result,.
- the production of protocatechuic acid in the Pnag-qsuB-rhcR strain that did not induce the transcription of the nagI promoter was 47.5 hours and 43.5 hours after the start of culture, respectively.
- the maximum production of protocatechuic acid was 11.2 g / L and 8.4 g / L, respectively. From this result, even when the expression of the aroE3 gene and the qsuB gene is controlled by the RhcR repressor, a significant amount of protocatechuic acid can be produced. It became clear that
- Reference Example 1 Construction of shuttle vector pHCG298 for Escherichia coli-corynebacterium bacteria Corynebacterium glutamicum ATCC13058 carrying plasmid pHM1519 was obtained from NITE, and plasmid pHM1519 was isolated by a conventional method. Subsequently, two types of DNA (SEQ ID NOs: 1 and 2) were used to add a restriction enzyme BglII site and an NcoI site to the pHSG298 vector for Escherichia coli carrying the kanamycin resistance gene functioning in the ATCC13032 strain (manufactured by Takara Bio Inc.). Was synthesized.
- Bacillus subtilis encoding a levans classase that functions lethally to the host cell in order to select a strain in which the target DNA region has been integrated into a specific region of the chromosome by double crossover homologous recombination 168 strains of sacB gene were used as selection markers.
- Bacillus subtilis 168 strain was obtained as IAM2118 strain from RIKEN Microbial System Storage Facility.
- genome sequence information (GB accession number: NC — 000964) of Bacillus subtilis 168 strain was obtained from NCBI via the Internet.
- Chromosomal DNA of Bacillus subtilis 168 strain was purified using a chromosomal DNA extraction kit (RBC Bioscience).
- telomere sequence containing the sacB gene.
- an A residue was added to the 3 ′ end of the amplified DNA fragment using Taq DNA polymerase.
- the amplified DNA fragment was recovered after gel electrophoresis, purified, and then incorporated into a pT7Blue-T vector to construct a plasmid pTBSACB1 carrying the sacB gene.
- the sacB gene A constitutive expression plasmid pHKPsacB1 was constructed.
- the plasmid pHKPsacB1 carries DNA having high homology with the chromosomal DNA of Corynebacterium, homologous recombination occurs between the chromosomal DNA and the plasmid, and the entire plasmid is incorporated into the chromosome.
- the resulting strain exhibits a kanamycin resistant and sucrose sensitive phenotype. Thereafter, when crossover homologous recombination occurs twice, the plasmid-derived DNA region is excluded from the chromosome. As a result, the bacterium exhibits a kanamycin sensitive and sucrose resistant phenotype.
- Reference Example 3 Production of HT23 strain in which the modification / restriction enzyme gene of ATCC13032 strain is disrupted Construction of Plasmid for DNA Modification / Restriction Enzyme Gene Disruption HT23 strain in which DNA modification / restriction enzyme gene was disrupted was constructed based on ATCC13032 strain as follows. ATCC13032 strain was obtained from NITE as NBRC12168 strain. The base numbers shown in the following examples are the base numbers of the genome sequence of the ATCC13032 strain, and the genome sequence information was obtained from the NCBI GB database as the accession number NC_006958 via the Internet.
- culture of Corynebacterium bacteria including ATCC13032 strain is CGYE medium (ammonium sulfate 20 g / L, urea 5 g / L, KH2PO4 1 g / L, K2HPO4 1 g / L, MgSO4 ⁇ 7H20 0.25 g / L, yeast extract 1 g / L, CaCl2 10 mg / L, FeSO4 ⁇ 7H2O 10 mg / L, MnSO4 ⁇ 5H2O 10 mg / L, ZnSO4 ⁇ 7H2O 1 mg / L, CuSO4 0.2 mg / L, NiCl2 ⁇ 6H2O 0.02 mg / L, biotin 0.2 mg / L, glucose 20 g / L, pH 7).
- CGYE medium ammonium sulfate 20 g / L, urea 5 g / L, KH2PO4 1 g / L, K2HPO4 1 g
- CglIM (cg1996) gene (base number 1,879,784 to 1,880,875) encoding the DNA modifying enzyme possessed by ATCC13032 strain
- cglIR cg1997) gene (base number 1,880,884 to 1,881,960)
- cglIIR cg1998) gene encoding two restriction enzymes
- the 5 'flanking region was amplified by PCR using these primers with the chromosomal DNA of ATCC13032 strain as a template. After adding an A residue to the 3 ′ end of the amplified DNA fragment with Taq DNA polymerase, the amplified DNA fragment is purified by gel electrophoresis and incorporated into the pT7Blue-T vector to retain the 5 ′ flanking region. Plasmid pTUSR1F was constructed.
- an about 1.1 kb BamHI-XhoI fragment containing the 5 'flanking region was prepared from the plasmid pTUSR1F. Further, an XbaI-XhoI fragment of about 1.4 kb containing the 3 'flanking region was prepared from the plasmid pTUSR1R. Further, an approximately 1.6 kb BamHI-SphI fragment containing a sacB gene constitutive expression region was prepared from plasmid pHKPsacB1 (Reference Example 2-2). These three fragments were inserted between the BamHI and XbaI sites of the cloning vector pHSG298 for Escherichia coli to construct a plasmid pHKSD0977-9 for DNA modification / restriction enzyme gene disruption.
- SCR strain is cultured in 1 mL LB liquid medium (10 g / l tryptone, 5 g / l yeast extract, 10 g / l sodium chloride) for 24 hours, and a part of the culture solution is LB agar containing 10% sucrose.
- HT23 strain was obtained by smear culture on the medium. It was confirmed by PCR using the primers of SEQ ID NOs: 13 and 14 that the HT23 strain was a double-crossover homologous recombinant lacking the DNA modification / restriction enzyme gene as expected.
- Reference Example 4 Construction of a pcaH ⁇ pcaG gene-disrupted strain
- ATCC13032 A DRHG strain lacking a part of the chromosomal DNA of the strain (complementary strand base numbers 2,511,382 to 2,512,700) was constructed.
- the amplified DNA fragment is purified by gel electrophoresis and incorporated into a pT7Blue-T vector (manufactured by Novagen). Plasmid pTPCAUSR1F with a 5 ′ flanking region was constructed.
- two types of primers SEQ ID NOs: 19 and 20 for amplifying the 3 ′ flanking region (complementary strand base numbers 2,510,015 to 2,511,377) of HT23 strain were synthesized, and purified chromosomal DNA of HT23 strain was used as a template.
- the 3 ′ flanking region was amplified by PCR using these primers. After adding an A residue to the 3 ′ end of the amplified DNA fragment by Taq DNA polymerase, the amplified DNA fragment is purified by gel electrophoresis, and incorporated into the pT7Blue-T vector, whereby a plasmid having the 3 ′ flanking region is obtained.
- pTPCADSR1R was constructed. An approximately 1.3 kb EcoRI-MluI fragment containing the 5 ′ flanking region was prepared from pTPCAUSR1F by restriction enzyme digestion. In addition, an approximately 1.4 kb MluI-BamHI fragment containing the 3 ′ flanking region was prepared from pTPCADSR1R by restriction enzyme digestion.
- SCRHG strain was cultured in 1 mL of LB liquid medium for 24 hours, and a part of the culture solution was smeared on LB agar medium containing 10% sucrose to obtain DRHG strain. It was confirmed by PCR using the primer sets of SEQ ID NOS: 23 and 24 that the DRHG strain was a double-crossover homologous recombinant lacking the pcaH / pcaG gene as expected. In addition, when PCR reaction using the primers of SEQ ID NOs: 25 and 26 was performed, an amplified DNA fragment was not obtained, so it was also confirmed that the DRHG strain was deficient in the sacB gene.
- Reference Example 5 Construction of a pobB gene disrupted strain Based on the DRHG strain constructed in Reference Example 4 in order to disrupt the pobB gene encoding parahydroxybenzoate hydroxylase and suppress the production of gallic acid from protocatechuic acid, A DRHG145 strain lacking a part of DNA (complementary strand base numbers 1,126,301 to 1,127,488b) was constructed as follows.
- the 5 ′ flanking region (base numbers 1,127,505 to 1,128,687) of the pobB gene and add a restriction enzyme XbaI site and a SalI site at both ends of the amplified fragment, 30) was synthesized.
- the 5 ′ flanking region of the pobB gene was amplified by PCR using these primers.
- the amplified DNA fragment in the 3 ′ flanking region is digested with restriction enzymes SphI and XbaI
- the amplified DNA fragment in the 5 ′ flanking region is digested with restriction enzymes XbaI and SalI. 2 was introduced between the SphI site and the SalI site in the multicloning site to obtain a plasmid pHKPsacB ⁇ pobB.
- DRHG_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a part of the culture solution was smeared on LB agar medium containing 10% sucrose to obtain DRHG145 strain. It was confirmed by PCR using the primers of SEQ ID NOs: 27 and 30 that the DRHG145 strain was a double-crossover homologous recombinant lacking the pobB gene as expected.
- two types of primers (SEQ ID NOs: 35 and 36) were synthesized in order to add a restriction enzyme RsrII site and a PfoI site at both ends of the amplified fragment.
- the 5 ′ flanking region of the cg0500 gene, the 3 ′ flanking region of the promoter region of the aro operon, and the Tu promoter were amplified by PCR using these three primer sets.
- the amplified DNA fragment in the 5 'flanking region of the cg0500 gene is digested with restriction enzymes SphI and RsrII
- the amplified DNA fragment in the 3' flanking region of the promoter region of the aro operon is digested with restriction enzymes PfoI and SbfI
- Tu After digesting the amplified DNA fragment of the promoter with restriction enzymes RsrII and PfoI, SbfI and SphI sites in the multiple cloning site of plasmid pHKPsacB1 (see Reference Example 2) are obtained from these three restriction fragments.
- the plasmid pHKPsacB_ aro-Ntu was obtained.
- pHKPsacB_Ntu-aro is introduced into DRHG145 strain and selected for resistance to kanamycin to obtain DRHG145_Km strain did.
- the DRHG145_Km strain was analyzed by PCR using the primers of SEQ ID NOs: 31 and 34 and the primer for confirming the sacB gene (SEQ ID NOs: 3 and 4), the expected results were obtained. Therefore, the DRHG145_Km strain was isolated from the plasmid pHKPsacB_Ntu-aro. Was confirmed to be a single crossover homologous recombinant introduced into the cg0500 gene region.
- the DRHG145_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a portion of the culture solution was smeared on an LB agar medium containing 10% sucrose to obtain the NUA strain.
- NDSGU strain in which promoter of aroG gene was replaced with Tu promoter region was constructed based on NUA strain as follows.
- (1) Construction of aroG promoter replacement plasmid Amplifies the 5 'flanking region of the aroG gene promoter (complementary strand base numbers 2,280,842 to 2,282,206) and adds restriction enzyme SphI and RsrII sites to both ends of the amplified fragment
- Two kinds of primers (SEQ ID NOs: 38 and 39) were synthesized.
- the NDS_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a part of the culture solution was smeared on an LB agar medium containing 10% sucrose to obtain an NDSGU strain.
- the NDSGU strain was a double crossover homologous recombinant in which the aroG promoter region was replaced with the Tu promoter as expected. did.
- a DNA primer (SEQ ID NO: 42) outside the 5 'flanking region of the aroG gene and a DNA primer (SEQ ID NO: 43) outside the 3' flanking region were synthesized, and the aroG promoter was also obtained by PCR using these primers. It was confirmed that the region was a double crossover homologous recombinant in which the Tu promoter was replaced.
- NSU strain in which promoter of aroB gene was replaced with Tu promoter region was constructed based on NDSGU strain as follows.
- (1) Construction of aroB promoter replacement plasmid In order to amplify the 5 'flanking region (complementary strand base numbers 1,720,573 to 1,721,670) of the aroB gene promoter and add restriction enzyme SphI and RsrII sites to both ends of the amplified fragment, two primers (SEQ ID NO: 44 and 45) was synthesized.
- the NDSGU_Km strain was analyzed by PCR using the chromosomal DNA of the NDSGU_Km strain as a template, the primers of SEQ ID NOs: 48 and 49, the primers of SEQ ID NOs: 35 and 36, and the sacB gene confirmation primer (SEQ ID NOs: 3 and 4). From the obtained results, it was confirmed that the NDSGU_Km strain was a single crossover homologous recombinant in which the plasmid pHKPsacB_aroB-Ntu was introduced into the aroB gene region.
- the NDSGU_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a part of the culture solution was smeared on an LB agar medium containing 10% sucrose to obtain an NSU strain.
- the double cross-over type in which the aroB promoter region is replaced with the Tu promoter as expected in the NSU strain by PCR using the primers of SEQ ID NOs: 44 and 47, the primers of SEQ ID NOs: 48 and 49, and the primers of SEQ ID NOs: 35 and 36 It was confirmed to be a homologous recombinant.
- NSH strain in which promoter of aroF gene is replaced with Tu promoter NSH strain in which the promoter region of aroF gene encoding DHQ dehydrase is replaced with Tu promoter region was constructed based on NSU strain as follows.
- (1) Construction of aroF promoter replacement plasmid In order to amplify the 5 ′ flanking region (complementary strand base numbers 1,046,619 to 1,047,863) of the promoter of the aroF gene and to add restriction enzyme SphI site and RsrII site to both ends of the amplified fragment, two kinds of primers (SEQ ID NO: 50 and 51) was synthesized.
- the NSU_Km strain was analyzed by PCR using the chromosomal DNA of the NSU_Km strain as a template, the primers of SEQ ID NOs: 54 and 55, the primers of SEQ ID NOs: 35 and 36, and the sacB gene confirmation primer (SEQ ID NOs: 3 and 4). From the obtained results, it was confirmed that the NSU_Km strain was a single crossover homologous recombinant in which the plasmid pHKPsacB_aroF-Ntu was introduced into the aroF gene region.
- NSU_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a part of the culture solution was smeared on LB agar medium containing 10% sucrose to obtain NSH strain.
- the double cross-over type in which the aroB promoter region is replaced with the Tu promoter as expected in the NSU strain by PCR using the primers of SEQ ID NOs: 50 and 53, the primers of SEQ ID NOs: 54 and 55, and the primers of SEQ ID NOs: 35 and 36 It was confirmed to be a homologous recombinant.
- in-frame disruption strains of aroE1 and aroE3 genes encoding shikimate dehydrogenase Among the three types of shikimate dehydrogenase possessed by ATCC13032, the aroE1 and aroE3 genes that may affect growth
- a strain into which an in-frame deletion mutation was introduced (a strain into which an in-frame deletion mutation was introduced (hereinafter also referred to as an in-frame disruption strain) was constructed as follows. Mutation, i.e., in-frame disruption, refers to a mutation in which a deletion having a length of a multiple of 3 bases is introduced into the translation region.
- a digestion reaction was performed with restriction enzymes SphI and SbfI.
- the resulting DNA fragment was introduced between the restriction enzyme sites SbfI-SphI in the multiple cloning site of plasmid pHKPsacB1 (see Reference Example 2) to obtain plasmid pHKPsacB ⁇ aroE1.
- the plasmid pHKPsacB ⁇ aroE1 has a structure in which the region of complementary strand base numbers 1,181,390 to 1,184,160 is deleted from the aroE1 gene region.
- NSH_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a part of the culture solution was smeared on LB agar medium containing 10% sucrose to obtain NSH ⁇ aroE1 strain. It was confirmed by PCR using the primers of SEQ ID NOs: 60 and 61 that the NSH ⁇ aroE1 strain was a double-crossover homologous recombinant lacking the aroE1 gene as expected.
- SEQ ID NOs: 64 and 65 Two types of DNA primers (SEQ ID NOs: 64 and 65) were synthesized in order to amplify the 3 ′ flanking region (base numbers 1,725,101 to 1,726,094) of the aroE3 gene.
- a restriction enzyme SbfI site was introduced into the primer of SEQ ID NO: 65.
- the amplification reaction was performed by the overlapping PCR method using the primers of SEQ ID NOs: 62 and 65. Then, a digestion reaction with restriction enzymes SphI and SbfI was performed.
- the resulting DNA fragment was introduced between the restriction enzyme SbfI site and the SphI site in the multiple cloning site of plasmid pHKPsacB1 (see Reference Example 2) to obtain plasmid pHKPsacB ⁇ aroE3.
- the plasmid pHKPsacB ⁇ aroE3 has a structure in which the region of complementary strand base numbers 1,726,095 to 1,726,886 is deleted from the aroE3 gene region.
- the NSH_Km strain was analyzed by the PCR method using the primers of SEQ ID NOs: 62 and 65, the primers of SEQ ID NOs: 66 and 67, and the primer for confirming the sacB gene (SEQ ID NOs: 3 and 4), an expected result was obtained. From the results, it was confirmed that the NSH_Km2 strain was a single crossover type homologous recombinant in which the plasmid pHKPsacB ⁇ aroE3 was introduced into the aroE3 gene region.
- NSH_Km2 strain was cultured in 1 mL of LB liquid medium for 24 hours, and a part of the culture solution was smeared on LB agar medium containing 10% sucrose to obtain NSH ⁇ aroE3 strain. It was confirmed by PCR using the primers of SEQ ID NOs: 66 and 67 that the NSH ⁇ aroE3 strain was a double-crossover homologous recombinant lacking the aroE3 gene as expected.
- NSH ⁇ aroE3 and NSH ⁇ aroE1 ⁇ aroE3 strains were found to recover their growth when shikimic acid, tryptophan, fanylalanine and tyrosine were added to CGXII medium, and the main gene encoding shikimate dehydrogenase was confirmed to be the aroE3 gene. .
- Reference Example 8 Construction of protocatechuic acid producing strain NSH ⁇ aroE3_vanE3 expressing aroE3 gene under the control of vanA promoter
- the gene encoding vanillate demethylase of ATCC13032 strain is cg2616 (hereinafter abbreviated as vanA gene.
- vanA gene Base number 2,496,775-2,497,905 bp: size 1,131 bp
- cg2617 hereinafter abbreviated as vanB gene, base numbers 2,497,909 to 2,498,886 bp: size 978 bp).
- vanAB gene region base numbers 2,497,909 to 2,498,886 bp
- the repressor that controls vanA and vanB genes is known as cg2615 (hereinafter abbreviated as vanR gene; complementary strand base numbers 2,496,013 to 2,496,591 bp: size 579 bp).
- vanR gene complementary strand base numbers 2,496,013 to 2,496,591 bp: size 579 bp.
- the aroE3 gene was introduced into the vanA gene and vanB gene regions by the vanR gene, and the aroE3 gene was replaced under the control of the vanR gene.
- the parent strain the NSH ⁇ aroE3 strain shown in Reference Example 7 was used.
- a replacement vector was constructed by the following procedure in order to replace the vanAB gene region of the ATCC13032 strain with the aroE3 gene.
- the 5 ′ flanking region of the vanAB gene is amplified and the restriction enzyme site SphI is introduced into the DNA primer of SEQ ID NO: 68, and the 3 ′ flanking region of the vanAB gene is amplified and the restriction enzyme site SbfI is added to the DNA primer of SEQ ID NO: 71.
- Two kinds of DNA primers (SEQ ID NO: 72 and SEQ ID NO: 73) were synthesized to amplify the aroE3 gene.
- DNA primers (SEQ ID NO: 74 and SEQ ID NO: 75) were prepared 112 bp outside the 5 ′ flanking region of the vanAB gene and 106 bp outside the 3 ′ flanking region, and the 5 ′ flanking region and 3 ′ flanking region of the vanAB gene were prepared. PCR was performed using the DNA primers (SEQ ID NO: 68 and SEQ ID NO: 71) used to amplify the ranking region, and using a colony of a kanamycin resistant strain of the strain NSH ⁇ aroE3 directly as a template.
- the amplified DNA fragment obtained by PCR was 2,905 bp.
- an amplified DNA fragment obtained by PCR amplified with the DNA primers of SEQ ID NO: 74 and SEQ ID NO: 71 is 2,913 bp
- an amplified DNA fragment obtained by PCR amplified with the DNA primers of SEQ ID NO: 68 and SEQ ID NO: 75 is 4,186 bp. Met.
- the sacB gene region was confirmed by PCR analysis or the like, it was found that the strain NSH ⁇ aroE3 was a single cross-type homologous recombinant.
- the strain NSH ⁇ aroE3 once cross-type homologous recombinant was cultured in 1 mL of LB liquid medium for 24 hours, and a small amount was smeared on LB agar medium containing 10% sucrose.
- the chromosomal DNA retained by the grown strain NSH ⁇ aroE3 was predicted to replace the sacB gene, kanamycin resistance gene, and vanAB gene region with the aroE3 gene, and was named NSH ⁇ aroE3_vanE3 strain.
- strain NSH ⁇ aroE3_vanE3 17.5 mg / L shikimic acid, 50 mg / L tryptophan, 50 mg / L fanylalanine, 50 mg / L tyrosine, 10 mg / L parahydroxybenzoic acid and 10 mg / L
- the viability tests of strains NSH ⁇ aroE3_vanE3 and NSH ⁇ aroE3 were carried out using CGXII medium added with paraaminobenzoic acid (hereinafter referred to as 6 kinds of additives) and CGXII medium without addition of 6 kinds of additives.
- the strain NSH ⁇ aroE3_vanE3 grew normally in the former medium, but the growth rate to the middle of the culture was slow in the latter medium.
- the strain NSH ⁇ aroE3 grew on the former medium but did not grow on the latter medium. It was also found that when the strain NSH ⁇ aroE3_vanE3 was cultured by adding 50 ⁇ M or more of ferulic acid, vanillic acid or vanillin to the CGXII medium without the addition of six kinds of additives, it was found that the cells grew normally.
- the 5 ′ flanking region and 3 ′ flanking region of the qsuB gene were amplified by PCR using these primers. After digesting the 5 ′ flanking region amplification product with restriction enzymes SbfI and XbaI and digesting the 3 ′ flanking region amplification product with restriction enzymes XbaI and SalI, these DNA fragments were ligated to plasmid pHKPsacB1 (see Reference Example 2). The plasmid was introduced between the SbfI site and the SalI site in the multiple cloning site to obtain pHKPsacB ⁇ qsuB, a plasmid for disrupting the qsuB gene.
- a DNA primer (SEQ ID NO: 80) that matches the sequence upstream of the 5 ′ flanking region of the qsuB gene and a DNA primer (SEQ ID NO: 80) that matches the sequence downstream of the 3 ′ flanking region ( SEQ ID NO: 81) was synthesized and analyzed by PCR using the primers of SEQ ID NOs: 79 and 80 and primers of SEQ ID NOs: 76 and 81. Analysis by PCR using any primer set gave the expected results, confirming that these strains are single-crossover homologous recombinants with the plasmid pHKPsacB ⁇ qsuB introduced into the qsuB gene region. .
- NSH ⁇ qsuB_Km strain, NSH ⁇ aroE3 ⁇ qsuB_Km strain, and NSH ⁇ aroE3_vanE3 ⁇ qsuB_Km strain are each cultured in 1 ⁇ mL of LB liquid medium for 24 hours, and a part of the culture solution is smeared on LB agar medium containing 10% sucrose. Strain, NSH ⁇ aroE3 ⁇ qsuB strain, and NSH ⁇ aroE3_vanE3 ⁇ qsuB strain were obtained.
- the resulting DNA fragment was introduced between the restriction enzyme sites SbfI-SalI in the multiple cloning site of plasmid pHKPsacB1 (see Reference Example 2) to obtain plasmid pHKPsacB ⁇ qsuD.
- the plasmid pHKPsacB ⁇ qsuD has a structure in which the region of base numbers 446,554 to 447,384 is deleted from the qsuD gene region.
- NSH ⁇ aroE3 ⁇ qsuB_Km and NSH ⁇ aroE3_vanE3 ⁇ qsuB_km strains are the desired single-crossover homologous recombinants, a part upstream of the qsuD gene 5 ′ flanking region and a downstream of the 3 ′ flanking region Two types of primers (SEQ ID NOs: 86 and 87) based on a part were synthesized.
- NSH ⁇ aroE3 ⁇ qsuB_Km strain and NSH ⁇ aroE3_vanE3 ⁇ qsuB_km strain were cultured in 1 ⁇ mL of LB liquid medium for 24 hours, and a part of the culture was smeared on LB agar medium containing 10% sucrose to obtain NSH ⁇ aroE3 ⁇ qsuB ⁇ qsuD strain and NSH ⁇ aroE3 ⁇ BuSqD .
- NSH ⁇ aroE3 ⁇ qsuB ⁇ aroE strain and NSH ⁇ aroE3_vanE3 ⁇ qsuB ⁇ qsuD strain were confirmed to be double-crossover homologous recombinants lacking the aroE gene as expected, by PCR using the primers of SEQ ID NOs: 86 and 87.
- NSH ⁇ aroE3 ⁇ qsuB ⁇ qsuD strain aromatic amino acid requirement test NSH strain, NSH ⁇ aroE3 strain, NSH ⁇ aroE3 ⁇ qsuB strain, NSH ⁇ aroE3 ⁇ qsuB ⁇ aroE1 strain, NSH ⁇ aroE3_vanE3 ⁇ qsuB strain, NSH ⁇ aroE3_vanE3 ⁇ qsuB strain, NSH ⁇ aroE3 ⁇ qsuBq A test was conducted.
- Reference Example 11 Construction of a DHS producing strain capable of controlling the expression of aroE3 gene by inducible expression of VanR repressor
- vanR repressor By replacing the benABCD gene region of NSH ⁇ aroE3_vanE3 ⁇ qsuB ⁇ qsuD strain constructed in Reference Example 10 with vanR gene, the vanR gene is expressed under the control of benA promoter A strain was established.
- SEQ ID NOs: 88 and 89 Two DNA primers (SEQ ID NOs: 88 and 89) were synthesized to amplify the 5 'flanking region (base numbers 2,515,963 to 2,516,969) of the benABCD gene region.
- SEQ ID NOs: 90 and 91 two types of DNA primers (SEQ ID NOs: 90 and 91) were synthesized in order to amplify the 3 ′ flanking region (base numbers 2,521,372 to 2,522,498) of the benABCD gene region.
- the primers of SEQ ID NOs: 88 and 91 have a restriction enzyme SalI site and an SbfI site, respectively.
- the 5 ′ flanking region and 3 ′ flanking region of the benABCD gene were amplified by PCR using these primers.
- two types of DNA primers (SEQ ID NOs: 92 and 93) were synthesized to amplify the vanR gene.
- the vanR gene region was amplified by PCR using the chromosomal DNA of the HT23 strain as a template and the primers of SEQ ID NOs: 92 and 93.
- an amplification reaction was performed by the overlap PCR method using the primers of SEQ ID NOs: 88 and 93. Thereafter, a DNA fragment cleaved at the SalI site in the primer of SEQ ID NO: 88 and the NarI site inside the vanR gene was obtained by digestion with restriction enzymes SalI and NarI.
- the amplification reaction was performed by the overlap PCR method using primers of SEQ ID NOs: 91 and 92 using a mixture of the amplified DNA of the vanR gene region and the amplified DNA of the 3 ′ flanking region of the benABCD gene as a template.
- NarI and SbfI a DNA fragment cleaved at the NarI site in the vanR gene and the SbfI site in the primer of SEQ ID NO: 91 was obtained.
- These two types of DNA fragments were introduced between the SalI and SbfI sites in the multiple cloning site of plasmid pHKPsacB1 (see Reference Example 2) to obtain plasmid pHKPsacB_Pben-vanR.
- the primer (SEQ ID NO: 94) matches the sequence upstream of the 5 ′ flanking region of the vanR gene and matches the sequence downstream of the 3 ′ flanking region. After synthesizing the primer (SEQ ID NO: 95), it was analyzed by PCR using the primers of SEQ ID NOs: 94 and 91 and the primers of SEQ ID NOs: 88 and 95.
- NSH ⁇ aroE3_vanE3 ⁇ qsuB ⁇ qsuD strain_Km strain is a single-crossover homologous recombinant in which the plasmid pHKPsacB_Pben-vanR has been introduced into the benABCD gene region. I confirmed that there was.
- the NSH ⁇ aroE3_vanE3 ⁇ qsuB ⁇ qsuD strain_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a portion of the culture solution was smeared on an LB agar medium containing 10% sucrose to obtain a Pben-vanR strain.
- Pben-vanR strain was a double crossover homologous recombinant in which the benABCD gene region was replaced with the vanR gene.
- a primer (SEQ ID NO: 98) matching the sequence upstream of the 5 ′ flanking region of the qsuB gene and a primer matching the sequence downstream of the 3 ′ flanking region ( SEQ ID NO: 99) was synthesized and analyzed by PCR using the primers of SEQ ID NO: 88 and 99 and the primers of SEQ ID NOs: 91 and 98.
- NSH ⁇ aroE3_vanE3 ⁇ qsuB_Km strain is a single-crossover homologous recombinant in which the plasmid pHKPsacB_Pben-qsuB has been introduced into the benABCD gene region, because the PCR method using any primer set gave the expected results. It was confirmed.
- the NSH ⁇ aroE3_vanE3 ⁇ qsuB_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a portion of the culture solution was smeared on an LB agar medium containing 10% sucrose to obtain an NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB strain. It was confirmed by PCR using the primers of SEQ ID NOs: 88 and 99 and the primers of SEQ ID NOs: 91 and 98 that the benABCD gene region was a double crossover homologous recombinant in which the qsuB gene was substituted.
- SEQ ID NOs: 100 and 101 Two types of DNA primers (SEQ ID NOs: 100 and 101) were used to amplify the 5 'flanking region (base numbers 2,520,333 to 2,521,374) of the vanR gene insertion region.
- SEQ ID NOs: 102 and 103 Two types of DNA primers (SEQ ID NOs: 102 and 103) were synthesized.
- the primers of SEQ ID NOs: 100 and 103 have a restriction enzyme SalI site and an SbfI site, respectively.
- the 5 ′ flanking region and 3 ′ flanking region of the vanR gene insertion region were amplified by PCR using these primers.
- two types of DNA primers SEQ ID NOs: 104 and 105 were synthesized in order to amplify the vanR gene, and the vanR gene region was obtained by PCR using the chromosomal DNA of the HT23 strain (Reference Example 3) as a template. Amplified.
- the amplification reaction was performed by the overlap PCR method using the primers of SEQ ID NOs: 100 and 105. Thereafter, digestion with restriction enzymes SalI and BamHI gave a DNA fragment cleaved at the SalI site in the primer of SEQ ID NO: 100 and the BamHI site inside the vanR gene.
- an amplification reaction was performed by an overlap PCR method using primers of SEQ ID NOs: 104 and 103, using a mixture of the amplified DNA of the vanR gene region and the amplified DNA of the 3 ′ flanking region of the vanR gene insertion region as a template, By digestion with restriction enzymes BamHI and SbfI, a DNA fragment cleaved at the BamHI site inside the vanR gene and the SbfI site in the primer of SEQ ID NO: 103 was obtained.
- the NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB strain_Km strain is a one-time crossover type in which the plasmid pHKPsacB_Pben-qsuB-vanR was introduced into the vanR gene insertion region. It was confirmed to be a homologous recombinant.
- NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB strain_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a part of the culture was smeared on LB agar medium containing 10% sucrose to obtain NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB-vanR strain . It was confirmed by PCR using the primers of SEQ ID NOs: 106 and 107 that the homologous recombinant was a two-fold crossover type in which the vanR gene was inserted downstream of the qsuB gene in the benABCD gene region.
- Reference Example 14 Construction of a protocatechuic acid-producing bacterium tkt strain that constitutively expresses the tkt gene
- the NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB-vanR strain constructed in Reference Example 13 is the parent strain, and the tkt gene involved in the synthesis of erythrose-4-phosphate is the Tu promoter (Reference Example)
- a tkt strain in which the transcription unit linked to 6) hereinafter abbreviated as Ptu-tkt transcription unit was inserted into the DNA modification / restriction enzyme gene disruption site was constructed as follows.
- Reference Example HT23 strain (Reference Example 3) chromosomal DNA was used as a template to amplify the 5 'flanking region and 3' flanking region of the DNA modification / restriction enzyme gene disruption site by PCR using these primers.
- the obtained 5 ′ flanking region amplified DNA and tkt gene amplified DNA were used as templates, followed by an amplification reaction by the overlapping PCR method using primers of SEQ ID NOs: 108 and 113, and then by restriction enzymes SbfI and PfoI. By digestion, a DNA fragment cleaved at the SbfI site in the primer of SEQ ID NO: 108 and the PfoI site in the primer of SEQ ID NO: 113 was obtained.
- an amplification reaction was performed by the overlapping PCR method using primers of SEQ ID NOs: 36 and 111 using a mixture of amplified DNA of the Tu promoter and amplified DNA of the 3 ′ flanking region as a template, and then the restriction enzymes Pfo and SalI were used. By digestion, a DNA fragment cleaved at the PfoI site in the primer of SEQ ID NO: 36 and the SalI site in the primer of SEQ ID NO: 111 was obtained.
- the primer (SEQ ID NO: 115) matches the sequence upstream of the 5 ′ flanking region and the sequence downstream of the 3 ′ flanking region.
- the primer (SEQ ID NO: 116) to be synthesized was synthesized and analyzed by PCR using the primers of SEQ ID NOs: 115 and 116 and the primers of SEQ ID NOs: 108 and 111.
- NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB-vanR strain_Km strain is a single crossover homologous recombinant into which the plasmid pHKPsacB_Ptu-tkt has been introduced. I confirmed that there was.
- the NSH ⁇ aroE3_vanE3 ⁇ qsuB_Pben-qsuB-vanR strain_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a part of the culture solution was smeared on an LB agar medium containing 10% sucrose to obtain a tkt strain.
- a Ptu-tkt transcription unit was incorporated into the DNA modification / restriction enzyme gene disruption site by PCR using the primers of SEQ ID NOs: 115 and 116 and the primers of SEQ ID NOs: 108 and 111 and restriction enzyme digestion of the amplified fragment 2 It was confirmed to be a homologous recombination with round crossover type.
- Reference Example 15 Construction of a strain expressing the aroE3 gene under the control of the rhcH promoter A strain from which the region from the cg1309 gene to the cg1311 gene (hereinafter abbreviated as rhcHMD gene region) was replaced with the aroE3 gene was constructed as follows.
- the amplified DNA fragment in the 5 'flanking region is digested with restriction enzymes KpnI and SbfI, and the amplified DNA fragment in the 3' flanking region is digested with restriction enzymes SbfI and SalI, and then these DNA fragments are converted into plasmid pHKPsacB1 (Reference Example). 2) between the KpnI site and the SalI site in the multiple cloning site to obtain plasmid pHKPsacB ⁇ genH.
- the NSH ⁇ aroE3_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a part of the culture solution was smeared on an LB agar medium containing 10% sucrose to obtain an NSH ⁇ aroE3 ⁇ genH strain. It was confirmed by PCR using the primers of SEQ ID NOs: 117 and 120 that the NSH ⁇ aroE3 ⁇ genH strain was a double-crossover homologous recombinant lacking the genH gene as expected.
- SEQ ID NO: 123 and SEQ ID NO: 124 were synthesized to amplify the 3 ′ flanking region (base numbers 1,218,296 to 1,219,545) of the rhcHMD gene.
- a restriction enzyme SalI site was introduced into the primer of SEQ ID NO: 124.
- DNA primers of SEQ ID NO: 125 and SEQ ID NO: 126 were synthesized in order to amplify the aroE3 gene.
- amplification was performed by overlap PCR using DNA fragments (SEQ ID NO: 125 and SEQ ID NO: 124) using a DNA fragment obtained by amplifying the 3 'flanking region and aroE3 gene as a template.
- the amplified DNA fragments were digested with SbfI and AvrII and AvrII and SalI, and introduced between the restriction enzyme sites SbfI-SalI in the multiple cloning site of plasmid pHKPsacB1 (see Reference Example 2) to obtain plasmid pHKPsacB_rhcR_Prhc-rhcE3.
- the NSH ⁇ aroE3 ⁇ genH_Km strain was analyzed by PCR using the DNA primers of SEQ ID NO: 127 and SEQ ID NO: 128, the primers of SEQ ID NO: 121 and SEQ ID NO: 124, and the sacB gene confirmation primer (SEQ ID NOS: 3 and 4), the expected results were obtained.
- the NSH ⁇ aroE3 ⁇ genH_Km strain was confirmed to be a single crossover homologous recombinant in which the plasmid pHKPsacB_rhcR_Prhc-rhcE3 was introduced into the rhcHMD gene region.
- NSH ⁇ aroE3 ⁇ genH_rhcE3 strain is obtained by culturing a single cross-type homologous recombinant of NSH ⁇ aroE3 ⁇ genH strain in 1 ⁇ mL LB liquid medium for 24 hours and smearing a part of the culture on LB agar medium containing 10% sucrose. It was. By the PCR method using the primers of SEQ ID NOs: 127 and 128, it was confirmed that the rhcHMD gene region of the NSH ⁇ aroE3 ⁇ genH_rhcE3 strain was replaced with the aroE3 gene and was a double-crossover homologous recombinant as expected.
- Each strain was inoculated into CGXII medium as a pre-culture medium, cultured at 30 ° C. overnight, washed twice, and planted in the main culture medium CGXII medium so that the absorbance at 600 nm was about 0.1.
- CGXII medium a culture experiment in which shikimic acid (17.5 mg / L), tryptophan (50 mg / L), fanylalanine (50 mg / L) and tyrosine (50 mg / L) were added to CGXII medium was also performed in parallel. As a result, it was revealed that the NSH strain grew normally on the CGXII medium, but the NSH ⁇ aroE3 strain grew very slowly.
- NSH ⁇ aroE3_vanE3 strain and the NSH ⁇ aroE3 ⁇ genH_rhcE3 strain were delayed in growth from the beginning to the middle of the culture and recovered in the end of the culture.
- the growth of NSH ⁇ aroE3_vanE3 strain or NSH ⁇ aroE3 ⁇ genH_rhcE3 strain was recovered when shikimic acid, tryptophan, fanylalanine and tyrosine were added to CGXII medium. From these results, it was found that both the VanR repressor and the RhcR repressor can control the expression of the aroE3 gene.
- Reference Example 16 Construction of a strain that expresses the rhcR gene under the control of the nagI promoter
- the strain that expresses the rhcR gene under the control of the nagI promoter is as follows: It was constructed as follows.
- SEQ ID NO: 129 and SEQ ID NO: 130 Two types of DNA primers (SEQ ID NO: 129 and SEQ ID NO: 130) were synthesized in order to amplify the 5 'flanking region (complementary strand base numbers 3,199,996 to 3,200,904) of the nagIKL gene operon region.
- a restriction enzyme SbfI site was introduced into the primer of SEQ ID NO: 129.
- the 5 'flanking region of the nagIKL gene operon region was amplified by PCR using these primers.
- SEQ ID NO: 131 and SEQ ID NO: 132 Two types of DNA primers (SEQ ID NO: 131 and SEQ ID NO: 132) were synthesized in order to amplify the 3 'flanking region (complementary strand base numbers 3,196,321 to 3,197,306) of the nagIKL gene operon region.
- a restriction enzyme SalI site was introduced into the primer of SEQ ID NO: 132.
- DNA primers of SEQ ID NO: 133 and SEQ ID NO: 134 were synthesized in order to amplify the rhcR gene.
- amplification was performed by overlap PCR using DNA fragments (SEQ ID NO: 132 and SEQ ID NO: 133) using a DNA fragment obtained by amplifying the 3 'flanking region and rhcR gene as a template.
- the amplified DNA fragments were digested with SbfI and AvrII and AvrII and SalI, and introduced between the restriction enzyme sites SbfI-SalI in the multiple cloning site of plasmid pHKPsacB1 (see Reference Example 2) to obtain plasmid pHKPsacB_Pnag-rhcR.
- the NSH ⁇ aroE3 ⁇ genH_rhcE3_Km strain was analyzed by the PCR method using the DNA primers of SEQ ID NO: 135 and SEQ ID NO: 136, the primers of SEQ ID NO: 129 and SEQ ID NO: 132, and primers for confirming the sacB gene (SEQ ID NOs: 3 and 4), the expected results were obtained.
- the NSH ⁇ aroE3 ⁇ genH_rhcE3_Km strain was confirmed to be a single crossover homologous recombinant in which the plasmid pHKPsacB_Pnag-rhcR was introduced into the operon region of the nagIKL gene.
- the Pnag-rhcR strain was obtained by culturing a single cross-type homologous recombinant of NSH ⁇ aroE3 ⁇ genH_rhcE3 strain in 1 mL of LB liquid medium for 24 hours and smearing a part of the culture on LB agar medium containing 10% sucrose. Got.
- the nagIKL gene region of the Pnag-rhcR strain was replaced with the rhcR gene, and it was confirmed that it was a double-crossover homologous recombinant as expected.
- the primers of SEQ ID NOs: 137 and 140 have a restriction enzyme SbfI site and a SphI site, respectively.
- SbfI site a restriction enzyme
- SphI site a restriction enzyme
- the 5 ′ flanking region and the 3 ′ flanking region of the qsuB gene introduction region were amplified by PCR using these primers.
- two types of DNA primers SEQ ID NOs: 141 and 142
- SEQ ID NOs: 141 and 142 were synthesized to amplify the qsuB gene, and the qsuB gene region was determined by PCR using the chromosomal DNA of the HT23 strain (Reference Example 3) as a template. Amplified.
- an amplification reaction was performed by the overlap PCR method using the primers of SEQ ID NOs: 137 and 142. Thereafter, digestion with restriction enzymes SbfI and XhoI yielded a DNA fragment cleaved at the SbfI site in the primer of SEQ ID NO: 137 and the XhoI site inside the qsuB gene.
- an amplification reaction was performed by an overlap PCR method using primers of SEQ ID NOs: 140 and 141, using a mixture of the amplified DNA of the qsuB gene region and the amplified DNA of the 3 ′ flanking region of the qsuB gene introduction region as a template, Digestion with restriction enzymes XhoI and SphI yielded a DNA fragment cleaved at the XhoI site within the qsuB gene and the SphI site within the primer of SEQ ID NO: 140.
- the Pnag-rhcR strain_Km strain was cultured in 1 mL of LB liquid medium for 24 hours, and a portion of the culture was smeared on an LB agar medium containing 10% sucrose to obtain the Pnag-qsuB-rhcR strain. . It was confirmed by the PCR method using the primers of SEQ ID NOs: 143 and 144 that the qsuB gene was a double crossover homologous recombinant introduced downstream of the nagI promoter.
Landscapes
- Chemical & Material Sciences (AREA)
- Organic Chemistry (AREA)
- Life Sciences & Earth Sciences (AREA)
- Engineering & Computer Science (AREA)
- Wood Science & Technology (AREA)
- Zoology (AREA)
- Health & Medical Sciences (AREA)
- Genetics & Genomics (AREA)
- Bioinformatics & Cheminformatics (AREA)
- Biotechnology (AREA)
- General Engineering & Computer Science (AREA)
- Microbiology (AREA)
- General Health & Medical Sciences (AREA)
- Biochemistry (AREA)
- General Chemical & Material Sciences (AREA)
- Chemical Kinetics & Catalysis (AREA)
- Biomedical Technology (AREA)
- Molecular Biology (AREA)
- Medicinal Chemistry (AREA)
- Biophysics (AREA)
- Plant Pathology (AREA)
- Physics & Mathematics (AREA)
- Tropical Medicine & Parasitology (AREA)
- Virology (AREA)
- Preparation Of Compounds By Using Micro-Organisms (AREA)
- Micro-Organisms Or Cultivation Processes Thereof (AREA)
Description
(1)炭素源から生育に必須な代謝物Mを生成する生合成経路において中間代謝物である化合物Pを代謝物Mに転換する酵素Xの発現量を調節することにより化合物Pを蓄積させるために、以下の(a)から(d)に記載の性質をすべて有する原核生物。
(a)該原核生物が利用できる炭素源から化合物Pに至る生合成に関わる酵素群のうち、いずれか1つ以上の酵素の活性が該原核生物の野生株と比べて増強している。
(b)酵素Xのタンパク質をコードする野生型遺伝子xの翻訳領域、該遺伝子の翻訳調節領域または遺伝子xの転写を促すプロモーター領域の中に1個以上の塩基の置換、欠失または付加による変異を有することにより、酵素Xの活性が欠損または低下している。
(c)前記遺伝子xの転写を促す本来のプロモーターとは異なっており、かつリプレッサーRのタンパク質により転写が抑制されるプロモーターAにより、活性型酵素XをコードするDNAの転写が制御される。
(d)リプレッサーRのタンパク質をコードする遺伝子zを1コピー以上有し、該遺伝子の転写が該遺伝子の本来のプロモーターおよび/または該プロモーターとは異なる誘導プロモーターBにより制御される。
(3)前記原核生物が有する化合物Pを代謝する酵素のうち、酵素X以外のいずれか1つ以上の代謝酵素の活性が欠損または低下していることを特徴とする、(1)または(2)に記載の原核生物。
(4)前記プロモーターBが、フェルラ酸、バニリン酸、バニリン、安息香酸、3-ヒドロキシ安息香酸、レゾルシノール、4-ヒドロキシ安息香酸、2,4-ジヒドロキシ安息香酸、フラクトースおよびスクロースからなる群から選ばれる化合物の添加により誘導されるプロモーターであることを特徴とする、(1)から(3)のいずれか一項に記載の原核生物。
(6)前記プロモーターAが、コリネバクテリウム・グルタミカムATCC13032株のvanA(cg2616)遺伝子のプロモーター、pobA(cg1226)遺伝子のプロモーター、pcaH(cg2631)遺伝子のプロモーター、またはrhcH(cg1309)遺伝子のプロモーターであることを特徴とする、(1)から(5)のいずれか一項に記載の原核生物。
(8)(1)から(6)のいずれか一項に記載の原核生物をプロモーターAの転写を抑制した状態で炭素源の存在下で培養した後、プロモーターBの転写を誘導する処理を加えることによりリプレッサーRのタンパク質の発現量を増加させることにより、代謝物Mの生成量を減少させることを特徴とする、化合物Pの製造方法。
(9)(1)から(6)のいずれか一項に記載の原核生物をプロモーターAの転写抑制を解除した状態で炭素源の存在下で培養した後、プロモーターBの転写を誘導する処理を加えることによりリプレッサーRのタンパク質の発現量を増加させることにより、代謝物Mの生成量を減少させることを特徴とする、化合物Pの製造方法。
(f)化合物Pが3-デヒドロシキミ酸であり、代謝物Mがシキミ酸である。
(g)化合物Pがグルタミン酸であり、代謝物MがN-アセチルグルタミン酸またはγーグルタミルリン酸である。
(h)化合物Pがアスパラギン酸であり、代謝物Mがβ-アルパルチルリン酸である。
(i)化合物Pがセリンであり、代謝物Mがグリシンである。
(e)酵素Yのタンパク質をコードする遺伝子yの翻訳領域、該遺伝子の翻訳調節領域または遺伝子yの転写を促すプロモーター領域の中に1個以上の塩基の置換、欠失または付加による変異を有することにより、化合物Pから化合物Qへの転換能が増強している、もしくは遺伝子yの発現が野生型とは異なる制御を受けている。
(13)性質(e)に記載の遺伝子yの転写を促すプロモーター領域の中の置換変異が、該プロモーターの全域または一部の領域を、該遺伝子の転写を促す本来のプロモーターおよびプロモーターAとは異なるプロモーターCのDNAと置換する変異であることを特徴とする、(11)または(12)に記載の原核生物。
(15)前記プロモーターCが誘導プロモーターであることを特徴とする、(13)または(14)に記載の原核生物。
(16)前記プロモーターCが、フェルラ酸、バニリン酸、バニリン、安息香酸、3-ヒドロキシ安息香酸、レゾルシノール、4-ヒドロキシ安息香酸、2,4-ジヒドロキシ安息香酸、フラクトースおよびスクロースからなる群から選ばれる化合物の添加により誘導されるプロモーターであることを特徴とする、(15)に記載の原核生物。
(18)(11)から(16)のいずれか一項に記載の原核生物をプロモーターAの転写を抑制した状態で炭素源の存在下で培養した後、プロモーターBの転写を誘導する処理を加えることによりリプレッサーRのタンパク質の発現量を増加させることにより、代謝物Mの生成量を減少させ、化合物Qの生成量を増加させることを特徴とする、化合物Qの製造方法。
(20)(11)から(16)のいずれか一項に記載の原核生物を炭素源の存在下で培養した後、前記プロモーターCの転写を誘導する処理を加えることにより前記酵素Yの発現量を増加させることにより、化合物Qの生成量を増加させることを特徴とする、(17)から(19)のいずれか一項に記載の化合物Qの製造方法。
(j)化合物Pが3-デヒドロシキミ酸であり、化合物Qがプロトカテク酸であり、代謝物Mがシキミ酸である。
(k)化合物Pがコリスミン酸であり、化合物Qがアントラニル酸であり、代謝物Mがプレフェン酸である。
(l)化合物Pがプレフェン酸であり、化合物Qが4-ヒドロキシフェニルピルビン酸であり、代謝物Mがフェニルピルビン酸である。
(m)化合物Pがプレフェン酸であり、化合物Qがアロゲン酸であり、代謝物Mがフェニルピルビン酸である。
(n)化合物Pがプレフェン酸であり、化合物Qがフェニルピルビン酸であり、代謝物Mが4-ヒドロキシフェニルピルビン酸またはアロゲン酸である。
(o)化合物Pが2-オキソイソ吉草酸であり、化合物Qが2-イソプロピルリンゴ酸であ、代謝物Mがバリンである。
(p)化合物Pが2-オキソイソ吉草酸であり、化合物Qがバリンであり、代謝物Mが2-イソプロピルリンゴ酸である。
(q)化合物Pがグルタミン酸であり、化合物Qがγーグルタミルリン酸であり、代謝物MがN-アセチルグルタミン酸である。
(r)化合物Pがグルタミン酸であり、化合物QがN-アセチルグルタミン酸であり、代謝物Mがγーグルタミルリン酸である。
(s)化合物Pがアスパラギン酸であり、化合物Qがアスパラギンであり、代謝物Mがβ-アルパルチルリン酸である。
(t)化合物Pがアスパラギン酸β-セミアルデヒドあり、化合物Qが2,3-ジヒドロジピコリン酸であり、代謝物Mがホモセリンである。
(u)化合物Pがホモセリンあり、化合物QがO-アセチルホモセリンであり、代謝物Mがホモセリンリン酸である。
(v)化合物Pがホモセリンであり、化合物Qがホモセリンリン酸であり、代謝物MがO-アセチルホモセリンである。
(w)化合物Pがセリンであり、化合物QがO-アセチルセリンであり、代謝物Mがグリシンである。
(23)プロトカテク酸2,3-ジオキシゲナーゼをコードする遺伝子、プロトカテク酸3,4-ジオキシゲナーゼをコードする遺伝子、プロトカテク酸4,5-ジオキシゲナーゼをコードする遺伝子およびプロトカテク酸脱炭酸酵素をコードする遺伝子からなる群から選ばれる少なくとも1つの遺伝子について、該遺伝子の翻訳領域またはその転写・翻訳調節領域の中に、1個以上の塩基の置換、欠失または付加による変異を導入することにより、プロトカテク酸の代謝活性が欠損または低下している性質であることを特徴とする、(22)に記載のプロトカテク酸の製造方法。
(24)コリネバクテリウム・グルタミカムまたはエッシェリヒア・コリである、(1)から(6)のいずれか一項または(11)から(16)のいずれか一項に記載の原核生物。
本発明の微生物は、炭素源から生育に必須な代謝物Mに至る生合成経路上の中間代謝物である目的の化合物Pを効率よく製造するために用いる微生物であり、以下に示す4つの性質(a)~(d)を有する。該微生物は、遺伝子発現制御のためにリプレッサーRによる転写抑制系が発達している原核生物であることが好ましい。
(a)該微生物が利用できる炭素源から化合物Pに至る生合成に関わる酵素群のうち、いずれか1つ以上の酵素の活性が該原核生物の野生株と比べて増強している。
(b)化合物Pを代謝物Mに転換する酵素Xのタンパク質をコードする野生型遺伝子xの翻訳領域、該遺伝子の翻訳調節領域または遺伝子xの転写を促すプロモーター領域の中に1個以上の塩基の置換、欠失または付加による変異を有することにより、酵素Xの活性が欠損または低下している。
(c)前記遺伝子xの転写を促す本来のプロモーターとは異なっており、かつリプレッサーRのタンパク質により転写が抑制されるプロモーターAにより、活性型酵素XをコードするDNAの転写が制御される。
(d)リプレッサーRのタンパク質をコードする遺伝子zを1コピー以上有し、該遺伝子の転写が該遺伝子の本来のプロモーターおよび/または該プロモーターとは異なる誘導プロモーターBにより制御される。
(aa)該菌株が利用できる炭素源からDHS(化合物P)に至る生合成に関わる酵素群のうち、いずれか1つ以上の酵素の活性が野生株と比べて増強している。
(bb)DHSをシキミ酸(代謝物M)に転換するシキミ酸脱水素酵素(酵素X)のタンパク質をコードする野生型遺伝子(遺伝子x)の翻訳領域または翻訳調節領域、もしくは該遺伝子の転写を促すプロモーター領域の中に、1個以上の塩基の置換、欠失または付加による変異を有することにより、DHSからシキミ酸への転換能が欠損または低下している。ATCC13032株は、活性型シキミ酸脱水素酵素をコードする遺伝子として、aroE3(cg1835)遺伝子(相補鎖塩基番号1,726,078~1,726,908のDNA領域)、aroE1(cg1283)遺伝子(相補鎖塩基番号1,182,337~1,183,143のDNA領域)およびqsuD(cg0504)遺伝子(塩基番号446,537~447,388のDNA領域)の3種類を有しているが、DHSからシキミ酸への変換に主要な役割を果たしているaroE3遺伝子に変異を導入し、該遺伝子の活性型シキミ酸脱水素酵素の発現が欠損あるいは低下することによっても性質bbを付与することができる。さらに、aroE3遺伝子の変異に加えて、qsuD遺伝子および/またはaroE1遺伝子についても該遺伝子の活性型シキミ酸脱水素酵素の発現が欠損あるいは低下する変異を導入することにより、性質bbを付与してもよい。
(cc)前記遺伝子xの転写を促す本来のプロモーターとは異なっており、かつリプレッサーRのタンパク質により転写が抑制されるプロモーターAにより、活性型酵素XをコードするDNAの転写が制御される。
(dd)リプレッサーRのタンパク質をコードする遺伝子zを1コピー以上有し、該遺伝子の転写が該遺伝子の本来のプロモーターおよび/または該プロモーターとは異なる誘導プロモーターBにより制御される。
(e)化合物Pから化合物Qに転換する酵素Yのタンパク質をコードする遺伝子yの翻訳領域、該遺伝子の翻訳調節領域または遺伝子yの転写を促すプロモーター領域の中に1個以上の塩基の置換、欠失または付加による変異を有することにより、化合物Pから化合物Qへの転換能が増強している、もしくは遺伝子yの発現が野生型とは異なる制御を受けている。
(ee)DHSからプロトカテク酸に転換するDHS脱水酵素(酵素Y)のタンパク質をコードするqsuB遺伝子(遺伝子y;塩基番号444,184~446,040)の転写を促す本来のプロモーターとは異なり、かつプロモーターAとも異なるプロモーターCによりqsuB遺伝子の転写が制御される。
以下のようにして、qsuB遺伝子のインフレーム欠失変異によりDHSからプロトカテク酸への転換能を失い、benAプロモーターの制御下にあるvanR遺伝子がコードするVanRリプレッサーによりaroE3遺伝子の発現が抑制されるPben-vanR株(参考例11)、およびqsuB遺伝子とqsuD遺伝子とaroE3遺伝子のインフレーム欠失変異により、DHSからシキミ酸への転換能とDHSからプロトカテク酸への転換能を失ったNSHΔaroE3ΔqsuBΔqsuD株(参考例10)を3 Lジャーファーメンターで培養し、DHSの生産性試験を行った。なお、これらの菌株は、DAHPからDHSの合成に関わる4種類の遺伝子(aroF、aroG、aroBおよびaroD遺伝子)の転写がTuプロモーターによって制御されるために、野生株と比べてDHS生成量が向上している。
以下のようにして、aroE3遺伝子にインフレーム欠失変異を有するNSHΔaroE3株(参考例7参照)、ならびに該菌株をもとにVanRリプレッサーの制御下でaroE3遺伝子を発現するように改変したNSHΔaroE3_vanE3株(参考例8参照)を、3 Lジャーファーメンターで培養してプロトカテク酸の生産量を比較した。
以下のようにして、上述のNSHΔaroE3_vanE3株(参考例8)、qsuB遺伝子の発現を安息香酸の添加により誘導できるように育種したNSHΔaroE3_vanE3ΔqsuB_Pben-qsuB株(参考例12)、ならびにqsuB遺伝子およびvanR遺伝子の発現を安息香酸の添加により誘導できるように育種したNSHΔaroE3_vanE3ΔqsuB_Pben-qsuB-vanR株(参考例13)を、それぞれ3 Lジャーファーメンターを用いて培養することによりプロトカテク酸の生産量を比較した。NSHΔaroE3_vanE3ΔqsuB_Pben-qsuB株については本培養開始後14.5時間の時点で安息香酸を終濃度5 mM添加することによりqsuB遺伝子を誘導し、またNSHΔaroE3_vanE3ΔqsuB_Pben-qsuB-vanR株については本培養開始後15.5時間の時点で安息香酸を終濃度5 mM添加することによりqsuB遺伝子とvanR遺伝子を誘導した。NSHΔaroE3_vanE3ΔqsuB_Pben-qsuB株、NSHΔaroE3_vanE3ΔqsuB_Pben-qsuB-vanR株、およびNSHΔaroE3_vanE3株については、それぞれ本培養開始後14.5時間、15.5時間および18.5時間の時点でグルコースを3.5 g/時間の割合で添加し、本培養開始より62時間後に培養を終了した。その結果、NSHΔaroE3_vanE3株、NSHΔaroE3_vanE3ΔqsuB_Pben-qsuB株およびNSHΔaroE3_vanE3ΔqsuB_Pben-qsuB-vanR株のプロトカテク酸の生産量はそれぞれ培養開始後43.8時間目と43.8時間目と38.5時間目に最大化し、これら菌株のプロトカテク酸の最大生産量はそれぞれ8.5 g/L、8.6 g/L、15.2 g/Lであった。この結果から、VanRリプレッサーの制御下でaroE3遺伝子を発現する菌株は著量のプロトカテク酸を蓄積できるとともに、VanRリプレッサー量を増加させてaroE3遺伝子の発現抑制を強化した方が、プロトカテク酸の生産性が高いことが明らかになった。
以下のようにして、上述のNSHΔaroE3_vanE3ΔqsuB_Pben-qsuB-vanR株(参考例13)および該菌株のトランスケトラーゼ発現を強化したtkt株(参考例14)をそれぞれ3 Lジャーファーメンターを用いて培養することによりプロトカテク酸の生産量を比較した。それぞれの菌株ともに、本培養開始後17時間の時点で安息香酸を終濃度5 mM添加し、また本培養開始後24時間の時点でグルコースを4.5 g/時間の割合で添加し、培養開始から64時間後に培養を終了した。その結果、NSHΔaroE3_vanE3ΔqsuB_Pben-qsuB-vanR株とtkt株のプロトカテク酸の生産量はそれぞれ培養開始後40時間目と45時間目に最大化し、これら菌株のプロトカテク酸の最大生産量はそれぞれ14.8 g/L、20.0 g/Lであった。この結果から、VanRリプレッサーによるaroE遺伝子の発現制御に加えて、トランスケトラーゼ発現を増強すると、プロトカテク酸の生産性が向上することが明らかになった。
上述の実施例はすべてVanRリプレッサーによりaroE3遺伝子の発現を制御した生産実験であったが、VanRリプレッサーの代わりにRhcRリプレッサーによりaroE3遺伝子の発現を制御し、かつnagIプロモーターの制御下でqsuB遺伝子とrhcR遺伝子を発現するPnag-qsuB-rhcR株(参考例17)を用いて、nagIプロモーターの転写誘導の有無によるプロトカテク酸の生産性の差を、以下のようにして調べた。Pnag-qsuB-rhcR株のnagIプロモーターの転写を誘導する場合には、本培養開始後16.5時間の時点で3-ヒドロキシ安息香酸を5 mMになるように添加し、20.5時間の時点でグルコースを4.9 g/時間の割合で添加し、培養開始から62.5時間後に培養を終了した。Pnag-qsuB-rhcR株のnagIプロモーターの転写を誘導しない場合には、3-ヒドロキシ安息香酸を添加せずに、培養開始から62.5時間後に培養を終了した。その結果、。その結果、nagIプロモーターの転写誘導を行ったPnag-qsuB-rhcR株のnagIプロモーターの転写誘導を行わないPnag-qsuB-rhcR株のプロトカテク酸の生産量はそれぞれ培養開始後47.5時間目と43.5時間目に最大化し、それぞれのプロトカテク酸の最大生産量は11.2 g/L、8.4 g/Lであった。この結果より、RhcRリプレッサーによってaroE3遺伝子とqsuB遺伝子の発現を制御した場合でも著量のプロトカテク酸が生産できるとともに、培養後期にRhcRリプレッサーの量を増強した方がプロトカテク酸の生産性が高くなることが明らかになった。
プラスミドpHM1519を保持するコリネバクテリウム・グルタミカムATCC13058株は、NITEより入手し、プラスミドpHM1519を常法により単離した。続いて、ATCC13032株中で機能するカナマイシン耐性遺伝子を保持する大腸菌用ベクターpHSG298(タカラバイオ株式会社製)に制限酵素BglII部位とNcoI部位を付加するために2種類のDNA(配列番号1と2)を合成した。これらDNAをpHSG298のKpnI部位とPstI部位の間に挿入し、プラスミドpHSG298BNを構築した。続いて、プラスミドpHM1519から、プラスミド複製領域を含む約3.1 kbのBglII断片をBglII消化により切り出し、pHSG298BNのBglII部位に挿入することにより、プラスミドpHCG100を構築した。このpHCG100を制限酵素BglIIとNcoIによって消化した後、Blunting Highキット(東洋紡績株式会社製)を用いて制限酵素切断末端を平滑末端に変えた。pHCG100由来の約1.9 kbのBglII(平滑末端)-NcoI(平滑末端)断片を精製した後、pHSG298のStuI部位に挿入することにより、大腸菌-コリネバクテリウム用シャトル・ベクターpHCG298を構築した。
コリネバクテリウム属細菌において2回交差型相同組換えにより目的DNA領域を染色体の特定領域に組み込むときに用いるレバンスクラーゼ遺伝子を構成的に発現するプラスミドpHKPsacB1を以下のようにして構築した。
2回交差型相同組換えにより目的DNA領域が染色体の特定領域に組み込まれた株を選別するため、宿主細胞に対して致死的に機能するレバンスクラ-ゼをコードするバチルス・サブチリス168株のsacB遺伝子を選別マーカーとして用いることにした。バチルス・サブチリス168株は、理化学研究所微生物系統保存施設よりIAM2118株として入手した。また、バチルス・サブチリス168株のゲノム配列情報(GBアクセッション番号:NC_000964)をNCBIよりインターネット経由で取得した。バチルス・サブチリス168株の染色体DNAを染色体DNA 抽出キット(RBC Bioscience社製)を用いて精製した。この染色体DNA(100 ng)を鋳型として、2種類のDNAプライマー(配列番号3と4)を用いてPCR反応を行うことにより、sacB遺伝子を含む増幅DNA断片を得た。続いて、Taq DNAポリメラーゼを用いて増幅DNA断片の3’末端へA残基を付加した。増幅DNA断片をゲル電気泳動後に回収し、精製した後、pT7Blue-Tベクターに組み込むことにより、sacB遺伝子を保持するプラスミドpTBSACB1を構築した。
配列番号5~12で表わされる8本の合成DNAを合成した後に、これら合成DNAをpT7Blue-Tベクターに組み込むことにより、sacB遺伝子を構成的に発現するプラスミドpTRKD2を構築した。続いて、上記1で取得したプラスミドpTBSACB1上のsacB遺伝子領域を制限酵素HindIIIとKpnIならびに制限酵素KpnIとXbaIを用いて消化することより、sacB遺伝子領域から由来する1.0 kbのHindIII-KpnI断片と0.4 kbのKpnI-XbaI断片を得た。これら2つのDNA断片を、上で取得したpTRKD2由来の0.2 kbのHindIII-SphI断片ともに、大腸菌用クローニングベクターpHSG298(タカラバイオ社製)のSphI部位とXbaI部位の間に挿入することにより、sacB遺伝子構成的発現プラスミドpHKPsacB1を構築した。
プラスミドpHKPsacB1は、コリネバクテリウム属細菌の染色体DNAと相同性が高いDNAを運ぶときには、染色体DNAとプラスミドの間で相同組換えを起こり、プラスミド全体が染色体上に取り込まれる。そのために得られた菌株がカナマイシン耐性かつスクロース感受性の表現型を示す。その後、2回交差型相同組換えが起こると、プラスミド由来DNA領域が染色体上から排除される。その結果、当該細菌はカナマイシン感受性かつスクロース耐性の表現型を示す。
1.DNA修飾・制限酵素遺伝子破壊用プラスミドの構築
以下のようにして、ATCC13032株をもとに、DNA修飾・制限酵素遺伝子を破壊したHT23株を造成した。ATCC13032株は、NITEよりNBRC12168株として入手した。また、下記の実施例で示す塩基番号は、ATCC13032株のゲノム配列の塩基番号であり、該ゲノム配列情報については、NCBIのGBデータベースから、アクセッション番号NC_006958としてインターネット経由で取得した。また、ATCC13032株を含むコリネバクテリウム属細菌の培養は、特記しない限り、CGYE培地(硫酸アンモニウム 20 g/L、尿素 5 g/L、KH2PO4 1 g/L、K2HPO4 1 g/L、MgSO4・7H20 0.25 g/L、酵母エキス 1 g/L、CaCl2 10 mg/L、FeSO4・7H2O 10 mg/L、MnSO4・5H2O 10 mg/L、ZnSO4・7H2O 1 mg/L、CuSO4 0.2 mg/L、NiCl2・6H2O 0.02 mg/L、ビオチン 0.2 mg/L、グルコース 20 g/L、pH7)を用いて行った。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKSD0977-9をATCC13032株に導入し、カナマイシン耐性で選択することにより、SCR株を取得した。配列番号13と16のプライマーならびにsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりSCR株を解析したところ、予想通りの結果が得られたことから、SCR株はプラスミドpHKSD0977-9 がDNA修飾・制限酵素遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
以下のようにして、参考例3で造成したHT23株をもとに、プロトカテク酸4,5-ジオキシゲナーゼをコードするpcaH・pcaG遺伝子を欠損させるために、ATCC13032株の染色体DNAの一部(相補鎖塩基番号2,511,382~2,512,700)を欠失したDRHG株を造成した。
HT23株のpcaH・pcaG遺伝子の5’フランキング領域(塩基番号2,512,743~2,513,990)を増幅するための2種類のプライマー(配列番号17と18)を合成した。HT23株の染色体DNA(100 ng)を鋳型として、これらのプライマーを用いるPCR法により該5’フランキング領域を増幅した。続いて、Taq DNAポリメラーゼによって増幅DNA断片の3’末端にA残基を付加した後、増幅DNA断片をゲル電気泳動法により精製し、pT7Blue-Tベクター(ノバジェン社製)に組み込むことにより、該5’フランキング領域を持つプラスミドpTPCAUSR1Fを構築した。同様にして、HT23株の3’フランキング領域(相補鎖塩基番号2,510,015~2,511,377)を増幅するための2種類のプライマー(配列番号19と20)を合成し、HT23株の精製染色体DNAを鋳型としてこれらのプライマーを用いるPCR法により該3’フランキング領域を増幅した。Taq DNAポリメラーゼによって増幅DNA断片の3’末端にA残基を付加した後、増幅DNA断片をゲル電気泳動法により精製し、pT7Blue-Tベクターに組み込むことにより、該3’フランキング領域を持つプラスミドpTPCADSR1Rを構築した。制限酵素消化によりpTPCAUSR1Fから上記5’フランキング領域を含む約1.3 kbのEcoRI-MluI断片を調製した。また、制限酵素消化によりpTPCADSR1Rから上記3’フランキング領域を含む約1.4 kbのMluI-BamHI断片を調製した。さらに、制限酵素消化によりプラスミドpHKPsacB1(参考例2参照)からsacB遺伝子領域を含む約1.6 kbのBamHI-SphI断片を調製した後、これら3つの断片を大腸菌用クローニングベクターpHSG298のEcoRI部位とSphI部位の間に挿入し、遺伝子破壊用プラスミドpHKSDPHG1を構築した。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKSDPHG1をHT23株に導入し、カナマイシン耐性で選択することにより、SCRHG株を取得した。配列番号21と22のプライマーならびにsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりSCRHG株を解析したところ、予想通りの結果が得られたことから、SCRHG株はプラスミドpHKSDPHG1がpcaH・pcaG遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
パラヒドロキシ安息香酸ヒドロキシゲナーゼをコードするpobB遺伝子を破壊し、プロトカテク酸から没食子酸の生成を抑制するために、参考例4で造成したDRHG株をもとに、染色体DNAの一部(相補鎖塩基番号1,126,301~1,127,488b)を欠失したDRHG145株を以下のようにして造成した。
pobB遺伝子の3’フランキング領域を(塩基番号1,125,101~1,126,300)を増幅するとともに、増幅断片の両端に制限酵素SphI部位とXbaI部位を付加するために、2種類のDNAプライマー(配列番号27と28)を合成した。DRHG株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりpobB遺伝子の3’フランキング領域を増幅した。次に、pobB遺伝子の5’フランキング領域(塩基番号1,127,505~1,128,687)を増幅するとともに、増幅断片の両端に制限酵素XbaI部位とSalI部位を付加するために2種類のDNAプライマー(配列番号29と30)を合成した。DRHG株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりpobB遺伝子の5’フランキング領域を増幅した。3’フランキング領域の増幅DNA断片を制限酵素SphIとXbaIで消化し、また5’フランキング領域の増幅DNA断片を制限酵素XbaIとSalIで消化した後、これらのDNA断片をプラスミドpHKPsacB1(参考例2参照)のマルチクローニングサイト内にあるSphI部位とSalI部位の間に導入し、プラスミドpHKPsacBΔpobBを得た。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacBΔpobBをDRHG株に導入し、カナマイシン耐性で選択することにより、DRHG_Km株を取得した。配列番号27と30のプライマーならびにsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりDRHG_Km株を解析したところ、予想通りの結果が得られたことから、これら菌株はプラスミドpHKPsacBΔpobBがpobB遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
以下の4段階で、DAHPからDHSの合成に関わる4種類の遺伝子(aroF、aroG、aroBおよびaroD遺伝子)の転写を、ATCC13032株のTuプロモーター(塩基番号526,013~526,374)の制御下に置くことにより、DHS生成量を増強したNSH株を造成した。
ATCC13032株のqsuB遺伝子、aroD遺伝子、qsuD遺伝子はオペロン構造を形成しており(以下、aroオペロンという)、同一の制御因子(cg0500遺伝子産物)により、発現量が調節されている。参考例5で造成したDRHG145株をもとに、このaroオペロンのプロモーター領域(塩基番号441,597~442,756)を転写活性が強いTuプロモーターと置換したNUA株を以下にようにして造成した。
cg0500遺伝子の5’フランキング領域(塩基番号440,437~441,596)を増幅するともに、増幅断片の両端に制限酵素SphI部位とRsrII部位を付加するために2種類のプライマー(配列番号31と32)を合成した。aroプロモーターの3’フランキング領域(塩基番号442,757~443,916)を増幅するともに、増幅断片の両端に制限酵素PfoI部位とSbfI部位を付加するために2種類のプライマー(配列番号33と34)を合成した。また、Tuプロモーター領域を増幅するともに、増幅断片の両端に制限酵素RsrII部位とPfoI部位を付加するために2種類のプライマー(配列番号35と36)を合成した。DRHG株の染色体DNAを鋳型にして、これら3組のプライマーセットを用いるPCR法によりcg0500遺伝子の5’フランキング領域、aroオペロンのプロモーター領域の3’フランキング領域、およびTuプロモーターを増幅した。
エレクトロポレーションによる形質転換法を用いて、pHKPsacB_Ntu-aroをDRHG145株に導入し、カナマイシン耐性で選択することにより、DRHG145_Km株を取得した。配列番号31と34のプライマーならびにsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりDRHG145_Km株を解析したところ、予想通りの結果が得られたことから、DRHG145_Km株はプラスミドpHKPsacB_Ntu-aroがcg0500遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
DAHP合成酵素をコードするaroG遺伝子のプロモーター領域をTuプロモーター領域と置換したNDSGU株を以下のようにしてNUA株をもとに造成した。
(1)aroGプロモーター置換用プラスミドの構築
aroG遺伝子のプロモーターの5’フランキング領域(相補鎖塩基番号2,280,842~2,282,206)を増幅するとともに、増幅断片の両端に制限酵素SphI部位とRsrII部位を付加するために2種類のプライマー(配列番号38と39)を合成した。aroG遺伝子のプロモーターの3’フランキング領域(相補鎖塩基番号2,279,366~2,280,754)を増幅するとともに、増幅断片の両端に制限酵素PfoI部位とSbfI部位を付加するために2種類のプライマー(配列番号40と41)を合成した。また、DRHG株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりaroG遺伝子のプロモーターの5’フランキング領域と3’フランキング領域をそれぞれ増幅した。これとは別に、DRHG株の染色体DNAを鋳型にして配列番号35と36のプライマーを用いるPCR法によりTuプロモーターを増幅した。
エレクトロポレーションによる形質転換法を用いて、プラスミドpHKPsacB_aroG-NtuをNUA株に導入し、カナマイシン耐性で選択することにより、NUA_Km株を取得した。配列番号38と41のプライマーならびにsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりNUA_Km株を解析したところ、予想通りの結果が得られたことから、NUA_Km株はプラスミドpHKPsacB_aroG-NtuがaroG遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
DHQ合成酵素をコードするaroB遺伝子のプロモーター領域をTuプロモーター領域と置換したNSU株を以下のようにしてNDSGU株をもとに造成した。
(1)aroBプロモーター置換用プラスミドの構築
aroB遺伝子のプロモーターの5’フランキング領域(相補鎖塩基番号1,720,573~1,721,670)を増幅するとともに、増幅断片の両端に制限酵素SphI部位とRsrII部位を付加するために2種類のプライマー(配列番号44と45)を合成した。aroB遺伝子のプロモーターの3’フランキング領域(相補鎖塩基番号1,719,404~1,720,501)を増幅するとともに、増幅断片の両端に制限酵素PfoI部位とSbfI部位を付加するために2種類のプライマー(配列番号46と47)を合成した。また、DRHG株の染色体DNAを鋳型にして、これらのプライマーを用い、PCR法によりaroB遺伝子のプロモーターの5’フランキング領域と3’フランキング領域をそれぞれ増幅した。これとは別に、DRHG株の染色体DNAを鋳型にして配列番号35と16のプライマーを用いるPCR法によりTuプロモーターを増幅した。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacB_aroB-NtuをNDSGU株に導入して、カナマイシン耐性で選択することにより、NDSGU_Km株を取得した。得られたNDSGU_Km株が目的の1回交差型相同組換え体であることを確認するために、aroB遺伝子のプロモーターの5’フランキング領域の上流の一部と3’フランキング領域の下流の一部に基づく2種類のプライマー(配列番号48と49)を合成した。NDSGU_Km株の染色体DNAを鋳型として、配列番号48と49のプライマー、配列番号35と36のプライマーならびにsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりNDSGU_Km株を解析したところ、予想通りの結果が得られたことから、NDSGU_Km株はプラスミドpHKPsacB_aroB-NtuがaroB遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
DHQ脱水酵素をコードするaroF遺伝子のプロモーター領域をTuプロモーター領域と置換したNSH株を、以下のようにしてNSU株をもとに造成した。
(1)aroFプロモーター置換用プラスミドの構築
aroF遺伝子のプロモーターの5’フランキング領域(相補鎖塩基番号1,046,619~1,047,863)を増幅するとともに、増幅断片の両端に制限酵素SphI部位とRsrII部位を付加するために2種類のプライマー(配列番号50と51)を合成した。aroB遺伝子のプロモーターの3’フランキング領域(相補鎖塩基番号1,046,619~1,047,863)を増幅するとともに、増幅断片の両端に制限酵素PfoI部位とSbfI部位を付加するために2種類のプライマー(配列番号52と53)を合成した。また、DRHG株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりaroF遺伝子のプロモーターの5’フランキング領域と3’フランキング領域をそれぞれ増幅した。これとは別に、DRHG株の染色体DNAを鋳型にして配列番号35と36のプライマーを用いるPCR法によりTuプロモーターを増幅した。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacB_aroF-NtuをNSU株に導入して、カナマイシン耐性で選択することにより、NSU_Km株を取得した。得られたNSU_Km株が目的の1回交差型相同組換え体であることを確認するために、aroF遺伝子のプロモーターの5’フランキング領域の上流の一部と3’フランキング領域の下流の一部に基づく2種類のプライマー(配列番号54と55)を合成した。NSU_Km株の染色体DNAを鋳型として、配列番号54と55のプライマー、配列番号35と36のプライマーならびにsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりNSU_Km株を解析したところ、予想通りの結果が得られたことから、NSU_Km株はプラスミドpHKPsacB_aroF-NtuがaroF遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
ATCC13032株が保有する3種類のシキミ酸脱水素酵素のうち、生育に影響を与える可能性があるaroE1遺伝子とaroE3遺伝子にインフレーム欠失変異を導入した菌株(インフレーム欠失変異を導入した菌株(以下、必要に応じて、インフレーム破壊株とも呼ぶ)を以下のようにして造成した。なお、インフレーム欠失変異、すなわちインフレーム破壊とは、翻訳領域内に3塩基の倍数の長さの欠失を導入した変異のことをいう。
aroE1遺伝子の5’フランキング領域を(塩基番号1,183,128~1,184,160)を増幅させるために2種類のDNAプライマー(配列番号56と57)を合成した。なお、配列番号56のプライマーには制限酵素SphI部位を導入した。DRHG株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりaroE1遺伝子の5’フランキング領域を増幅した。次に、aroE1遺伝子の3’フランキング領域(塩基番号1,181,390~1,182,365)を増幅させるために2種類のDNAプライマー(配列番号58と59)を合成した。なお、配列番号59のプライマーには制限酵素SbfI部位を導入した。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacBΔaroE1をNSH株に導入し、カナマイシン耐性で選択することにより、NSH_Km株を取得した。得られたNSH_Km株が目的の1回交差型相同組換え体であることを確認するために、aroE1遺伝子5’フランキング領域の上流の一部と3’フランキング領域の下流の一部に基づく2種類のプライマー(配列番号60と61)を合成した。配列番号56と59のプライマーならびにsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりNSH_Km株を解析したところ、予想通りの結果が得られたことから、NSH_Km株はプラスミドpHKPsacBΔaroE1がaroE1遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
aroE3遺伝子の5’フランキング領域を(塩基番号1,726,887~1,727,845)を増幅させるために2種類のDNAプライマー(配列番号62と63)を合成した。なお、配列番号62のプライマーには制限酵素SphI部位を導入した。DRHG株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりaroE3遺伝子の5’フランキング領域を増幅した。次に、aroE3遺伝子の3’フランキング領域(塩基番号1,725,101~1,726,094)を増幅させるために2種類のDNAプライマー(配列番号64と65)を合成した。なお、配列番号65のプライマーには制限酵素SbfI部位を導入した。このようにして得られたaroE3遺伝子の5’フランキング領域の増幅DNAと3’フランキング領域の増幅DNAの混合物を鋳型として、配列番号62と65のプライマーを用いるオーバーラップPCR法により増幅反応を行った後、制限酵素SphIとSbfIによる消化反応を行った。生じたDNA断片をプラスミドpHKPsacB1(参考例2参照)のマルチクローニングサイト内の制限酵素SbfI 部位とSphI部位の間に導入し、プラスミドpHKPsacBΔaroE3を得た。なお、プラスミドpHKPsacBΔaroE3は、aroE3遺伝子領域のうち、相補鎖塩基番号1,726,095~1,726,886の領域が欠失している構造を有する。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacBΔaroE3をNSH株に導入し、カナマイシン耐性で選択することにより、NSH_Km2株を取得した。得られたNSH_Km2株が目的の1回交差型相同組換え体であることを確認するために、aroE3遺伝子5’フランキング領域の上流の一部と3’フランキング領域の下流の一部に基づく2種類のプライマー(配列番号66と67)を合成した。配列番号62と65のプライマー、配列番号66と67のプライマー、ならびにsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりNSH_Km株を解析したところ、予想通りの結果が得られたことから、NSH_Km2株はプラスミドpHKPsacBΔaroE3がaroE3遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
NSHΔaroE1菌をもとに、上述の「aroE3遺伝子のインフレーム破壊株の造成」と同じ手法で、aroE3遺伝子もインフレームで破壊した2重破壊株(NSHΔaroE1ΔaroE3株と命名)を造成した。
上述のようにして造成したNSH株、NSHΔaroE1株、NSHΔaroE3株およびにNSHΔaroE1ΔaroE3株の4株について、以下のようにして芳香族アミノ酸の要求性試験を行った。
ATCC13032株のバニリン酸脱メチル酵素をコードする遺伝子はcg2616(以下vanA遺伝子と略す。塩基番号2,496,775~2,497,905 bp:大きさ1,131 bp)とcg2617(以下vanB遺伝子と略す。塩基番号2,497,909~2,498,886 bp:大きさ978 bp)である。また2,496,775~2,498,886 bpをvanAB遺伝子領域と略す。またvanA遺伝子とvanB遺伝子を制御しているリプレッサーはcg2615 (以下vanR遺伝子と略す。相補鎖塩基番号2,496,013~2,496,591 bp:大きさ579 bp)が既知である。vanR遺伝子により、aroE3遺伝子をvanA遺伝子とvanB遺伝子領域に導入し、aroE3遺伝子をvanR遺伝子制御下に置換した。親株は、上記参考例7に示すNSHΔaroE3株を用いた。
ATCC13032株のvanAB遺伝子領域をaroE3遺伝子に置換するために以下の手順で置換用ベクターを構築した。まず、ATCC13032株のゲノム配列をもとに、vanAB遺伝子領域の5’フランキング領域(塩基番号2,495,799~2,496,774 bp:大きさ976kp)を増幅させるために2種類のDNAプライマー(配列番号68と配列番号69)を合成し、vanAB遺伝子領域の3’フランキング領域(塩基番号2,498,887~2,499,870 bp:大きさ984kp)を増幅するために2種類のDNAプライマー(配列番号70と配列番号71)を合成した。これらのプライマーを用いて菌株NSHの染色体DNAを鋳型にしてPCR法によりvanAB遺伝子の5’フランキング領域、vanAB遺伝子の3’フランキング領域を増幅した。vanAB遺伝子の5’フランキング領域を増幅するとともに配列番号68のDNAプライマーに制限酵素サイトSphIを導入し、vanAB遺伝子の3’フランキング領域を増幅するとともに配列番号71のDNAプライマーに制限酵素サイトSbfIを導入した。またaroE3遺伝子を増幅するために2種類のDNAプライマー(配列番号72と配列番号73)を合成した。aroE3遺伝子を増幅するとともに、配列番号72のDNAプライマーにvanAB遺伝子領域の5’フランキング領域の3´末端の15 bpを付加し、配列番号73のDNAプライマーにvanAB遺伝子領域の3’フランキング領域の5´末端の15 bpを付加し増幅した。増幅した5’フランキング領域、aroE3遺伝子を混合したDNA断片を鋳型として、DNAプライマー(配列番号68と配列番号73)を用い、また増幅した3’フランキング領域、aroE3遺伝子を混合したDNA断片を鋳型として、DNAプライマー(配列番号72と配列番号71)を用い、オーバーラップPCR法で増幅した。それぞれ増幅したDNA断片を制限酵素SphIとaroE3遺伝子内部制限酵素サイトAscIおよびSbfIとaroE3遺伝子内部制限酵素サイトAscIで消化し、プラスミドpHKPsacB1(参考例2参照)のマルチクローニングサイト内の制限酵素部位SbfI - SphI間に導入し、プラスミドpHKPsacB_vanR_Pvan-aroE3を得た。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacB_vanR_Pvan-aroE3を菌株NSHΔaroE3に導入して、カナマイシン耐性株NSHΔaroE3を取得した。得られたカナマイシン耐性株の染色体DNA上(塩基番号2,495,799~2,499,870 bp:大きさ4,071 bp)領域のうち塩基番号2,496,775~2,498,886 bpがaroE3遺伝子に置換していることを確認するために以下の実験を行った。vanAB遺伝子の5’フランキング領域の112 bp外側と3’フランキング領域の106 bp外側にDNAプライマー(配列番号74と配列番号75)を作製し、vanAB遺伝子の5’フランキング領域と3’フランキング領域を増幅する際に使用したDNAプライマー(配列番号68と配列番号71)を用いて菌株NSHΔaroE3のカナマイシン耐性株のコロニーを直接鋳型としたPCRを行った。
菌株NSHΔaroE3のカナマイシン耐性株の染色体DNAを鋳型にし、配列番号74と配列番号71のDNAプライマーで増幅したPCRによって取得される増幅DNA断片が4,194 bp、配列番号68と配列番号75のDNAプライマーで増幅したPCRによって取得される増幅DNA断片は2,905 bpであった。もしくは配列番号74と配列番号71のDNAプライマーで増幅したPCRによって取得される増幅DNA断片が2,913bp、配列番号68と配列番号75のDNAプライマーで増幅したPCRによって取得される増幅DNA断片は4,186 bpであった。またsacB遺伝子領域が、PCR法による解析などによって確認されたことから、菌株NSHΔaroE3は1回交差型相同組換え体であることがわかった。
17.5 mg/Lのシキミ酸、50 mg/Lのトリプトファン、50 mg/Lのファニルアラニン、50 mg/Lのチロシン、10 mg/Lのパラヒドロキシ安息香酸および10 mg/Lのパラアミノ安息香酸(以下、6種類の添加物という)を加えたCGXII培地および6種類の添加物を加えないCGXII培地を用いて、菌株NSHΔaroE3_vanE3とNSHΔaroE3の生育性試験を行った。その結果、菌株NSHΔaroE3_vanE3は前者の培地では正常に増殖したが、後者の培地では培養中期までの生育速度が遅かった。一方、菌株NSHΔaroE3は前者の培地では増殖したが、後者の培地では増殖しなかった。また、6種類の添加物を加えないCGXII培地にフェルラ酸またはバニリン酸またはバニリンを50 μM以上添加して、菌株NSHΔaroE3_vanE3を培養したときには、正常に増殖することがわかった。
参考例6、7および8に示すNSH株、NSHΔaroE3株およびNSHΔaroE3_vanE3株を親株として、qsuB遺伝子にインフレーム欠失変異を導入することにより、DHSからプロトカテク酸への転換を遮断した菌株を以下のようにして造成した。
qsuB遺伝子の5’フランキング領域(塩基番号443,184~444,207)を増幅するとともに、増幅断片の両端に制限酵素SbfI部位とXbaI部位を付加するために2種類のDNAプライマー(配列番号76と77)を合成した。また、qsuB遺伝子の3’フランキング領域(塩基番号446,032~447,053)を増幅するともに、増幅断片の両端に制限酵素XbaI部位とSalI部位を付加するために2種類のDNAプライマー(配列番号78と79)を合成した。NSH株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりqsuB遺伝子の5’フランキング領域と3’フランキング領域を増幅した。5’フランキング領域増幅物を制限酵素SbfIおよびXbaIで消化し、また3’フランキング領域増幅物を制限酵素XbaIおよびSalIで消化した後、これらのDNA断片をプラスミドpHKPsacB1(参考例2参照)のマルチクローニングサイト内にあるSbfI 部位とSalI部位の間に導入し、qsuB遺伝子破壊用プラスミドであるpHKPsacBΔqsuBを得た。
上述のプラスミドpHKPsacBΔqsuBをNSH株、NSHΔaroE3株およびNSHΔaroE3_vanE3株に導入し、カナマイシン耐性で選択することにより、それぞれの菌株からNSH_qsuB_Km株、NSHΔaroE3_qsuB_Km株およびNSHΔaroE3_vanE3_qsuB_Km株を取得した。取得した菌株をPCR法により解析するために、qsuB遺伝子の5’フランキング領域の上流の配列に一致するDNAプライマー(配列番号80)と3’フランキング領域の下流の配列に一致するDNAプライマー(配列番号81)を合成し、配列番号79と80のプライマー、配列番号76と81のプライマーを用いるPCR法によって解析した。いずれのプライマーセットを用いたPCR法による解析でも予想通りの結果が得られたことから、これら菌株はプラスミドpHKPsacBΔqsuB がqsuB遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
NSHΔaroE3ΔqsuB株およびNSHΔaroE3_vanE3ΔqsuB株をもとに、残存するシキミ酸脱水酵素活性を低減するために、シキミ酸脱水酵素をコードするqsuD遺伝子にインフレーム欠失変異を導入した菌株を以下のようにして造成した。
qsuD遺伝子の5’フランキング領域を(塩基番号443,644~446,553、ただし444,199~446,031の領域はqsuBのインフレーム破壊領域である)を増幅させるために2種類のDNAプライマー(配列番号82と83)を合成した。なお、配列番号82のプライマーには制限酵素SbfI部位を導入した。NSHΔqsuB株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりqsuD遺伝子の5’フランキング領域を増幅した。次に、qsuD遺伝子の3’フランキング領域(塩基番号447,385~448,441)を増幅させるために2種類のDNAプライマー(配列番号84と85)を合成した。なお、配列番号85のプライマーには制限酵素XhoI部位を導入した。このようにして得られたaroE遺伝子の5’フランキング領域の増幅DNAと3’フランキング領域の増幅DNAの混合物を鋳型としてDNAプライマー(配列番号82と85)を用いてオーバーラップPCR法による増幅反応を行った後、制限酵素SbfIとXhoIで消化反応を行った。生じたDNA断片をプラスミドpHKPsacB1(参考例2参照)のマルチクローニングサイト内の制限酵素部位SbfI - SalI間に導入し、プラスミドpHKPsacBΔqsuDを得た。なお、プラスミドpHKPsacBΔqsuDは、qsuD遺伝子領域のうち、塩基番号446,554~447,384の領域が欠失している構造を有する。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacBΔqsuDをNSHΔaroE3ΔqsuB株、およびNSHΔaroE3_vanE3ΔqsuB株に導入し、カナマイシン耐性で選択することにより、NSHΔaroE3ΔqsuB_Km株、およびNSHΔaroE3_vanE3ΔqsuB_km株を取得した。得られたNSHΔaroE3ΔqsuB_Km株、およびNSHΔaroE3_vanE3ΔqsuB_km株が目的の1回交差型相同組換え体であることを確認するために、qsuD遺伝子5’フランキング領域の上流の一部と3’フランキング領域の下流の一部に基づく2種類のプライマー(配列番号86と87)を合成した。配列番号86と85のプライマー、配列番号82と87ならびにsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりNSHΔaroE3ΔqsuB_Km株、およびNSHΔaroE3_vanE3ΔqsuB_km株を解析したところ、予想通りの結果が得られたことから、NSHΔaroE3ΔqsuB_Km株、およびNSHΔaroE3_vanE3ΔqsuB_km株はプラスミドpHKPsacBΔqsuDがqsuD遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
上述のようにして造成したNSH株、NSHΔaroE3株、NSHΔaroE3ΔqsuB株、NSHΔaroE3ΔqsuBΔaroE1株、NSHΔaroE3_vanE3ΔqsuB株、NSHΔaroE3ΔqsuBΔqsuD株の6株について、以下のようにして芳香族アミノ酸の要求性試験を行った。
参考例10で造成したNSHΔaroE3_vanE3ΔqsuBΔqsuD株のbenABCD遺伝子領域をvanR遺伝子と置換することにより、benAプロモーターの制御下でvanR遺伝子を発現する菌株を造成した。
benABCD遺伝子領域の5’フランキング領域(塩基番号2,515,963~2,516,969)を増幅させるために2種類のDNAプライマー(配列番号88と89)を合成し、またbenABCD遺伝子領域の3’フランキング領域(塩基番号2,521,372~2,522,498)を増幅するために2種類のDNAプライマー(配列番号90と91)を合成した。なお、配列番号88と91のプライマーはそれぞれ制限酵素SalI部位とSbfI部位を持つ。HT23株(参考例3)の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりbenABCD遺伝子の5’フランキング領域と3’フランキング領域を増幅した。また、vanR遺伝子を増幅するために2種類のDNAプライマー(配列番号92と93)を合成した。HT23株の染色体DNAを鋳型にして配列番号92と93のプライマーを用いるPCR法によりvanR遺伝子領域を増幅した。このようにして得られたbenABCD遺伝子の5’フランキング領域の増幅DNAとqsuB遺伝子領域の増幅DNAの混合物を鋳型として、配列番号88と93のプライマーを用いるオーバーラップPCR法により増幅反応を行った後、制限酵素SalIとNarIによる消化により、配列番号88のプライマー内のSalI部位とvanR遺伝子内部のNarI部位で切断されたDNA断片を得た。また、vanR遺伝子領域の増幅DNAとbenABCD遺伝子の3’フランキング領域の増幅DNAの混合物を鋳型として、配列番号91と92のプライマーを用いたオーバーラップPCR法により増幅反応を行った後、制限酵素NarIとSbfIによる消化により、vanR遺伝子内部のNarI部位と配列番号91のプライマー内のSbfI部位で切断されたDNA断片を得た。これら2種類のDNA断片をプラスミドpHKPsacB1(参考例2参照)のマルチクローニングサイト内にあるSalI部位とSbfI部位の間に導入し、プラスミドpHKPsacB_Pben-vanRを得た。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacB_Pben-vanRをNSHΔaroE3_vanE3ΔqsuBΔqsuD株に導入してカナマイシン耐性となったNSHΔaroE3_vanE3ΔqsuBΔqsuD株_Km株を取得した。続いて、NSHΔaroE3_vanE3ΔqsuBΔqsuD株_Km株をPCR法により解析するために、vanR遺伝子の5’フランキング領域の上流の配列に一致するプライマー(配列番号94)と3’フランキング領域の下流の配列に一致するプライマー(配列番号95)を合成した後、配列番号94と91のプライマーならびに配列番号88と95のプライマーを用いるPCR法によって解析した。いずれのプライマーセットを用いたPCR法による解析でも、予想通りの結果が得られたことから、NSHΔaroE3_vanE3ΔqsuBΔqsuD株_Km株はプラスミドpHKPsacB_Pben-vanR がbenABCD遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
以下のようにして、参考例9で造成したNSHΔaroE3_vanE3ΔqsuB株のbenABCD遺伝子オペロン領域をqsuB遺伝子と置換することにより、benAプロモーターの制御下でqsuB遺伝子を発現する菌株を造成した。
benABCD遺伝子領域の5’フランキング領域を増幅させるために2種類のDNAプライマー(配列番号88と89)を利用し、またbenABCD遺伝子領域の3’フランキング領域を増幅するために2種類のDNAプライマー(配列番号90と91)を利用した。HT23株(参考例3)の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりbenABCD遺伝子の5’フランキング領域と3’フランキング領域を増幅した。また、qsuB遺伝子を増幅するために2種類のDNAプライマー(配列番号96と97)を合成し、HT23株の染色体DNAを鋳型にしてこれらプライマーを用いるPCR法によりqsuB遺伝子領域を増幅した。このようにして得られたbenABCD遺伝子の5’フランキング領域の増幅DNAとqsuB遺伝子領域の増幅DNAの混合物を鋳型として、配列番号88と97のプライマーを用いるオーバーラップPCR法により増幅反応を行った後、制限酵素SalIとNarIによる消化により、配列番号88のプライマー内のSalI部位とqsuB遺伝子内部のNarI部位で切断されたDNA断片を得た。また、qsuB遺伝子領域の増幅DNA とbenABCD遺伝子の3’フランキング領域の増幅DNAの混合物を鋳型として、配列番号91と96のプライマーを用いたオーバーラップPCR法により増幅反応を行った後、制限酵素NarIとSbfIによる消化により、qsuB遺伝子内部のNarI部位と配列番号91のプライマー内のSbfI部位で切断されたDNA断片を得た。これら2種類のDNA断片をプラスミドpHKPsacB1(参考例2参照)のマルチクローニングサイト内にあるSalI部位とSbfI部位の間に導入し、プラスミドpHKPsacB_Pben-qsuBを得た。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacB_Pben-qsuBをNSHΔaroE3_vanE3ΔqsuB株に導入してカナマイシン耐性となったNSHΔaroE3_vanE3ΔqsuB_Km株を取得した。続いて、NSHΔaroE3_vanE3ΔqsuB_Km株をPCR法により解析するために、qsuB遺伝子の5’フランキング領域の上流の配列に一致するプライマー(配列番号98)と3’フランキング領域の下流の配列に一致するプライマー(配列番号99)を合成した後、配列番号88と99のプライマーならびに配列番号91と98のプライマーを用いるPCR法によって解析した。いずれのプライマーセットを用いたPCR法による解析でも、予想通りの結果が得られたことから、NSHΔaroE3_vanE3ΔqsuB_Km株はプラスミドpHKPsacB_Pben-qsuB がbenABCD遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
参考例12で造成したNSHΔaroE3_vanE3ΔqsuB_Pben-qsuB株をもとに、qsuB遺伝子とオペロン構造を保持する形でvanR遺伝子を導入した。qsuB遺伝子とオペロン構造を保持することにより、benAプロモーターの制御下でqsuB遺伝子とvanR遺伝子を発現する菌株を以下のようにして造成した。
vanR遺伝子挿入領域の5’フランキング領域(塩基番号2,520,333~2,521,374)を増幅させるために2種類のDNAプライマー(配列番号100と101)を合成し、またvanR遺伝子挿入領域の3’フランキング領域(塩基番号2,521,372~2,522,498)を増幅するために2種類のDNAプライマー(配列番号102と103)を合成した。なお、配列番号100と103のプライマーはそれぞれ制限酵素SalI部位とSbfI部位を持つ。NSHΔaroE3_vanE3ΔqsuB_Pben-qsuB株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりvanR遺伝子挿入領域の5’フランキング領域と3’フランキング領域を増幅した。また、vanR遺伝子を増幅するために2種類のDNAプライマー(配列番号104と105)を合成し、HT23株(参考例3)の染色体DNAを鋳型にしてこれらプライマーを用いるPCR法によりvanR遺伝子領域を増幅した。このようにして得られたvanR遺伝子挿入領域の5’フランキング領域の増幅DNAとvanR遺伝子領域の増幅DNAの混合物を鋳型として、配列番号100と105のプライマーを用いるオーバーラップPCR法により増幅反応を行った後、制限酵素SalIとBamHIによる消化により、配列番号100のプライマー内のSalI部位とvanR遺伝子内部のBamHI部位で切断されたDNA断片を得た。また、vanR遺伝子領域の増幅DNAとvanR遺伝子挿入領域の3’フランキング領域の増幅DNAの混合物を鋳型として、配列番号104と103のプライマーを用いたオーバーラップPCR法により増幅反応を行った後、制限酵素BamHIとSbfIによる消化により、vanR遺伝子内部のBamHI部位と配列番号103のプライマー内のSbfI部位で切断されたDNA断片を得た。これら2種類のDNA断片をプラスミドpHKPsacB1(参考例2参照)のマルチクローニングサイト内にあるSalI部位とSbfI部位の間に導入し、プラスミドpHKPsacB_Pben-qsuB-vanRを得た。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacB_Pben-qsuB-vanRをNSHΔaroE3_vanE3ΔqsuB_Pben-qsuB株に導入してカナマイシン耐性となったNSHΔaroE3_vanE3ΔqsuB_Pben-qsuB株_Km株を取得した。続いて、NSHΔaroE3_vanE3ΔqsuB_Pben-qsuB株_Km株をPCR法により解析するために、vanR遺伝子導入領域の5’フランキング領域の上流の配列に一致するプライマー(配列番号106)と3’フランキング領域の下流の配列に一致するプライマー(配列番号107)を合成した後、配列番号100と103のプライマーを用いるPCR法によって解析した。いずれのプライマーセットを用いたPCR法による解析でも、予想通りの結果が得られたことから、NSHΔaroE3_vanE3ΔqsuB_Pben-qsuB株_Km株はプラスミドpHKPsacB_Pben-qsuB-vanRがvanR遺伝子挿入領域に導入された1回交差型相同組換え体であることを確認した。
参考例13で造成したNSHΔaroE3_vanE3ΔqsuB_Pben-qsuB-vanR株を親株として、エリトロース-4-リン酸の合成に関わるtkt遺伝子をTuプロモーター(参考例6)に連結した転写ユニット(以下、Ptu-tkt転写ユニットと略す)をDNA修飾・制限酵素遺伝子の破壊部位に挿入したtkt株を以下のようにして造成した。
DNA修飾・制限酵素遺伝子の破壊部位の5’フランキング領域(塩基番号1,885,120~1,883,861)を増幅させるために2種類のDNAプライマー(配列番号108と109)を合成し、DNA修飾・制限酵素遺伝子の破壊部位の3’フランキング領域(塩基番号1,879,744~1,878,664)を増幅するために2種類のDNAプライマー(配列番号110と111)を合成した。またTuプロモーターを増幅させるためにDNAプライマー(配列番号112と36)を、tkt遺伝子を増幅させるために2種類のDNAプライマー(配列番号113と114)合成した。なお、配列番号108と111のプライマーはそれぞれ制限酵素SbfI部位とSalI部位を持つ。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacB_Ptu-tktをNSHΔaroE3_vanE3ΔqsuB_Pben-qsuB-vanR株に導入してカナマイシン耐性となったNSHΔaroE3_vanE3ΔqsuB_Pben-qsuB-vanR株_Km株を取得した。続いて、NSHΔaroE3_vanE3ΔqsuB_Pben-qsuB-vanR株_Km株をPCR法により解析するために、5’フランキング領域の上流の配列に一致するプライマー(配列番号115)と3’フランキング領域の下流の配列に一致するプライマー(配列番号116)を合成した後、配列番号115と116のプライマーならびに配列番号108と111のプライマーを用いるPCR法によって解析した。いずれのプライマーセットを用いたPCR法による解析でも、予想通りの結果が得られたことから、NSHΔaroE3_vanE3ΔqsuB_Pben-qsuB-vanR株_Km株はプラスミドpHKPsacB_Ptu-tktが導入された1回交差型相同組換え体であることを確認した。
cg1309遺伝子からcg1311遺伝子までの領域(以下、rhcHMD遺伝子領域と略す)をaroE3遺伝子で置換した菌株を以下のように造成した。
3-ヒドロキシ安息香酸6-ヒドロキシラーゼをコードする遺伝子であるcg3354遺伝子(以下、genH遺伝子という)の5’フランキング領域を(塩基番号3,201,424~3,202,403)を増幅するとともに、増幅断片の両端に制限酵素KpnI部位とSbfI部位を付加するために、2種類のDNAプライマー(配列番号117と118)を合成した。DRHG株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりgenH遺伝子の5’フランキング領域を増幅した。次に、genH遺伝子の3’フランキング領域(塩基番号3,203,894~3,204,901)を増幅するとともに、増幅断片の両端に制限酵素SbfI部位とSalI部位を付加するために2種類のDNAプライマー(配列番号119と120)を合成した。DRHG株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりgenH遺伝子の3’フランキング領域を増幅した。5’フランキング領域の増幅DNA断片を制限酵素KpnIとSbfIで消化し、また3’フランキング領域の増幅DNA断片を制限酵素SbfIとSalIで消化した後、これらのDNA断片をプラスミドpHKPsacB1(参考例2参照)のマルチクローニングサイト内にあるKpnI部位とSalI部位の間に導入し、プラスミドpHKPsacBΔgenHを得た。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacBΔgenHをNSHΔaroE3株に導入し、カナマイシン耐性で選択することにより、NSHΔaroE3_Km株を取得した。配列番号117と120のプライマーならびにsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりNSHΔaroE3_Km株を解析したところ、予想通りの結果が得られたことから、これら菌株はプラスミドpHKPsacBΔgenHがgenH遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
rhcHMD遺伝子領域の5’フランキング領域(塩基番号1,213,341~1,214,736)を増幅させるために2種類のDNAプライマー(配列番号121と配列番号122)を合成した。なお、配列番号121のプライマーには制限酵素SbfI部位を導入した。NSH株の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりrhcHMD遺伝子領域の5’フランキング領域を増幅した。次に、rhcHMD遺伝子の3’フランキング領域(塩基番号1,218,296~1,219,545)を増幅させるために2種類のDNAプライマー(配列番号123と配列番号124)を合成した。なお、配列番号124のプライマーには制限酵素SalI部位を導入した。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacB_rhcR_Prhc-rhcE3を上述の菌株NSHΔaroE3ΔgenH株に導入し、カナマイシン耐性で選択することによりNSHΔaroE3ΔgenH_Km株を取得した。
配列番号127と配列番号128のDNAプライマーと、配列番号121と配列番号124のプライマーとsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりNSHΔaroE3ΔgenH_Km株を解析したところ、予想通りの結果が得られたことから、NSHΔaroE3ΔgenH_Km株はプラスミドpHKPsacB_rhcR_Prhc-rhcE3がrhcHMD遺伝子領域に導入された1回交差型相同組換え体であることを確認した。
造成したNSH株、NSHΔaroE3株、NSHΔaroE3_vanE3株、NSHΔaroE3ΔgenH_rhcE3株について、以下のようにして芳香族アミノ酸の要求性試験を行った。
rhcR遺伝子がコードするRhcRリプレッサーによりaroE3遺伝子の転写を制御する系を構築ために、nagIプロモーターの制御下でrhcR遺伝子を発現する菌株を以下のようにして造成した。
以下のようにして、上記で造成したNSHΔaroE3ΔgenH株のnagIKL遺伝子オペロン領域をrhcR遺伝子と置換することにより、nagIプロモーターの制御下でrhcR遺伝子を発現する菌株を造成した。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacB_Pnag-rhcRを菌株NSHΔaroE3ΔgenH_rhcE3に導入し、カナマイシン耐性で選択することによりNSHΔaroE3ΔgenH_rhcE3_Km株を取得した。配列番号135と配列番号136のDNAプライマーと、配列番号129と配列番号132のプライマーとsacB遺伝子確認用プライマー(配列番号3と4)を用いるPCR法によりNSHΔaroE3ΔgenH_rhcE3_Km株を解析したところ、予想通りの結果が得られたことから、NSHΔaroE3ΔgenH_rhcE3_Km株はプラスミドpHKPsacB_Pnag-rhcRがnagIKL遺伝子オペロン領域に導入された1回交差型相同組換え体であることを確認した。
qsuB遺伝子をPnag-rhcR株のnagIプロモーター制御下に導入した菌株を以下のようにして造成した。
qsuB遺伝子導入領域の5’フランキング領域(塩基番号3,199,996~3,200,904)を増幅させるために2種類のDNAプライマー(配列番号137と138)を合成し、またqsuB遺伝子導入領域の3’フランキング領域(塩基番号3,198,925~3,199,995 この領域はrhcR遺伝子に置換されている)を増幅するために2種類のDNAプライマー(配列番号139と140)を合成した。なお、配列番号137と140のプライマーはそれぞれ制限酵素SbfI部位とSphI部位を持つ。Pnag-rhcR株(参考例16)の染色体DNAを鋳型にして、これらのプライマーを用いるPCR法によりqsuB遺伝子導入領域の5’フランキング領域と3’フランキング領域を増幅した。また、qsuB遺伝子を増幅するために2種類のDNAプライマー(配列番号141と142)を合成し、HT23株(参考例3)の染色体DNAを鋳型にしてこれらプライマーを用いるPCR法によりqsuB遺伝子領域を増幅した。このようにして得られたqsuB遺伝子導入領域の5’フランキング領域の増幅DNAとqsuB遺伝子領域の増幅DNAの混合物を鋳型として、配列番号137と142のプライマーを用いるオーバーラップPCR法により増幅反応を行った後、制限酵素 SbfIとXhoIによる消化により、配列番号137のプライマー内のSbfI部位とqsuB遺伝子内部のXhoI部位で切断されたDNA断片を得た。また、qsuB遺伝子領域の増幅DNA とqsuB遺伝子導入領域の3’フランキング領域の増幅DNAの混合物を鋳型として、配列番号140と141のプライマーを用いたオーバーラップPCR法により増幅反応を行った後、制限酵素XhoIとSphIによる消化により、qsuB遺伝子内部のXhoI部位と配列番号140のプライマー内のSphI部位で切断されたDNA断片を得た。これら2種類のDNA断片をプラスミドpHKPsacB1(参考例2参照)のマルチクローニングサイト内にあるSbfI部位とSphI部位の間に導入し、プラスミドpHKPsacB_Pnag-qsuB-rhcRを得た。
エレクトロポレーションによる形質転換法を用いて、上述のプラスミドpHKPsacB_Pnag-qsuB-rhcRをPnag-rhcR株に導入してカナマイシン耐性となったPnag-rhcR株_Km株を取得した。続いて、Pnag-rhcR株_Km株をPCR法により解析するために、qsuB遺伝子導入領域の5’フランキング領域の上流の配列に一致するプライマー(配列番号143)と3’フランキング領域の下流の配列に一致するプライマー(配列番号144)を合成した後、配列番号137と140のプライマーを用いるPCR法によって解析した。いずれのプライマーセットを用いたPCR法による解析でも、予想通りの結果が得られたことから、Pnag-rhcR株_Km株はプラスミドpHKPsacB_Pnag-qsuB-rhcRがnagIプロモーターの下流に導入された1回交差型相同組換え体であることを確認した。
Claims (24)
- 炭素源から生育に必須な代謝物Mを生成する生合成経路において中間代謝物である化合物Pを代謝物Mに転換する酵素Xの発現量を調節することにより化合物Pを蓄積させるために、以下の(a)から(d)に記載の性質をすべて有する原核生物。
(a)該原核生物が利用できる炭素源から化合物Pに至る生合成に関わる酵素群のうち、いずれか1つ以上の酵素の活性が該原核生物の野生株と比べて増強している。
(b)酵素Xのタンパク質をコードする野生型遺伝子xの翻訳領域、該遺伝子の翻訳調節領域または遺伝子xの転写を促すプロモーター領域の中に1個以上の塩基の置換、欠失または付加による変異を有することにより、酵素Xの活性が欠損または低下している。
(c)前記遺伝子xの転写を促す本来のプロモーターとは異なっており、かつリプレッサーRのタンパク質により転写が抑制されるプロモーターAにより、活性型酵素XをコードするDNAの転写が制御される。
(d)リプレッサーRのタンパク質をコードする遺伝子zを1コピー以上有し、該遺伝子の転写が該遺伝子の本来のプロモーターおよび/または該プロモーターとは異なる誘導プロモーターBにより制御される。 - 性質(b)に記載の遺伝子xの転写を促すプロモーター領域の中の置換変異が、該プロモーターの全域または一部の領域を、性質(c)に記載のプロモーターAのDNAと置換する変異であることを特徴とする、請求項1に記載の原核生物。
- 前記原核生物が有する化合物Pを代謝する酵素のうち、酵素X以外のいずれか1つ以上の代謝酵素の活性が欠損または低下していることを特徴とする、請求項1または2に記載の原核生物。
- 前記プロモーターBが、フェルラ酸、バニリン酸、バニリン、安息香酸、3-ヒドロキシ安息香酸、レゾルシノール、4-ヒドロキシ安息香酸、2,4-ジヒドロキシ安息香酸、フラクトースおよびスクロースからなる群から選ばれる化合物の添加により誘導されるプロモーターであることを特徴とする、請求項1から3のいずれか一項に記載の原核生物。
- 前記リプレッサーRのタンパク質をコードする遺伝子zが、コリネバクテリウム・グルタミカムATCC13032株のvanR(cg2615)遺伝子、pcaR(cg2624)遺伝子、またはrhcR(cg1308)遺伝子であることを特徴とする、請求項1から4のいずれか一項に記載の原核生物。
- 前記プロモーターAが、コリネバクテリウム・グルタミカムATCC13032株のvanA(cg2616)遺伝子のプロモーター、pobA(cg1226)遺伝子のプロモーター、pcaH(cg2631)遺伝子のプロモーター、またはrhcH(cg1309)遺伝子のプロモーターであることを特徴とする、請求項1から5のいずれか一項に記載の原核生物。
- 請求項1から6のいずれか一項に記載の原核生物をプロモーターAの転写を抑制した状態で炭素源の存在下で培養することを特徴とする、化合物Pの製造方法。
- 請求項1から6のいずれか一項に記載の原核生物をプロモーターAの転写を抑制した状態で炭素源の存在下で培養した後、プロモーターBの転写を誘導する処理を加えることによりリプレッサーRのタンパク質の発現量を増加させることにより、代謝物Mの生成量を減少させることを特徴とする、化合物Pの製造方法。
- 請求項1から6のいずれか一項に記載の原核生物をプロモーターAの転写抑制を解除した状態で炭素源の存在下で培養した後、プロモーターBの転写を誘導する処理を加えることによりリプレッサーRのタンパク質の発現量を増加させることにより、代謝物Mの生成量を減少させることを特徴とする、化合物Pの製造方法。
- 前記化合物Pおよび前記代謝物Mの組み合わせが、以下(f)から(i)のいずれかであることを特徴とする、請求項7から9のいずれか一項に記載の化合物Pの製造方法。
(f)化合物Pが3-デヒドロシキミ酸であり、代謝物Mがシキミ酸である。
(g)化合物Pがグルタミン酸であり、代謝物MがN-アセチルグルタミン酸またはγーグルタミルリン酸である。
(h)化合物Pがアスパラギン酸であり、代謝物Mがβ-アルパルチルリン酸である。
(i)化合物Pがセリンであり、代謝物Mがグリシンである。 - 蓄積する化合物Pを酵素Yにより化合物Qに転換するために、(a)から(d)に記載の性質に加えて下記の性質(e)を有することを特徴する、請求項1から6のいずれか一項に記載の原核生物。
(e)酵素Yのタンパク質をコードする遺伝子yの翻訳領域、該遺伝子の翻訳調節領域または遺伝子yの転写を促すプロモーター領域の中に1個以上の塩基の置換、欠失または付加による変異を有することにより、化合物Pから化合物Qへの転換能が増強している、もしくは遺伝子yの発現が野生型とは異なる制御を受けている。 - 前記原核生物が有する化合物Qを代謝する酵素のうち、いずれか1つ以上の代謝酵素の活性が欠損または低下していることを特徴とする、請求項11に記載の原核生物。
- 性質(e)に記載の遺伝子yの転写を促すプロモーター領域の中の置換変異が、該プロモーターの全域または一部の領域を、該遺伝子の転写を促す本来のプロモーターおよびプロモーターAとは異なるプロモーターCのDNAと置換する変異であることを特徴とする、請求項11または12に記載の原核生物。
- 前記プロモーターCがプロモーターBと同一である、または前記プロモーターCがプロモーターBの転写を制御するタンパク質による転写制御を受けることを特徴とする、請求項13に記載の原核生物。
- 前記プロモーターCが誘導プロモーターであることを特徴とする、請求項13または14に記載の原核生物。
- 前記プロモーターCが、フェルラ酸、バニリン酸、バニリン、安息香酸、3-ヒドロキシ安息香酸、レゾルシノール、4-ヒドロキシ安息香酸、2,4-ジヒドロキシ安息香酸、フラクトースおよびスクロースからなる群から選ばれる化合物の添加により誘導されるプロモーターであることを特徴とする、請求項15に記載の原核生物。
- 請求項11から16のいずれか一項に記載の原核生物をプロモーターAの転写を抑制した状態で炭素源の存在下で培養することを特徴とする、化合物Qの製造方法。
- 請求項11から16のいずれか一項に記載の原核生物をプロモーターAの転写を抑制した状態で炭素源の存在下で培養した後、プロモーターBの転写を誘導する処理を加えることによりリプレッサーRのタンパク質の発現量を増加させることにより、代謝物Mの生成量を減少させ、化合物Qの生成量を増加させることを特徴とする、化合物Qの製造方法。
- 請求項11から16のいずれか一項に記載の原核生物をプロモーターAの転写抑制を解除した状態で炭素源の存在下で培養した後、プロモーターBの転写を誘導する処理を加えることによりリプレッサーRのタンパク質の発現量を増加させることにより、代謝物Mの生成量を減少させ、化合物Qの生成量を増加させることを特徴とする、化合物Qの製造方法。
- 請求項11から16のいずれか一項に記載の原核生物を炭素源の存在下で培養した後、前記プロモーターCの転写を誘導する処理を加えることにより前記酵素Yの発現量を増加させることにより、化合物Qの生成量を増加させることを特徴とする、請求項17から19のいずれか一項に記載の化合物Qの製造方法。
- 前記化合物P、前記化合物Qおよび前記代謝物Mの組み合わせが、以下(j)から(w)のいずれかであることを特徴とする、請求項17から20のいずれか一項に記載の化合物Qの製造方法。
(j)化合物Pが3-デヒドロシキミ酸であり、化合物Qがプロトカテク酸であり、代謝物Mがシキミ酸である。
(k)化合物Pがコリスミン酸であり、化合物Qがアントラニル酸であり、代謝物Mがプレフェン酸である。
(l)化合物Pがプレフェン酸であり、化合物Qが4-ヒドロキシフェニルピルビン酸であり、代謝物Mがフェニルピルビン酸である。
(m)化合物Pがプレフェン酸であり、化合物Qがアロゲン酸であり、代謝物Mがフェニルピルビン酸である。
(n)化合物Pがプレフェン酸であり、化合物Qがフェニルピルビン酸であり、代謝物Mが4-ヒドロキシフェニルピルビン酸またはアロゲン酸である。
(o)化合物Pが2-オキソイソ吉草酸であり、化合物Qが2-イソプロピルリンゴ酸であ、代謝物Mがバリンである。
(p)化合物Pが2-オキソイソ吉草酸であり、化合物Qがバリンであり、代謝物Mが2-イソプロピルリンゴ酸である。
(q)化合物Pがグルタミン酸であり、化合物Qがγーグルタミルリン酸であり、代謝物MがN-アセチルグルタミン酸である。
(r)化合物Pがグルタミン酸であり、化合物QがN-アセチルグルタミン酸であり、代謝物Mがγーグルタミルリン酸である。
(s)化合物Pがアスパラギン酸であり、化合物Qがアスパラギンであり、代謝物Mがβ-アルパルチルリン酸である。
(t)化合物Pがアスパラギン酸β-セミアルデヒドあり、化合物Qが2,3-ジヒドロジピコリン酸であり、代謝物Mがホモセリンである。
(u)化合物Pがホモセリンあり、化合物QがO-アセチルホモセリンであり、代謝物Mがホモセリンリン酸である。
(v)化合物Pがホモセリンであり、化合物Qがホモセリンリン酸であり、代謝物MがO-アセチルホモセリンである。
(w)化合物Pがセリンであり、化合物QがO-アセチルセリンであり、代謝物Mがグリシンである。 - 前記酵素Xがシキミ酸脱水素酵素であり、かつ前記酵素Yが3-デヒドロシキミ酸脱水酵素であることを特徴とする、請求項21の(j)に記載のプロトカテク酸の製造方法。
- プロトカテク酸2,3-ジオキシゲナーゼをコードする遺伝子、プロトカテク酸3,4-ジオキシゲナーゼをコードする遺伝子、プロトカテク酸4,5-ジオキシゲナーゼをコードする遺伝子およびプロトカテク酸脱炭酸酵素をコードする遺伝子からなる群から選ばれる少なくとも1つの遺伝子について、該遺伝子の翻訳領域またはその転写・翻訳調節領域の中に、1個以上の塩基の置換、欠失または付加による変異を導入することにより、プロトカテク酸の代謝活性が欠損または低下している性質であることを特徴とする、請求項22に記載のプロトカテク酸の製造方法。
- コリネバクテリウム・グルタミカムまたはエッシェリヒア・コリである、請求項1から6のいずれか一項または請求項11から16のいずれか一項に記載の原核生物。
Priority Applications (6)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2014523757A JP6322576B2 (ja) | 2012-07-03 | 2013-07-03 | 有用微生物および目的物質の製造方法 |
| KR1020147035866A KR102139454B1 (ko) | 2012-07-03 | 2013-07-03 | 유용미생물 및 목적물질의 제조방법 |
| CN201380032679.6A CN104379729B (zh) | 2012-07-03 | 2013-07-03 | 有用微生物和目标物质的制备方法 |
| EP13812517.4A EP2851421B1 (en) | 2012-07-03 | 2013-07-03 | Useful microorganism, and method for producing substance of interest |
| US14/408,400 US10047382B2 (en) | 2012-07-03 | 2013-07-03 | Useful microorganism and method for producing substance of interest |
| US16/034,532 US10781461B2 (en) | 2012-07-03 | 2018-07-13 | Useful microorganism and method for producing substance of interest |
Applications Claiming Priority (2)
| Application Number | Priority Date | Filing Date | Title |
|---|---|---|---|
| JP2012149431 | 2012-07-03 | ||
| JP2012-149431 | 2012-07-03 |
Related Child Applications (2)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| US14/408,400 A-371-Of-International US10047382B2 (en) | 2012-07-03 | 2013-07-03 | Useful microorganism and method for producing substance of interest |
| US16/034,532 Continuation US10781461B2 (en) | 2012-07-03 | 2018-07-13 | Useful microorganism and method for producing substance of interest |
Publications (1)
| Publication Number | Publication Date |
|---|---|
| WO2014007273A1 true WO2014007273A1 (ja) | 2014-01-09 |
Family
ID=49882025
Family Applications (1)
| Application Number | Title | Priority Date | Filing Date |
|---|---|---|---|
| PCT/JP2013/068205 Ceased WO2014007273A1 (ja) | 2012-07-03 | 2013-07-03 | 有用微生物および目的物質の製造方法 |
Country Status (6)
| Country | Link |
|---|---|
| US (2) | US10047382B2 (ja) |
| EP (1) | EP2851421B1 (ja) |
| JP (1) | JP6322576B2 (ja) |
| KR (1) | KR102139454B1 (ja) |
| CN (1) | CN104379729B (ja) |
| WO (1) | WO2014007273A1 (ja) |
Cited By (4)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN103937734A (zh) * | 2014-04-23 | 2014-07-23 | 清华大学 | 一种高产透明质酸的基因工程菌及其应用 |
| JP2016003291A (ja) * | 2014-06-18 | 2016-01-12 | 住友ベークライト株式会社 | ポリエステル樹脂用単量体およびポリエステル樹脂 |
| US10961526B2 (en) | 2016-03-28 | 2021-03-30 | Research Institute Of Innovative Technology For The Earth | Transformant, and method for producing protocatechuic acid or salt thereof using same |
| WO2023112933A1 (ja) | 2021-12-13 | 2023-06-22 | 花王株式会社 | 新規プロモーター |
Families Citing this family (11)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| US8647642B2 (en) | 2008-09-18 | 2014-02-11 | Aviex Technologies, Llc | Live bacterial vaccines resistant to carbon dioxide (CO2), acidic PH and/or osmolarity for viral infection prophylaxis or treatment |
| US10047382B2 (en) | 2012-07-03 | 2018-08-14 | Kao Corporation | Useful microorganism and method for producing substance of interest |
| US10676723B2 (en) | 2015-05-11 | 2020-06-09 | David Gordon Bermudes | Chimeric protein toxins for expression by therapeutic bacteria |
| CN106190939B (zh) * | 2016-07-18 | 2020-04-07 | 清华大学 | 高产透明质酸的重组谷氨酸棒杆菌及其制备方法与应用 |
| US11180535B1 (en) | 2016-12-07 | 2021-11-23 | David Gordon Bermudes | Saccharide binding, tumor penetration, and cytotoxic antitumor chimeric peptides from therapeutic bacteria |
| US11129906B1 (en) | 2016-12-07 | 2021-09-28 | David Gordon Bermudes | Chimeric protein toxins for expression by therapeutic bacteria |
| CN106947727B (zh) * | 2017-02-20 | 2019-08-16 | 波顿(上海)生物技术有限公司 | 一种基因工程菌及其构建方法与生产香兰素的方法 |
| CN109456991B (zh) * | 2017-09-06 | 2023-05-09 | 华东师范大学 | 原儿茶酸调控的开关系统及其调控方法和应用 |
| CN108251346B (zh) * | 2018-01-15 | 2021-03-23 | 清华大学 | 一种表达透明质酸酶的重组谷氨酸棒杆菌及其应用 |
| CN110607313B (zh) * | 2019-09-27 | 2021-06-22 | 内蒙古伊品生物科技有限公司 | 一种高产l-赖氨酸的重组菌株及其构建方法与应用 |
| CN111471630B (zh) * | 2020-06-11 | 2021-11-02 | 鲁东大学 | 一株棒杆菌Ytld-phe09及其应用 |
Citations (15)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JPS5089592A (ja) | 1973-12-13 | 1975-07-18 | ||
| JPS5867699A (ja) | 1981-10-16 | 1983-04-22 | Ajinomoto Co Inc | プラスミド |
| JPS5877895A (ja) | 1981-11-02 | 1983-05-11 | Ajinomoto Co Inc | プラスミドphm1519 |
| JPS6098989A (ja) | 1983-11-04 | 1985-06-01 | Ajinomoto Co Inc | 発酵法によるプロトカテク酸の製造法 |
| JPH0549489A (ja) | 1991-08-22 | 1993-03-02 | Ajinomoto Co Inc | 発酵法によるl−フエニルアラニンの製造法 |
| JPH05304971A (ja) | 1992-01-30 | 1993-11-19 | Ajinomoto Co Inc | L−フェニルアラニンの製造法 |
| US5272073A (en) * | 1992-06-30 | 1993-12-21 | Purdue Research Foundation | Biocatalytic synthesis of catechol from glucose |
| US5821266A (en) | 1996-07-19 | 1998-10-13 | Michigan State University | Antioxidant activity of 3-dehydroshinkimates |
| JP2000295996A (ja) * | 1999-02-08 | 2000-10-24 | Kyowa Hakko Kogyo Co Ltd | プリンヌクレオチドの製造法 |
| US6472169B1 (en) | 1999-01-29 | 2002-10-29 | Board Of Trustees Operating Michigan State University | Biocatalytic synthesis of shikimic acid |
| JP2006508684A (ja) * | 2002-12-03 | 2006-03-16 | アクゾ・ノベル・エヌ・ベー | ロドコッカスからの新しい発現系 |
| JP2007300809A (ja) * | 2006-05-08 | 2007-11-22 | Yamaguchi Univ | シキミ酸の製造方法 |
| JP4305184B2 (ja) | 2001-11-23 | 2009-07-29 | 味の素株式会社 | エシェリヒア属細菌を用いたl−アミノ酸の製造方法 |
| JP2010207094A (ja) * | 2009-03-06 | 2010-09-24 | Genaris Inc | プロトカテク酸の製造法 |
| WO2011138006A1 (de) * | 2010-05-03 | 2011-11-10 | Forschungszentrum Jülich GmbH | Sensoren zur intrazellulären metabolit-detektion |
Family Cites Families (12)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JPS4539036B1 (ja) | 1967-06-09 | 1970-12-09 | ||
| US4305184A (en) | 1979-01-15 | 1981-12-15 | Bunker Ramo Corporation | Control of skew, bow and yield in circularly knit pile fabric |
| US5571697A (en) | 1989-05-05 | 1996-11-05 | Baylor College Of Medicine Texas Medical Center | Expression of processed recombinant lactoferrin and lactoferrin polypeptide fragments from a fusion product in Aspergillus |
| ZA932568B (en) | 1992-04-24 | 1993-11-12 | Baylor College Midecine A Non | Production of recombinant human lactoferrin |
| TWI222464B (en) | 1999-02-08 | 2004-10-21 | Kyowa Hakko Kogyo Kk | Process for producing purine nucleotides |
| AU2003256365B2 (en) | 2002-07-03 | 2010-02-18 | Pfenex Inc. | Benzoate- and anthranilate-inducible promoters |
| KR100725336B1 (ko) | 2005-12-26 | 2007-06-07 | 씨제이 주식회사 | 코리네형 세균에서 작동하는 신규 유도발현 벡터 시스템 |
| DE102006025821A1 (de) | 2006-06-02 | 2007-12-06 | Degussa Gmbh | Ein Enzym zur Herstellung von Mehylmalonatsemialdehyd oder Malonatsemialdehyd |
| DE102007005072A1 (de) | 2007-02-01 | 2008-08-07 | Evonik Degussa Gmbh | Verfahren zur fermentativen Herstellung von Cadaverin |
| KR20100031525A (ko) | 2007-06-01 | 2010-03-22 | 에보니크 룀 게엠베하 | 메타크릴산 또는 메타크릴산 에스테르의 제조 방법 |
| CN102498086A (zh) | 2009-01-22 | 2012-06-13 | 阿迈瑞斯公司 | 产生十二烷二酸及其衍生物的方法 |
| US10047382B2 (en) * | 2012-07-03 | 2018-08-14 | Kao Corporation | Useful microorganism and method for producing substance of interest |
-
2013
- 2013-07-03 US US14/408,400 patent/US10047382B2/en active Active
- 2013-07-03 EP EP13812517.4A patent/EP2851421B1/en active Active
- 2013-07-03 JP JP2014523757A patent/JP6322576B2/ja active Active
- 2013-07-03 WO PCT/JP2013/068205 patent/WO2014007273A1/ja not_active Ceased
- 2013-07-03 CN CN201380032679.6A patent/CN104379729B/zh active Active
- 2013-07-03 KR KR1020147035866A patent/KR102139454B1/ko active Active
-
2018
- 2018-07-13 US US16/034,532 patent/US10781461B2/en active Active
Patent Citations (15)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| JPS5089592A (ja) | 1973-12-13 | 1975-07-18 | ||
| JPS5867699A (ja) | 1981-10-16 | 1983-04-22 | Ajinomoto Co Inc | プラスミド |
| JPS5877895A (ja) | 1981-11-02 | 1983-05-11 | Ajinomoto Co Inc | プラスミドphm1519 |
| JPS6098989A (ja) | 1983-11-04 | 1985-06-01 | Ajinomoto Co Inc | 発酵法によるプロトカテク酸の製造法 |
| JPH0549489A (ja) | 1991-08-22 | 1993-03-02 | Ajinomoto Co Inc | 発酵法によるl−フエニルアラニンの製造法 |
| JPH05304971A (ja) | 1992-01-30 | 1993-11-19 | Ajinomoto Co Inc | L−フェニルアラニンの製造法 |
| US5272073A (en) * | 1992-06-30 | 1993-12-21 | Purdue Research Foundation | Biocatalytic synthesis of catechol from glucose |
| US5821266A (en) | 1996-07-19 | 1998-10-13 | Michigan State University | Antioxidant activity of 3-dehydroshinkimates |
| US6472169B1 (en) | 1999-01-29 | 2002-10-29 | Board Of Trustees Operating Michigan State University | Biocatalytic synthesis of shikimic acid |
| JP2000295996A (ja) * | 1999-02-08 | 2000-10-24 | Kyowa Hakko Kogyo Co Ltd | プリンヌクレオチドの製造法 |
| JP4305184B2 (ja) | 2001-11-23 | 2009-07-29 | 味の素株式会社 | エシェリヒア属細菌を用いたl−アミノ酸の製造方法 |
| JP2006508684A (ja) * | 2002-12-03 | 2006-03-16 | アクゾ・ノベル・エヌ・ベー | ロドコッカスからの新しい発現系 |
| JP2007300809A (ja) * | 2006-05-08 | 2007-11-22 | Yamaguchi Univ | シキミ酸の製造方法 |
| JP2010207094A (ja) * | 2009-03-06 | 2010-09-24 | Genaris Inc | プロトカテク酸の製造法 |
| WO2011138006A1 (de) * | 2010-05-03 | 2011-11-10 | Forschungszentrum Jülich GmbH | Sensoren zur intrazellulären metabolit-detektion |
Non-Patent Citations (9)
| Title |
|---|
| LI, K. ET AL.: "Microbial synthesis of 3-dehydroshikimic acid: a comparative analysis of D-xylose, L-arabinose, and D- glucose carbon sources", BIOTECHNOL. PROG., vol. 15, no. 5, 1999, pages 876 - 883, XP055178510 * |
| MORAWSKI, B. ET AL.: "Repression of Acinetobacter vanillate demethylase synthesis by VanR, a member of the GntR family of transcriptional regulators", FEMS MICROBIOL. LETT., vol. 187, 2000, pages 65 - 68, XP055178512 * |
| NUCLEIC ACIDS RES., vol. 16, 1988, pages 6127 |
| PROC. NATL. ACAD. SCI. U.S.A., vol. 69, 1972, pages 2110 |
| PROC. NATL. ACAD. SCI. U.S.A., vol. 74, 1977, pages 5463 |
| STRATAGENE, STRATEGIES, vol. 5, 1992, pages 81 |
| TERAMOTO, H. ET AL.: "Regulation of expression of genes involved in quinate and shikimate utilization in Corynebacterium glutamicum", APPL. ENVIRON. MICROBIOL., vol. 75, no. 11, 2009, pages 3461 - 3468, XP009173982 * |
| VAN DER REST ET AL., APPL. MICROBIOL. BIOTECHNOL., vol. 52, 1999, pages 541 |
| W. JAGER ET AL., J. BACTERIOL., vol. 174, 1992, pages 5462 |
Cited By (5)
| Publication number | Priority date | Publication date | Assignee | Title |
|---|---|---|---|---|
| CN103937734A (zh) * | 2014-04-23 | 2014-07-23 | 清华大学 | 一种高产透明质酸的基因工程菌及其应用 |
| CN103937734B (zh) * | 2014-04-23 | 2016-05-11 | 清华大学 | 一种生产透明质酸的基因工程菌及其应用 |
| JP2016003291A (ja) * | 2014-06-18 | 2016-01-12 | 住友ベークライト株式会社 | ポリエステル樹脂用単量体およびポリエステル樹脂 |
| US10961526B2 (en) | 2016-03-28 | 2021-03-30 | Research Institute Of Innovative Technology For The Earth | Transformant, and method for producing protocatechuic acid or salt thereof using same |
| WO2023112933A1 (ja) | 2021-12-13 | 2023-06-22 | 花王株式会社 | 新規プロモーター |
Also Published As
| Publication number | Publication date |
|---|---|
| US10781461B2 (en) | 2020-09-22 |
| KR102139454B1 (ko) | 2020-08-03 |
| US10047382B2 (en) | 2018-08-14 |
| EP2851421A4 (en) | 2015-11-25 |
| JPWO2014007273A1 (ja) | 2016-06-02 |
| KR20150027761A (ko) | 2015-03-12 |
| CN104379729B (zh) | 2020-07-21 |
| EP2851421B1 (en) | 2018-11-28 |
| JP6322576B2 (ja) | 2018-05-09 |
| CN104379729A (zh) | 2015-02-25 |
| US20180312882A1 (en) | 2018-11-01 |
| US20150197775A1 (en) | 2015-07-16 |
| EP2851421A1 (en) | 2015-03-25 |
Similar Documents
| Publication | Publication Date | Title |
|---|---|---|
| JP6322576B2 (ja) | 有用微生物および目的物質の製造方法 | |
| CN103403147B (zh) | 产腐胺的微生物以及使用此微生物生产腐胺的方法 | |
| CN112888776B (zh) | 生产目标物质的方法 | |
| US10612006B2 (en) | Method for producing aldehyde | |
| JP6297134B2 (ja) | プトレシン生産性を有する微生物及びそれを用いたプトレシン生産方法 | |
| US8455237B2 (en) | Escherichia coli strain with phosphoenolpyruvate carboxylase promoter replaced with cysteine synthase promoter and method of enhanced production of L-threonine using the same | |
| EP3423563B1 (en) | Improved muconic acid production from genetically engineered microorganisms | |
| KR19990077974A (ko) | L-글루타민산을생산하는박테리아및l-글루타민산을생산하는방법 | |
| US12600939B2 (en) | Mutant of Corynebacterium glutamicum with enhanced l-glutamic acid productivity and method for preparing l-glutamic acid using the same | |
| US6316232B1 (en) | Microbial preparation of substances from aromatic metabolism/I | |
| CN1289675C (zh) | 微生物制备芳族氨基酸的方法 | |
| JP4821606B2 (ja) | 効率的にL−グルタミン酸を生産するためのプロモーター変異による細菌のsucAB発現の最適化 | |
| TW201718863A (zh) | 用於製造腐胺或鳥胺酸的微生物及使用該微生物用於製造腐胺或鳥胺酸的方法 | |
| DE10219714A1 (de) | Verfahren zur mikrobielien Herstellung von aromatischen Aminosäuren und anderen Metaboliten des aromatischen Aminosäurebiosyntheseweges | |
| US20150111261A1 (en) | L-threonine-producing escherichia coli and method for producing l-threonine using the same | |
| JP7422888B2 (ja) | 新規なプロモーター及びその用途 | |
| JP4314897B2 (ja) | L−ロイシンの製造方法 | |
| CN106574232A (zh) | 具有l‑赖氨酸生产能力的微生物和使用其的l‑赖氨酸生产方法 | |
| WO2024005033A1 (ja) | 芳香族化合物の製造方法 | |
| KR20150028121A (ko) | Nad+ 의존성 포르메이트 데히드로게나제 유전자를 포함하는 코리네박테리움 및 이를 이용한 c4 디카르복실산 생산 방법 | |
| CN118599748A (zh) | 一种工程改造的盐单胞菌及其生产乙偶姻的方法 |
Legal Events
| Date | Code | Title | Description |
|---|---|---|---|
| 121 | Ep: the epo has been informed by wipo that ep was designated in this application |
Ref document number: 13812517 Country of ref document: EP Kind code of ref document: A1 |
|
| WWE | Wipo information: entry into national phase |
Ref document number: 14408400 Country of ref document: US Ref document number: 2013812517 Country of ref document: EP |
|
| ENP | Entry into the national phase |
Ref document number: 20147035866 Country of ref document: KR Kind code of ref document: A |
|
| ENP | Entry into the national phase |
Ref document number: 2014523757 Country of ref document: JP Kind code of ref document: A |
|
| NENP | Non-entry into the national phase |
Ref country code: DE |